

You can:
| Name | Sphingosine 1-phosphate receptor 3 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | S1PR3 |
| Synonym | Sphingosine 1-phosphate receptor Edg-3 S1P3 receptor S1P3 S1P receptor Edg-3 S1P receptor 3 [ Show all ] |
| Disease | Breast cancer |
| Length | 378 |
| Amino acid sequence | MATALPPRLQPVRGNETLREHYQYVGKLAGRLKEASEGSTLTTVLFLVICSFIVLENLMVLIAIWKNNKFHNRMYFFIGNLALCDLLAGIAYKVNILMSGKKTFSLSPTVWFLREGSMFVALGASTCSLLAIAIERHLTMIKMRPYDANKRHRVFLLIGMCWLIAFTLGALPILGWNCLHNLPDCSTILPLYSKKYIAFCISIFTAILVTIVILYARIYFLVKSSSRKVANHNNSERSMALLRTVVIVVSVFIACWSPLFILFLIDVACRVQACPILFKAQWFIVLAVLNSAMNPVIYTLASKEMRRAFFRLVCNCLVRGRGARASPIQPALDPSRSKSSSSNNSSHSPKVKEDLPHTAPSSCIMDKNAALQNGIFCN |
| UniProt | Q99500 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q99500 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q99500. |
| BioLiP | N/A |
| Therapeutic Target Database | T11241 |
| ChEMBL | CHEMBL3892 |
| IUPHAR | 277 |
| DrugBank | N/A |
| Name | (S) FTY720 Phosphate |
|---|---|
| Molecular formula | C19H34NO5P |
| IUPAC name | [(2S)-2-amino-2-(hydroxymethyl)-4-(4-octylphenyl)butyl] dihydrogen phosphate |
| Molecular weight | 387.457 |
| Hydrogen bond acceptor | 6 |
| Hydrogen bond donor | 4 |
| XlogP | 0.8 |
| Synonyms | (S)-Fty 720P 2-Amino-2-[2-(4-octylphenyl)ethyl]-1,3-propanediol 1-(Dihydrogen Phosphate) BDBM23165 FT-0668884 SCHEMBL4267565 [ Show all ] |
| Inchi Key | LRFKWQGGENFBFO-IBGZPJMESA-N |
| Inchi ID | InChI=1S/C19H34NO5P/c1-2-3-4-5-6-7-8-17-9-11-18(12-10-17)13-14-19(20,15-21)16-25-26(22,23)24/h9-12,21H,2-8,13-16,20H2,1H3,(H2,22,23,24)/t19-/m0/s1 |
| PubChem CID | 11452022 |
| ChEMBL | CHEMBL366208 |
| IUPHAR | 6996 |
| BindingDB | 23165 |
| DrugBank | N/A |
Structure | 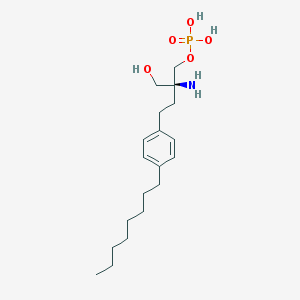 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| EC50 | 0.134 nM | PMID20188554 | BindingDB,ChEMBL |
| EC50 | 2.0 nM | PMID24909680 | ChEMBL |
| EC50 | 2.0 nM | PMID24909680 | BindingDB |
| EC50 | 3.0 nM | PMID22583616 | BindingDB,ChEMBL |
| EC50 | 3.1 nM | PMID16078855 | BindingDB,ChEMBL |
| EC50 | 3.1 nM | PMID17114004 | IUPHAR |
| EC50 | 3.3 nM | PMID24900318, PMID22405291 | BindingDB,ChEMBL |
| EC50 | 5.0 nM | PMID26751273 | BindingDB |
| EC50 | 5.01 nM | PMID21838322 | BindingDB |
| EC50 | 5.012 nM | PMID21838322, PMID26751273 | ChEMBL |
| EC50 | 27.0 nM | PMID22104144 | BindingDB,ChEMBL |
| EC50 | 27.84 nM | PMID20188554 | BindingDB,ChEMBL |
| EC50 | 94.0 nM | PMID26687487 | BindingDB,ChEMBL |
| Efficacy | 35.0 % | PMID22104144 | ChEMBL |
| Emax | 62.0 % | PMID21838322 | ChEMBL |
| IC50 | 6.3 nM | PMID24900286 | BindingDB,ChEMBL |
| IC50 | 23.0 nM | PMID22119341 | BindingDB,ChEMBL |
| Ki | 5.9 nM | PMID15598563 | BindingDB |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218