

You can:
| Name | Kappa-type opioid receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | OPRK1 |
| Synonym | K-OR-1 KOPr OP2 KOP KOR-1 [ Show all ] |
| Disease | Obesity Opiate dependence Inflammatory bowel disease Erythema Diarrhea-predominant IBS [ Show all ] |
| Length | 380 |
| Amino acid sequence | MDSPIQIFRGEPGPTCAPSACLPPNSSAWFPGWAEPDSNGSAGSEDAQLEPAHISPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRYTKMKTATNIYIFNLALADALVTTTMPFQSTVYLMNSWPFGDVLCKIVISIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVREDVDVIECSLQFPDDDYSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLSGSREKDRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSTSHSTAALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFPLKMRMERQSTSRVRNTVQDPAYLRDIDGMNKPV |
| UniProt | P41145 |
| Protein Data Bank | 6b73, 4djh |
| GPCR-HGmod model | P41145 |
| 3D structure model | This structure is from PDB ID 6b73. |
| BioLiP | BL0402244,BL0402246, BL0224693,BL0224694, BL0402243,BL0402245 |
| Therapeutic Target Database | T60693 |
| ChEMBL | CHEMBL237 |
| IUPHAR | 318 |
| DrugBank | BE0000632 |
| Name | naloxone |
|---|---|
| Molecular formula | C19H21NO4 |
| IUPAC name | (4R,4aS,7aR,12bS)-4a,9-dihydroxy-3-prop-2-enyl-2,4,5,6,7a,13-hexahydro-1H-4,12-methanobenzofuro[3,2-e]isoquinolin-7-one |
| Molecular weight | 327.38 |
| Hydrogen bond acceptor | 5 |
| Hydrogen bond donor | 2 |
| XlogP | 2.1 |
| Synonyms | HSDB 3279 MLS000069540 (4R,4aS,7aR,12bS)-4a,9-dihydroxy-3-prop-2-enyl-2,4,5,6,7a,13-hexahydro-1H-4,12-methanobenzofuro[3,2-e]isoquinoline-7-one;hydrochloride Morphinan-6-one, 4,5-epoxy-3,14-dihydroxy-17-(2-propenyl)-, (5alpha)-(9CI) 12-Allyl-7,7a,8,9-tetrahydro-3,7a-dihydroxy-4aH-8,9c-iminoethanophenanthro(4,5-bcd)furanone [ Show all ] |
| Inchi Key | UZHSEJADLWPNLE-GRGSLBFTSA-N |
| Inchi ID | InChI=1S/C19H21NO4/c1-2-8-20-9-7-18-15-11-3-4-12(21)16(15)24-17(18)13(22)5-6-19(18,23)14(20)10-11/h2-4,14,17,21,23H,1,5-10H2/t14-,17+,18+,19-/m1/s1 |
| PubChem CID | 5284596 |
| ChEMBL | CHEMBL80 |
| IUPHAR | 1676, 1638 |
| BindingDB | 50000788, 54795 |
| DrugBank | DB01183 |
Structure | 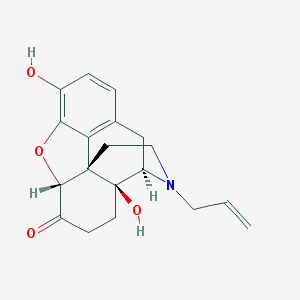 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| N/A | N/A | DrugBank | |
| Activity | 51.0 - | PMID7515442 | ChEMBL |
| Activity | 101.6 % | PMID26035635, PMID21667972 | ChEMBL |
| EC50 | 6.6 nM | PMID12747782 | BindingDB,ChEMBL |
| EC50 | 10.0 nM | PMID17407276 | BindingDB,ChEMBL |
| Emax | 25.0 % | PMID17407276 | ChEMBL |
| IC50 | 1.5 nM | PMID11585443 | BindingDB,ChEMBL |
| IC50 | 49.8 nM | PMID12747782 | ChEMBL |
| IC50 | 50.0 nM | PMID12747782 | BindingDB |
| IC50 | 75.0 nM | DrugMatrix in vitro pharmacology data | ChEMBL |
| IC50 | 182.0 nM | N/A | BindingDB |
| IC50 | 320.0 nM | PMID17407276 | BindingDB,ChEMBL |
| IC50 | 616.0 nM | N/A | BindingDB |
| IC50 | 704.0 nM | N/A | BindingDB |
| Imax | 55.0 % | PMID17407276 | ChEMBL |
| Inhibition | 105.6 % | PMID23659286 | ChEMBL |
| Ke | 10.0 nM | PMID20055417 | ChEMBL |
| Ke | 11.0 nM | PMID21570305 | ChEMBL |
| Ki | 0.25 nM | PMID21482470 | ChEMBL |
| Ki | 0.25 nM | PMID21482470 | BindingDB |
| Ki | 1.1 nM | PMID17407276 | BindingDB,ChEMBL |
| Ki | 1.2 nM | PMID10741545, PMID19027293 | BindingDB,ChEMBL |
| Ki | 2.3 nM | PMID8114680 | BindingDB |
| Ki | 2.5 nM | PMID9686407 | BindingDB |
| Ki | 2.51188 - 25.1189 nM | PMID9686407, PMID7869844, PMID7624359, PMID9262330 | IUPHAR |
| Ki | 3.0 nM | PMID20055417 | BindingDB,ChEMBL |
| Ki | 4.467 nM | PMID19527931 | ChEMBL |
| Ki | 4.5 nM | PMID19527931 | BindingDB |
| Ki | 9.2 nM | PMID18313920, PMID14643346, PMID17149859, PMID17149858 | BindingDB,ChEMBL |
| Ki | 9.2 nM | PMID14643346 | BindingDB |
| Ki | 10.7 nM | PMID12747782 | ChEMBL |
| Ki | 11.0 nM | PMID12747782 | BindingDB |
| Ki | 13.8 nM | PMID7932177 | BindingDB |
| Ki | 14.7 nM | PMID20441176 | ChEMBL |
| Ki | 15.0 nM | PMID20441176, MedChemComm, (2016) 7:2:317 | BindingDB,ChEMBL |
| Ki | 30.0 nM | DrugMatrix in vitro pharmacology data | ChEMBL |
| Ki | 362.0 nM | PMID25268943 | BindingDB,ChEMBL |
| Ki | 1000.0 nM | PMID7932177 | BindingDB |
| Max inhibition | 98.0 % | PMID12747782 | ChEMBL |
| Max stimulation | 30.0 % | PMID12747782 | ChEMBL |
| pKb | 7.14 - | PMID19527931 | ChEMBL |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218