

You can:
| Name | Corticotropin-releasing factor receptor 1 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | CRHR1 |
| Synonym | CRHR CRH-R1 CRH-R-1 CRFR1 CRFR-1 [ Show all ] |
| Disease | Major depressive disorder; Severe mood disorder Depression; Anxiety Depression Irritable bowel syndrome Anxiety disorder; Depression [ Show all ] |
| Length | 444 |
| Amino acid sequence | MGGHPQLRLVKALLLLGLNPVSASLQDQHCESLSLASNISGLQCNASVDLIGTCWPRSPAGQLVVRPCPAFFYGVRYNTTNNGYRECLANGSWAARVNYSECQEILNEEKKSKVHYHVAVIINYLGHCISLVALLVAFVLFLRLRPGCTHWGDQADGALEVGAPWSGAPFQVRRSIRCLRNIIHWNLISAFILRNATWFVVQLTMSPEVHQSNVGWCRLVTAAYNYFHVTNFFWMFGEGCYLHTAIVLTYSTDRLRKWMFICIGWGVPFPIIVAWAIGKLYYDNEKCWFGKRPGVYTDYIYQGPMILVLLINFIFLFNIVRILMTKLRASTTSETIQYRKAVKATLVLLPLLGITYMLFFVNPGEDEVSRVVFIYFNSFLESFQGFFVSVFYCFLNSEVRSAIRKRWHRWQDKHSIRARVARAMSIPTSPTRVSFHSIKQSTAV |
| UniProt | P34998 |
| Protein Data Bank | 4z9g, 4k5y |
| GPCR-HGmod model | P34998 |
| 3D structure model | This structure is from PDB ID 4z9g. |
| BioLiP | BL0350036,BL0350037,BL0350038, BL0251208 |
| Therapeutic Target Database | T45262 |
| ChEMBL | CHEMBL1800 |
| IUPHAR | 212 |
| DrugBank | BE0008658 |
| Name | 195055-03-9 |
|---|---|
| Molecular formula | C22H32N6 |
| IUPAC name | 3-[6-(dimethylamino)-4-methylpyridin-3-yl]-2,5-dimethyl-N,N-dipropylpyrazolo[1,5-a]pyrimidin-7-amine |
| Molecular weight | 380.54 |
| Hydrogen bond acceptor | 5 |
| Hydrogen bond donor | 0 |
| XlogP | 4.7 |
| Synonyms | 292073-75-7 BCP23964 FT-0767007 NBI-30775 UNII-G82N555U1N [ Show all ] |
| Inchi Key | ANNRUWYFVIGKHA-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C22H32N6/c1-8-10-27(11-9-2)20-13-16(4)24-22-21(17(5)25-28(20)22)18-14-23-19(26(6)7)12-15(18)3/h12-14H,8-11H2,1-7H3 |
| PubChem CID | 9821250 |
| ChEMBL | CHEMBL309138 |
| IUPHAR | 3520 |
| BindingDB | 50116105 |
| DrugBank | N/A |
Structure | 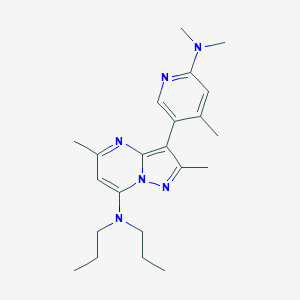 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| IC50 | 4.0 nM | PMID22971011 | BindingDB,ChEMBL |
| IC50 | 8.5 nM | PMID26901666 | BindingDB,ChEMBL |
| IC50 | 28.18 nM | PMID18989952 | BindingDB,ChEMBL |
| IC50 | 50.0 nM | PMID15341493, PMID16078829 | BindingDB,ChEMBL |
| Imax | 93.0 % | PMID21889333 | ChEMBL |
| Kd | 0.303 nM | PMID21889333 | BindingDB |
| Kd | 0.303 nM | PMID21889333 | ChEMBL |
| Ki | 1.0 - 5.0 nM | PMID10867111 | IUPHAR |
| Ki | 2.6 nM | PMID26456805 | BindingDB,ChEMBL |
| Ki | 3.0 nM | PMID26456805, PMID18672365, PMID21618986 | BindingDB,ChEMBL |
| Ki | 3.0 nM | PMID26456805 | BindingDB |
| Ki | 3.5 nM | PMID15341493 | BindingDB,ChEMBL |
| Ki | 3.98 nM | PMID21074436 | BindingDB |
| Ki | 3.981 nM | PMID21074436 | ChEMBL |
| Ki | 5.0 nM | PMID12127521 | BindingDB,ChEMBL |
| Ki | 7.8 nM | PMID15943483 | BindingDB,ChEMBL |
| Ki | 8.3 nM | PMID22587443 | BindingDB,ChEMBL |
| Ki | 12.0 nM | PMID21889333 | BindingDB,ChEMBL |
| T1/2 | 12.8 hr | PMID21889333 | ChEMBL |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218