

You can:
| Name | 5-hydroxytryptamine receptor 1B |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | HTR1B |
| Synonym | 5-HT1B receptor 5-HT-1B 5-HT-1D-beta 5-HT1B Serotonin 1D beta receptor [ Show all ] |
| Disease | Chronic schizophrenics Major depressive disorder Migraine headaches Mood disorder Psychotic disorders [ Show all ] |
| Length | 390 |
| Amino acid sequence | MEEPGAQCAPPPPAGSETWVPQANLSSAPSQNCSAKDYIYQDSISLPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASILHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFFWRQAKAEEEVSECVVNTDHILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRILKQTPNRTGKRLTRAQLITDSPGSTSSVTSINSRVPDVPSESGSPVYVNQVKVRVSDALLEKKKLMAARERKATKTLGIILGAFIVCWLPFFIISLVMPICKDACWFHLAIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFKCTS |
| UniProt | P28222 |
| Protein Data Bank | 4iar, 6g79, 5v54 |
| GPCR-HGmod model | P28222 |
| 3D structure model | This structure is from PDB ID 4iar. |
| BioLiP | BL0239857, BL0403524,BL0403525, BL0417722 |
| Therapeutic Target Database | T07806 |
| ChEMBL | CHEMBL1898 |
| IUPHAR | 2 |
| DrugBank | BE0000797 |
| Name | sumatriptan |
|---|---|
| Molecular formula | C14H21N3O2S |
| IUPAC name | 1-[3-[2-(dimethylamino)ethyl]-1H-indol-5-yl]-N-methylmethanesulfonamide |
| Molecular weight | 295.401 |
| Hydrogen bond acceptor | 4 |
| Hydrogen bond donor | 2 |
| XlogP | 0.9 |
| Synonyms | Imigran Recovery L000584 (3-[2-(Dimethylamino)ethyl]-1H-indol-5-yl)-N-methylmethanesulfonamide NCGC00095838-03 103628-46-2 [ Show all ] |
| Inchi Key | KQKPFRSPSRPDEB-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C14H21N3O2S/c1-15-20(18,19)10-11-4-5-14-13(8-11)12(9-16-14)6-7-17(2)3/h4-5,8-9,15-16H,6-7,10H2,1-3H3 |
| PubChem CID | 5358 |
| ChEMBL | CHEMBL128 |
| IUPHAR | 54 |
| BindingDB | 50005835 |
| DrugBank | DB00669 |
Structure | 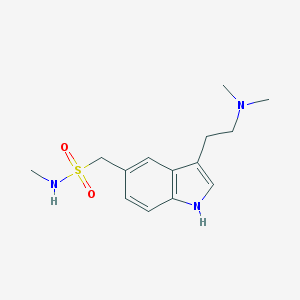 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| N/A | N/A | DrugBank | |
| EC50 | 38.0 nM | PMID9871581 | BindingDB |
| EC50 | 38.3 nM | PMID9871581 | ChEMBL |
| EC50 | 77.0 nM | PMID8960551 | BindingDB,ChEMBL |
| IC50 | 0.5012 nM | PMID11262079 | ChEMBL |
| IC50 | 10.0 nM | PMID10585208 | BindingDB,ChEMBL |
| IC50 | 11.0 nM | PMID10052975, PMID10052976 | BindingDB,ChEMBL |
| IC50 | 16.0 nM | PMID9357515, PMID9357514 | BindingDB,ChEMBL |
| IC50 | 27.0 nM | PMID9357515 | BindingDB,ChEMBL |
| Ki | 1.2 nM | PMID8935801 | PDSP,BindingDB |
| Ki | 1.99 nM | PMID7984267 | PDSP,BindingDB |
| Ki | 2.0 nM | PMID1315531 | PDSP,BindingDB |
| Ki | 3.4 nM | PMID1565658 | PDSP,BindingDB |
| Ki | 5.5 nM | PMID8071931 | BindingDB,ChEMBL |
| Ki | 7.7 nM | PMID14741277, PMID14505640 | BindingDB,ChEMBL |
| Ki | 7.76 nM | PMID7984267 | PDSP,BindingDB |
| Ki | 7.94 - 316.0 nM | PMID10611634, PMID10193663, PMID1565658, PMID8863519, PMID10513577, PMID11040052, PMID8967979 | IUPHAR |
| Ki | 9.1 nM | PMID18507369 | PDSP |
| Ki | 9.1 nM | PMID18507369 | BindingDB,ChEMBL |
| Ki | 9.33 nM | PMID7984267 | PDSP,BindingDB |
| Ki | 9.6 nM | PMID9986723 | BindingDB,ChEMBL |
| Ki | 10.0 nM | PMID8568822 | BindingDB,ChEMBL |
| Ki | 10.96 nM | PMID7984267 | PDSP,BindingDB |
| Ki | 19.0 nM | PMID9871581 | BindingDB |
| Ki | 19.1 nM | PMID9871581 | ChEMBL |
| Ki | 22.0 nM | PMID8941384 | BindingDB,ChEMBL |
| Ki | 23.0 nM | PMID7658447 | BindingDB |
| Ki | 23.1 nM | PMID8960551, PMID7658447 | BindingDB,ChEMBL |
| Ki | 24.0 nM | PMID8941384 | BindingDB,ChEMBL |
| Ki | 25.11 nM | PMID10611634 | PDSP,BindingDB |
| Ki | 28.0 nM | PMID9303569 | PDSP,BindingDB |
| Ki | 36.0 nM | PMID10853656 | BindingDB,ChEMBL |
| Ki | 47.9 nM | PMID10937729 | ChEMBL |
| Ki | 48.0 nM | PMID10937729 | BindingDB |
| Ki | 60.0 nM | PMID9303569 | PDSP,BindingDB |
| Ki | 61.65 nM | PMID7984267 | PDSP,BindingDB |
| pD2 | 5.7 - | PMID9871581 | ChEMBL |
| Selectivity | 2.2 - | PMID14505640 | ChEMBL |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218