

You can:
| Name | Free fatty acid receptor 3 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | FFAR3 |
| Synonym | FFA3R FFA3 receptor G-protein coupled receptor 41 GPCR41 GPR41 [ Show all ] |
| Disease | N/A |
| Length | 346 |
| Amino acid sequence | MDTGPDQSYFSGNHWFVFSVYLLTFLVGLPLNLLALVVFVGKLQRRPVAVDVLLLNLTASDLLLLLFLPFRMVEAANGMHWPLPFILCPLSGFIFFTTIYLTALFLAAVSIERFLSVAHPLWYKTRPRLGQAGLVSVACWLLASAHCSVVYVIEFSGDISHSQGTNGTCYLEFRKDQLAILLPVRLEMAVVLFVVPLIITSYCYSRLVWILGRGGSHRRQRRVAGLLAATLLNFLVCFGPYNVSHVVGYICGESPAWRIYVTLLSTLNSCVDPFVYYFSSSGFQADFHELLRRLCGLWGQWQQESSMELKEQKGGEEQRADRPAERKTSEHSQGCGTGGQVACAES |
| UniProt | O14843 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | O14843 |
| 3D structure model | This predicted structure model is from GPCR-EXP O14843. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL5201 |
| IUPHAR | 227 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 8967 | 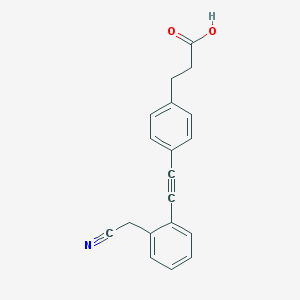 CHEMBL2315528 CHEMBL2315528 | C19H15NO2 | 289.334 | 3 / 1 | 3.4 | Yes |
| 467070 | 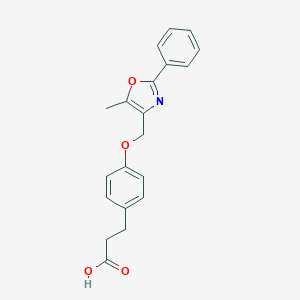 CHEMBL3600993 CHEMBL3600993 | C20H19NO4 | 337.375 | 5 / 1 | 3.7 | Yes |
| 36308 | 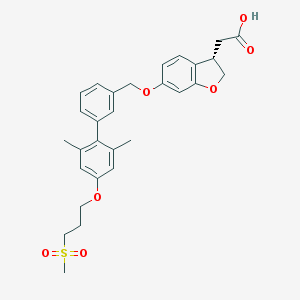 TAK-875 TAK-875 | C29H32O7S | 524.628 | 7 / 1 | 4.7 | No |
| 59168 | 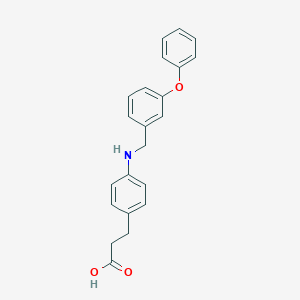 GW9508 GW9508 | C22H21NO3 | 347.414 | 4 / 2 | 4.7 | Yes |
| 553549 | 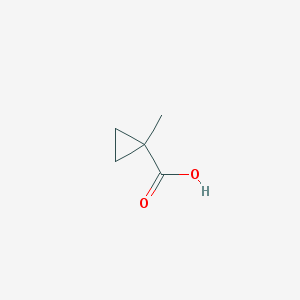 1-Methylcyclopropane-1-carboxylic acid 1-Methylcyclopropane-1-carboxylic acid | C5H8O2 | 100.117 | 2 / 1 | 0.6 | Yes |
| 537866 | 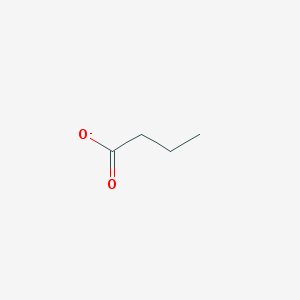 butyrate butyrate | C4H7O2- | 87.098 | 2 / 0 | 1.4 | Yes |
| 553650 | 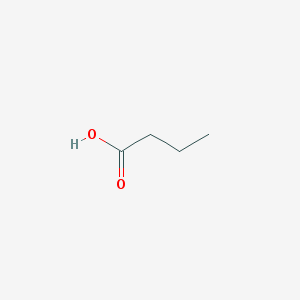 butyric acid butyric acid | C4H8O2 | 88.106 | 2 / 1 | 0.8 | Yes |
| 83244 | 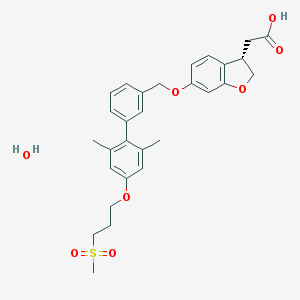 CHEMBL2047159 CHEMBL2047159 | C29H34O8S | 542.643 | 8 / 2 | N/A | No |
| 445087 | 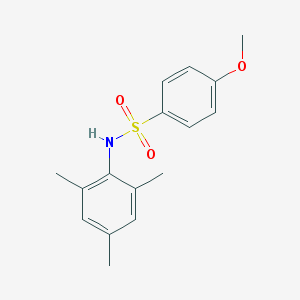 GSK137647A GSK137647A | C16H19NO3S | 305.392 | 4 / 1 | 3.5 | Yes |
| 525476 | 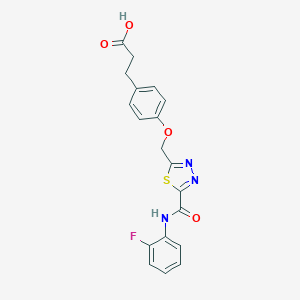 CHEMBL3805888 CHEMBL3805888 | C19H16FN3O4S | 401.412 | 8 / 2 | 2.9 | Yes |
| 550039 | 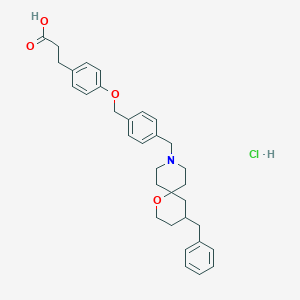 CHEMBL3963542 CHEMBL3963542 | C33H40ClNO4 | 550.136 | 5 / 2 | N/A | No |
| 554168 | 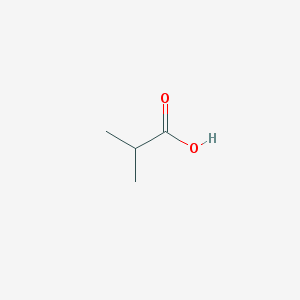 ISOBUTYRIC ACID ISOBUTYRIC ACID | C4H8O2 | 88.106 | 2 / 1 | 0.8 | Yes |
| 485512 | 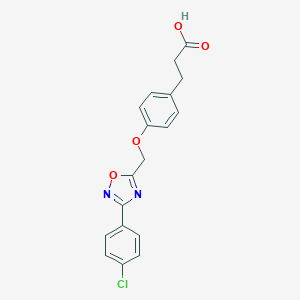 CHEMBL3601003 CHEMBL3601003 | C18H15ClN2O4 | 358.778 | 6 / 1 | 3.7 | Yes |
| 526754 | 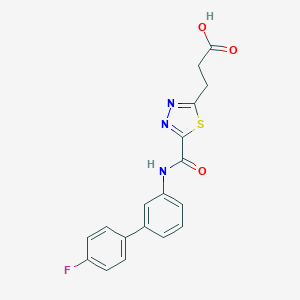 CHEMBL3805879 CHEMBL3805879 | C18H14FN3O3S | 371.386 | 7 / 2 | 3.1 | Yes |
| 550349 | 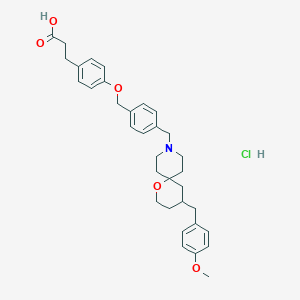 CHEMBL3893207 CHEMBL3893207 | C34H42ClNO5 | 580.162 | 6 / 2 | N/A | No |
| 527395 | 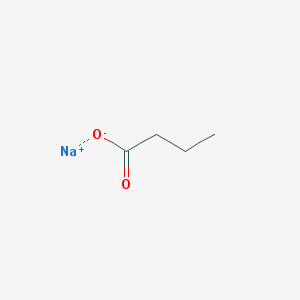 SODIUM BUTYRATE SODIUM BUTYRATE | C4H7NaO2 | 110.088 | 2 / 0 | N/A | N/A |
| 554419 | 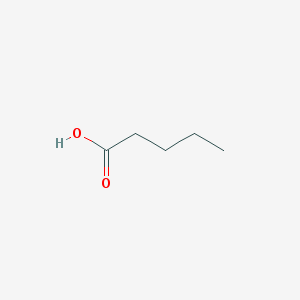 Valeric acid Valeric acid | C5H10O2 | 102.133 | 2 / 1 | 1.4 | Yes |
| 554692 | 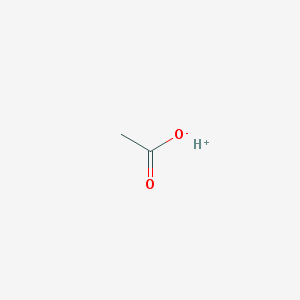 CID 21980959 CID 21980959 | C2H4O2 | 60.052 | 2 / 1 | N/A | N/A |
| 551665 | 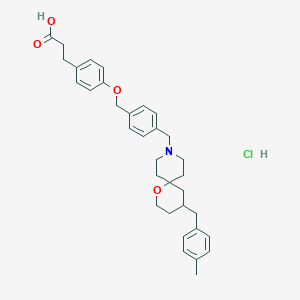 CHEMBL3926143 CHEMBL3926143 | C34H42ClNO4 | 564.163 | 5 / 2 | N/A | No |
| 503655 | 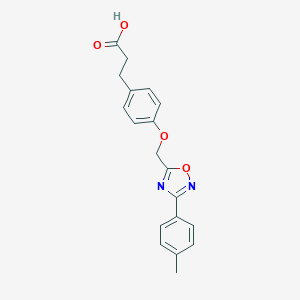 CHEMBL3601001 CHEMBL3601001 | C19H18N2O4 | 338.363 | 6 / 1 | 3.4 | Yes |
| 530813 | 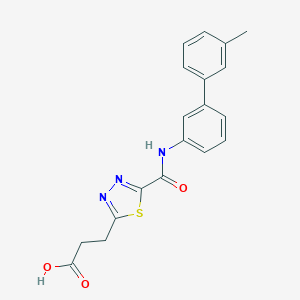 CHEMBL3805328 CHEMBL3805328 | C19H17N3O3S | 367.423 | 6 / 2 | 3.3 | Yes |
| 555241 | 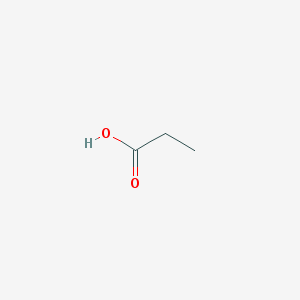 propionic acid propionic acid | C3H6O2 | 74.079 | 2 / 1 | 0.3 | Yes |
| 552794 | 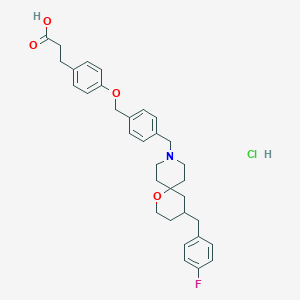 CHEMBL3941821 CHEMBL3941821 | C33H39ClFNO4 | 568.126 | 6 / 2 | N/A | No |
| 533007 | 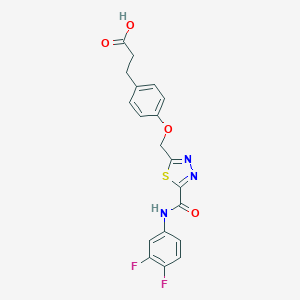 CHEMBL3805952 CHEMBL3805952 | C19H15F2N3O4S | 419.403 | 9 / 2 | 3.0 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218