

You can:
| Name | Vasoactive intestinal polypeptide receptor 1 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | VIPR1 |
| Synonym | VPAC1 VIP-R-1 VIP and PACAP receptor 1 VIP-R1 RDC1 [ Show all ] |
| Disease | Respiratory disease Chronic obstructive pulmonary disease; Pulmonary arterial hypertension |
| Length | 457 |
| Amino acid sequence | MRPPSPLPARWLCVLAGALAWALGPAGGQAARLQEECDYVQMIEVQHKQCLEEAQLENETIGCSKMWDNLTCWPATPRGQVVVLACPLIFKLFSSIQGRNVSRSCTDEGWTHLEPGPYPIACGLDDKAASLDEQQTMFYGSVKTGYTIGYGLSLATLLVATAILSLFRKLHCTRNYIHMHLFISFILRAAAVFIKDLALFDSGESDQCSEGSVGCKAAMVFFQYCVMANFFWLLVEGLYLYTLLAVSFFSERKYFWGYILIGWGVPSTFTMVWTIARIHFEDYGCWDTINSSLWWIIKGPILTSILVNFILFICIIRILLQKLRPPDIRKSDSSPYSRLARSTLLLIPLFGVHYIMFAFFPDNFKPEVKMVFELVVGSFQGFVVAILYCFLNGEVQAELRRKWRRWHLQGVLGWNPKYRHPSGGSNGATCSTQVSMLTRVSPGARRSSSFQAEVSLV |
| UniProt | P32241 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P32241 |
| 3D structure model | This predicted structure model is from GPCR-EXP P32241. |
| BioLiP | N/A |
| Therapeutic Target Database | T86364 |
| ChEMBL | CHEMBL5144 |
| IUPHAR | 371 |
| DrugBank | N/A |
| Name | CHEMBL2087421 |
|---|---|
| Molecular formula | C24H40N4O5S |
| IUPAC name | 4-methoxy-N,2,6-trimethyl-N-[2-[2-[4-(1-methylpiperidin-4-yl)piperazin-1-yl]-2-oxoethoxy]ethyl]benzenesulfonamide |
| Molecular weight | 496.667 |
| Hydrogen bond acceptor | 8 |
| Hydrogen bond donor | 0 |
| XlogP | 1.6 |
| Synonyms | BDBM50420580 SCHEMBL3997134 |
| Inchi Key | TYDKAHFZLFMNQX-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C24H40N4O5S/c1-19-16-22(32-5)17-20(2)24(19)34(30,31)26(4)14-15-33-18-23(29)28-12-10-27(11-13-28)21-6-8-25(3)9-7-21/h16-17,21H,6-15,18H2,1-5H3 |
| PubChem CID | 11340940 |
| ChEMBL | CHEMBL2087421 |
| IUPHAR | N/A |
| BindingDB | 50420580 |
| DrugBank | N/A |
Structure | 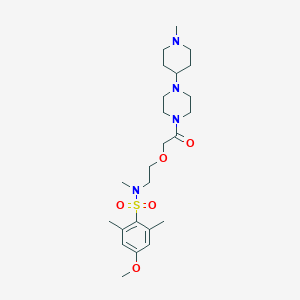 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| Inhibition | 5.0 % | PMID22369198 | ChEMBL |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218