

You can:
| Name | G-protein coupled receptor 182 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | GPR182 |
| Synonym | NOW L1-R GPR182 Gpcr22 Gpcr17 [ Show all ] |
| Disease | N/A |
| Length | 404 |
| Amino acid sequence | MSVKPSWGPGPSEGVTAVPTSDLGEIHNWTELLDLFNHTLSECHVELSQSTKRVVLFALYLAMFVVGLVENLLVICVNWRGSGRAGLMNLYILNMAIADLGIVLSLPVWMLEVTLDYTWLWGSFSCRFTHYFYFVNMYSSIFFLVCLSVDRYVTLTSASPSWQRYQHRVRRAMCAGIWVLSAIIPLPEVVHIQLVEGPEPMCLFMAPFETYSTWALAVALSTTILGFLLPFPLITVFNVLTACRLRQPGQPKSRRHCLLLCAYVAVFVMCWLPYHVTLLLLTLHGTHISLHCHLVHLLYFFYDVIDCFSMLHCVINPILYNFLSPHFRGRLLNAVVHYLPKDQTKAGTCASSSSCSTQHSIIITKGDSQPAAAAPHPEPSLSFQAHHLLPNTSPISPTQPLTPS |
| UniProt | O15218 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | O15218 |
| 3D structure model | This predicted structure model is from GPCR-EXP O15218. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL4356 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 53257 | 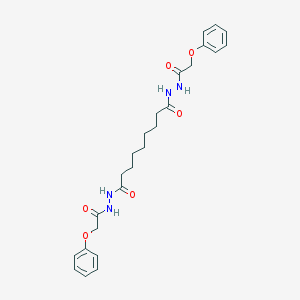 AC1NC8RF AC1NC8RF | C25H32N4O6 | 484.553 | 6 / 4 | 3.5 | Yes |
| 106229 | 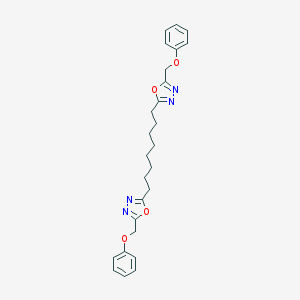 NSC697154 NSC697154 | C26H30N4O4 | 462.55 | 8 / 0 | 5.7 | No |
| 113148 | 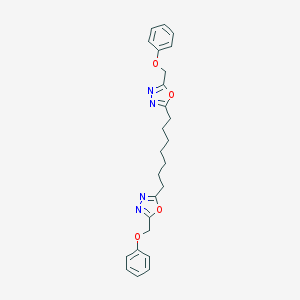 CHEMBL195442 CHEMBL195442 | C25H28N4O4 | 448.523 | 8 / 0 | 5.2 | No |
| 141835 | 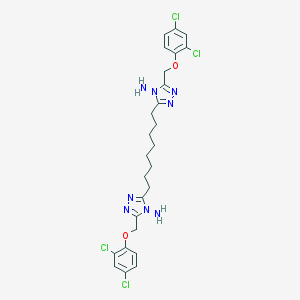 NSC697164 NSC697164 | C26H30Cl4N8O2 | 628.38 | 8 / 2 | 7.1 | No |
| 148880 | 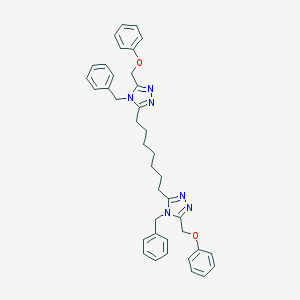 CHEMBL194933 CHEMBL194933 | C39H42N6O2 | 626.805 | 6 / 0 | 7.7 | No |
| 158080 | 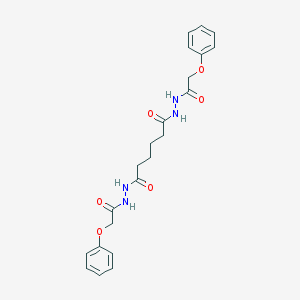 MLS001219965 MLS001219965 | C22H26N4O6 | 442.472 | 6 / 4 | 1.9 | Yes |
| 176630 | 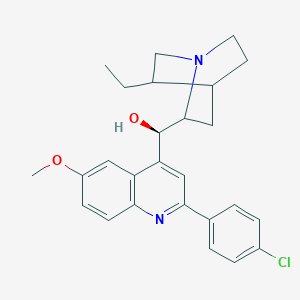 CHEMBL419684 CHEMBL419684 | C26H29ClN2O2 | 436.98 | 4 / 1 | 5.4 | No |
| 184293 | 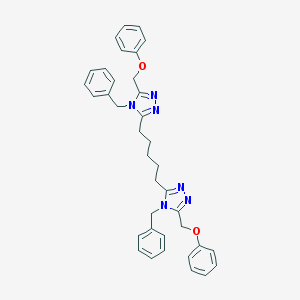 CHEMBL372706 CHEMBL372706 | C37H38N6O2 | 598.751 | 6 / 0 | 6.7 | No |
| 191514 | 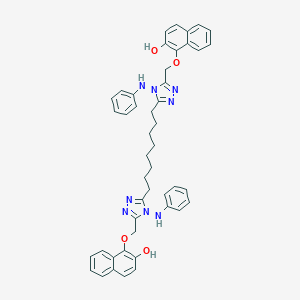 NSC697168 NSC697168 | C46H46N8O4 | 774.926 | 10 / 4 | 10.8 | No |
| 218715 | 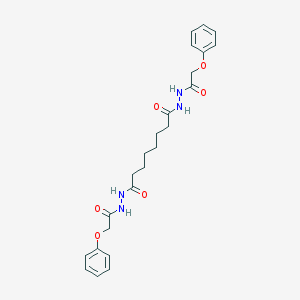 CHEMBL195275 CHEMBL195275 | C24H30N4O6 | 470.526 | 6 / 4 | 2.9 | Yes |
| 218723 | 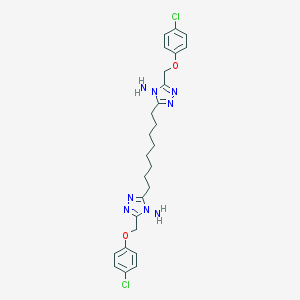 NSC697161 NSC697161 | C26H32Cl2N8O2 | 559.496 | 8 / 2 | 5.9 | No |
| 232766 | 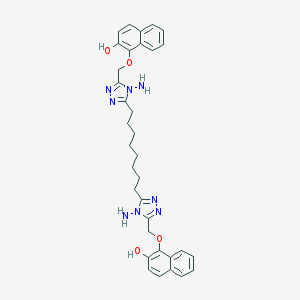 NSC697163 NSC697163 | C34H38N8O4 | 622.73 | 10 / 4 | 6.4 | No |
| 236963 | 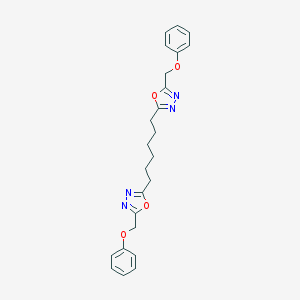 CHEMBL195133 CHEMBL195133 | C24H26N4O4 | 434.496 | 8 / 0 | 4.7 | Yes |
| 241929 | 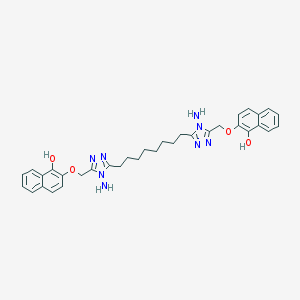 NSC697162 NSC697162 | C34H38N8O4 | 622.73 | 10 / 4 | 6.4 | No |
| 256541 | 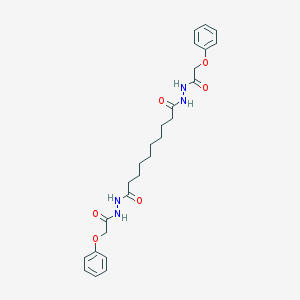 CHEMBL192683 CHEMBL192683 | C26H34N4O6 | 498.58 | 6 / 4 | 4.0 | Yes |
| 266068 | 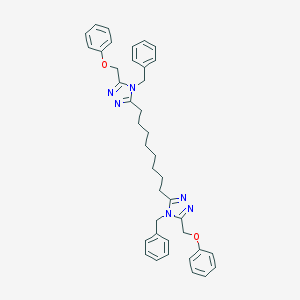 CHEMBL175928 CHEMBL175928 | C40H44N6O2 | 640.832 | 6 / 0 | 8.3 | No |
| 296678 | 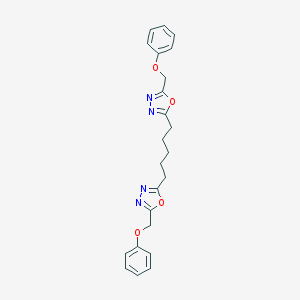 CHEMBL192620 CHEMBL192620 | C23H24N4O4 | 420.469 | 8 / 0 | 4.1 | Yes |
| 302608 | 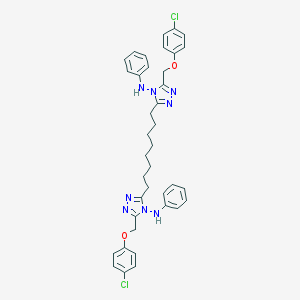 NSC697166 NSC697166 | C38H40Cl2N8O2 | 711.692 | 8 / 2 | 10.3 | No |
| 316389 | 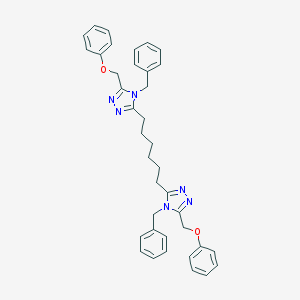 CHEMBL194874 CHEMBL194874 | C38H40N6O2 | 612.778 | 6 / 0 | 7.2 | No |
| 346622 | 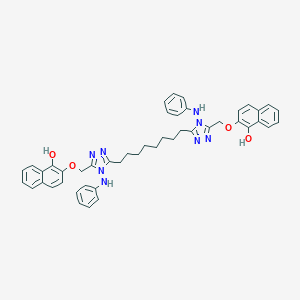 NSC697167 NSC697167 | C46H46N8O4 | 774.926 | 10 / 4 | 10.8 | No |
| 351719 | 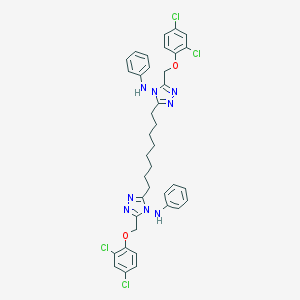 NSC697169 NSC697169 | C38H38Cl4N8O2 | 780.576 | 8 / 2 | 11.6 | No |
| 369056 | 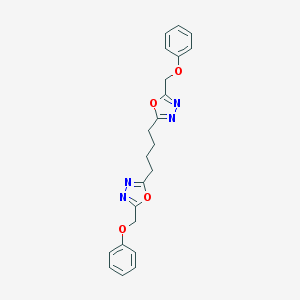 CHEMBL192644 CHEMBL192644 | C22H22N4O4 | 406.442 | 8 / 0 | 3.6 | Yes |
| 382561 | 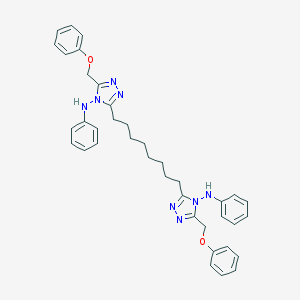 NSC697165 NSC697165 | C38H42N8O2 | 642.808 | 8 / 2 | 9.1 | No |
| 391680 | 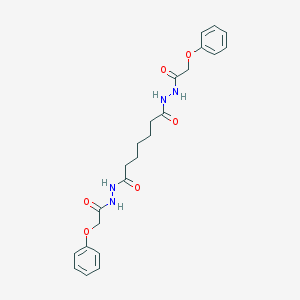 CHEMBL371823 CHEMBL371823 | C23H28N4O6 | 456.499 | 6 / 4 | 2.4 | Yes |
| 418047 | 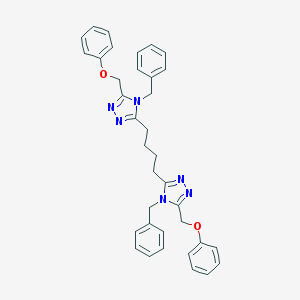 CHEMBL361867 CHEMBL361867 | C36H36N6O2 | 584.724 | 6 / 0 | 6.1 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218