

You can:
| Name | Neuropeptide S receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | NPSR1 |
| Synonym | G-protein coupled receptor for asthma susceptibility vasopressin receptor-related receptor 1 G protein-coupled receptor 154 PGR14 NPS receptor [ Show all ] |
| Disease | Neurological disease |
| Length | 371 |
| Amino acid sequence | MPANFTEGSFDSSGTGQTLDSSPVACTETVTFTEVVEGKEWGSFYYSFKTEQLITLWVLFVFTIVGNSVVLFSTWRRKKKSRMTFFVTQLAITDSFTGLVNILTDINWRFTGDFTAPDLVCRVVRYLQVVLLYASTYVLVSLSIDRYHAIVYPMKFLQGEKQARVLIVIAWSLSFLFSIPTLIIFGKRTLSNGEVQCWALWPDDSYWTPYMTIVAFLVYFIPLTIISIMYGIVIRTIWIKSKTYETVISNCSDGKLCSSYNRGLISKAKIKAIKYSIIIILAFICCWSPYFLFDILDNFNLLPDTQERFYASVIIQNLPALNSAINPLIYCVFSSSISFPCREQRSQDSRMTFRERTERHEMQILSKPEFI |
| UniProt | Q6W5P4 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q6W5P4 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q6W5P4. |
| BioLiP | N/A |
| Therapeutic Target Database | T20958 |
| ChEMBL | CHEMBL5162 |
| IUPHAR | 302 |
| DrugBank | BE0000948 |
| Name | MLS000587548 |
|---|---|
| Molecular formula | C24H22Cl2N2O2 |
| IUPAC name | (7E)-2-amino-4-(2-chlorophenyl)-7-[(2-chlorophenyl)methylidene]-5,6-dihydro-4H-cyclopenta[b]pyran-3-carbonitrile;ethanol |
| Molecular weight | 441.352 |
| Hydrogen bond acceptor | 4 |
| Hydrogen bond donor | 2 |
| XlogP | None |
| Synonyms | AKOS024302050 HMS2612N09 2-Amino-4-(2-chloro-phenyl)-7-[1-(2-chloro-phenyl)-meth-(E)-ylidene]-4,5,6,7-tetrahydro-cyclopenta[b]pyran-3-carbonitrile CHEMBL1537070 SMR000211567 |
| Inchi Key | ADCNYPCMZJMRFF-JHGYPSGKSA-N |
| Inchi ID | InChI=1S/C22H16Cl2N2O.C2H6O/c23-18-7-3-1-5-13(18)11-14-9-10-16-20(15-6-2-4-8-19(15)24)17(12-25)22(26)27-21(14)16;1-2-3/h1-8,11,20H,9-10,26H2;3H,2H2,1H3/b14-11+; |
| PubChem CID | 12005524 |
| ChEMBL | CHEMBL1537070 |
| IUPHAR | N/A |
| BindingDB | N/A |
| DrugBank | N/A |
Structure | 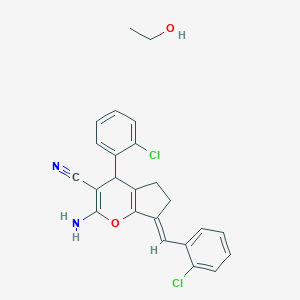 |
| Lipinski's druglikeness | Partition coefficient log P of this ligand is not available. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| Potency | 10000.0 nM | PubChem BioAssay data set | ChEMBL |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218