

You can:
| Name | 5-hydroxytryptamine receptor 2A |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Htr2a |
| Synonym | serotonin 5HT-2 receptor 5Ht-2 'D' receptor 5-hydroxytryptamine (serotonin) receptor 2A, G protein-coupled 5-HT2A receptor [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 471 |
| Amino acid sequence | MEILCEDNISLSSIPNSLMQLGDGPRLYHNDFNSRDANTSEASNWTIDAENRTNLSCEGYLPPTCLSILHLQEKNWSALLTTVVIILTIAGNILVIMAVSLEKKLQNATNYFLMSLAIADMLLGFLVMPVSMLTILYGYRWPLPSKLCAIWIYLDVLFSTASIMHLCAISLDRYVAIQNPIHHSRFNSRTKAFLKIIAVWTISVGISMPIPVFGLQDDSKVFKEGSCLLADDNFVLIGSFVAFFIPLTIMVITYFLTIKSLQKEATLCVSDLSTRAKLASFSFLPQSSLSSEKLFQRSIHREPGSYAGRRTMQSISNEQKACKVLGIVFFLFVVMWCPFFITNIMAVICKESCNENVIGALLNVFVWIGYLSSAVNPLVYTLFNKTYRSAFSRYIQCQYKENRKPLQLILVNTIPALAYKSSQLQVGQKKNSQEDAEQTVDDCSMVTLGKQQSEENCTDNIETVNEKVSCV |
| UniProt | P14842 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL322 |
| IUPHAR | 6 |
| DrugBank | N/A |
| Name | tryptamine |
|---|---|
| Molecular formula | C10H12N2 |
| IUPAC name | 2-(1H-indol-3-yl)ethanamine |
| Molecular weight | 160.22 |
| Hydrogen bond acceptor | 1 |
| Hydrogen bond donor | 2 |
| XlogP | 1.6 |
| Synonyms | VI30132 343-94-2 (mono-hydrochloride) MFCD00005661 AB1001199 NCGC00014994-06 [ Show all ] |
| Inchi Key | APJYDQYYACXCRM-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C10H12N2/c11-6-5-8-7-12-10-4-2-1-3-9(8)10/h1-4,7,12H,5-6,11H2 |
| PubChem CID | 1150 |
| ChEMBL | CHEMBL6640 |
| IUPHAR | 125 |
| BindingDB | 50024210 |
| DrugBank | N/A |
Structure | 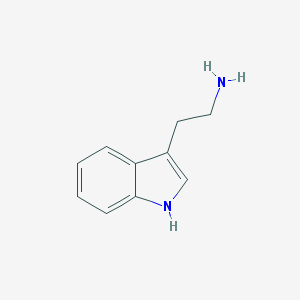 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| Ki | <10000.0 nM | PMID8510008 | BindingDB |
| Ki | 11.0 nM | PMID2795135 | BindingDB |
| Ki | 25.11 nM | PMID7984267 | BindingDB |
| Ki | 79.4328 nM | PMID8114677 | IUPHAR |
| Ki | 416.86 nM | PMID9225287 | BindingDB |
| Ki | 538.0 nM | PMID6645787 | BindingDB |
| Ki | 1682.0 nM | PMID6645787 | BindingDB |
| Ki | 2932.0 nM | PMID6645787 | BindingDB |
| Ki | 3890.45 nM | PMID7984267 | BindingDB |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218