

You can:
| Name | CHEMBL574212 |
|---|---|
| Molecular formula | C60H69ClN10O8S |
| IUPAC name | (2Z)-2-[(2E,4E)-5-[1-[6-[2-[5-[6-[(3-chlorophenyl)methylamino]-9-[(1S,2R,3S,4R,5R)-3,4-dihydroxy-5-(methylcarbamoyl)-2-bicyclo[3.1.0]hexanyl]purin-2-yl]pent-4-ynoylamino]ethylamino]-6-oxohexyl]-3,3-dimethylindol-1-ium-2-yl]penta-2,4-dienylidene]-1-ethyl-3,3-dimethylindole-5-sulfonate |
| Molecular weight | 1125.78 |
| Hydrogen bond acceptor | 13 |
| Hydrogen bond donor | 6 |
| XlogP | 6.3 |
| Synonyms | N/A |
| Inchi Key | BJRYADSSKIBSOD-HMXAIGQHSA-N |
| Inchi ID | InChI=1S/C60H69ClN10O8S/c1-7-69-45-29-28-40(80(77,78)79)34-42(45)59(4,5)46(69)23-10-8-11-24-47-58(2,3)41-21-13-14-22-44(41)70(47)32-17-9-12-26-49(72)63-30-31-64-50(73)27-16-15-25-48-67-55(65-36-38-19-18-20-39(61)33-38)51-56(68-48)71(37-66-51)52-43-35-60(43,57(76)62-6)54(75)53(52)74/h8,10-11,13-14,18-24,28-29,33-34,37,43,52-54,74-75H,7,9,12,16-17,26-27,30-32,35-36H2,1-6H3,(H4-,62,63,64,65,67,68,72,73,76,77,78,79)/t43-,52-,53+,54+,60-/m1/s1 |
| PubChem CID | 45483968 |
| ChEMBL | CHEMBL574212 |
| IUPHAR | N/A |
| BindingDB | 50300278 |
| DrugBank | N/A |
Structure | 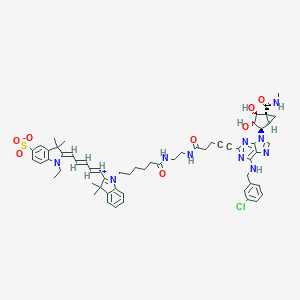 |
| Lipinski's druglikeness | This ligand has more than 5 hydrogen bond donor. This ligand has more than 10 hydrogen bond acceptor. This ligand is heavier than 500 daltons. This ligand has a partition coefficient log P greater than 5. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| Ki | 4730.0 nM | PMID19499950 | BindingDB,ChEMBL |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218