

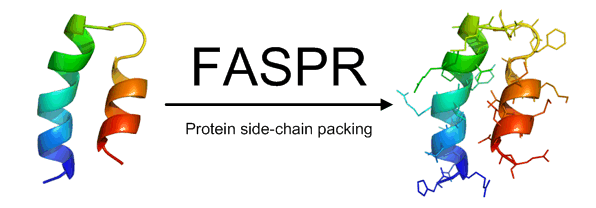
FASPR is a method for structural modeling of protein side-chain conformations.
Starting from a backbone structure, FASPR samples the side-chain rotamers for each amino acid
from the Dunbrack 2010 rotamer library with the atomic interaction energies calculated using an optimized
scoring function extended from EvoEF2,
where side-chain packing search is performed using a deterministic searching algorithm
combining self-energy checking, dead-end elimination theorems, and tree decomposition.
The large-scale benchmark tests showed that FASPR outperforms the current state-of-the-art
protein side-chain packers on both native and non-native backbones with higher accuracy
in terms of both side-chain dihedral angle (Χ1-4) recovery rate and RMSD with a much faster computational speed.
Please direct questions and inquiries to our Service System Discussion Board or contact Dr. Xiaoqiang Huang.