

You can:
| Name | Lutropin-choriogonadotropic hormone receptor |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Lhcgr |
| Synonym | Luteinizing hormone receptor LSH-R LHR LH/CG-R LH-R [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 700 |
| Amino acid sequence | MGRRVPALRQLLVLAVLLLKPSQLQSRELSGSRCPEPCDCAPDGALRCPGPRAGLARLSLTYLPVKVIPSQAFRGLNEVVKIEISQSDSLERIEANAFDNLLNLSELLIQNTKNLLYIEPGAFTNLPRLKYLSICNTGIRTLPDVTKISSSEFNFILEICDNLHITTIPGNAFQGMNNESVTLKLYGNGFEEVQSHAFNGTTLISLELKENIYLEKMHSGAFQGATGPSILDISSTKLQALPSHGLESIQTLIALSSYSLKTLPSKEKFTSLLVATLTYPSHCCAFRNLPKKEQNFSFSIFENFSKQCESTVRKADNETLYSAIFEENELSGWDYDYGFCSPKTLQCAPEPDAFNPCEDIMGYAFLRVLIWLINILAIFGNLTVLFVLLTSRYKLTVPRFLMCNLSFADFCMGLYLLLIASVDSQTKGQYYNHAIDWQTGSGCGAAGFFTVFASELSVYTLTVITLERWHTITYAVQLDQKLRLRHAIPIMLGGWLFSTLIATMPLVGISNYMKVSICLPMDVESTLSQVYILSILILNVVAFVVICACYIRIYFAVQNPELTAPNKDTKIAKKMAILIFTDFTCMAPISFFAISAAFKVPLITVTNSKILLVLFYPVNSCANPFLYAIFTKAFQRDFLLLLSRFGCCKRRAELYRRKEFSAYTSNCKNGFPGASKPSQATLKLSTVHCQQPIPPRALTH |
| UniProt | P16235 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL2456 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 8869 | 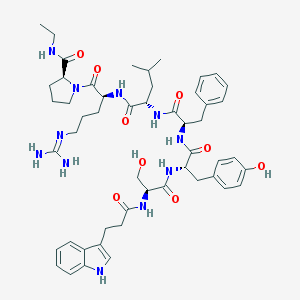 CHEMBL2369806 CHEMBL2369806 | C51H69N11O9 | 980.181 | 10 / 11 | 2.6 | No |
| 11209 | 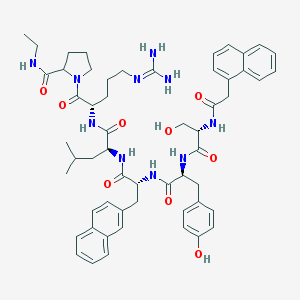 CHEMBL275709 CHEMBL275709 | C56H70N10O9 | 1027.24 | 10 / 10 | 4.7 | No |
| 12406 | 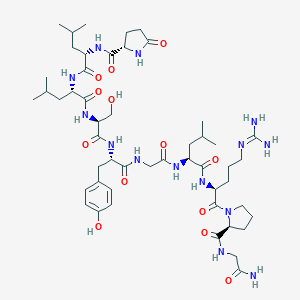 CHEMBL3272555 CHEMBL3272555 | C50H80N14O13 | 1085.28 | 14 / 14 | -1.2 | No |
| 13411 | 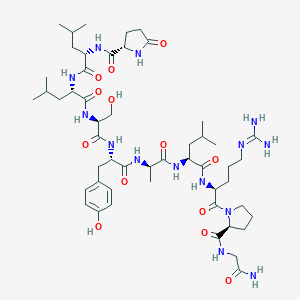 CHEMBL3272556 CHEMBL3272556 | C51H82N14O13 | 1099.3 | 14 / 14 | -0.8 | No |
| 29948 | 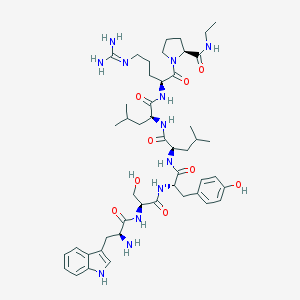 CHEMBL2369794 CHEMBL2369794 | C48H72N12O9 | 961.179 | 11 / 12 | 1.2 | No |
| 40074 | 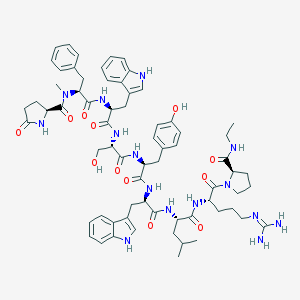 CHEMBL429240 CHEMBL429240 | C68H87N15O12 | 1306.54 | 13 / 14 | 3.1 | No |
| 45382 | 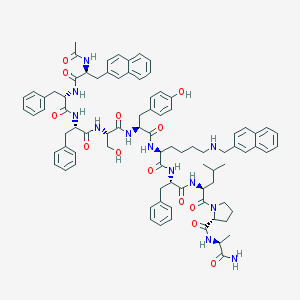 CHEMBL413661 CHEMBL413661 | C85H100N12O13 | 1497.81 | 14 / 13 | 9.0 | No |
| 45486 | 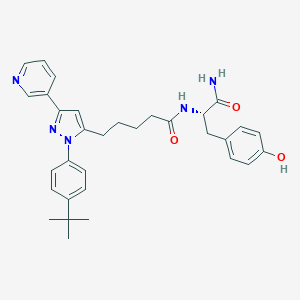 CHEMBL246321 CHEMBL246321 | C32H37N5O3 | 539.68 | 5 / 3 | 5.0 | No |
| 45538 | 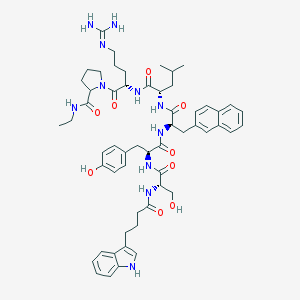 CHEMBL413887 CHEMBL413887 | C56H73N11O9 | 1044.27 | 10 / 11 | 4.2 | No |
| 47738 | 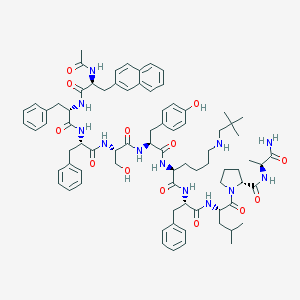 CHEMBL436705 CHEMBL436705 | C79H102N12O13 | 1427.76 | 14 / 13 | 8.0 | No |
| 52533 | 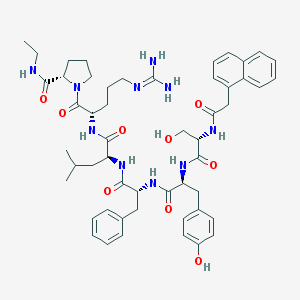 CHEMBL2369800 CHEMBL2369800 | C52H68N10O9 | 977.177 | 10 / 10 | 3.4 | No |
| 63100 | 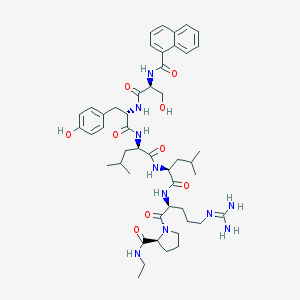 CHEMBL2369781 CHEMBL2369781 | C48H68N10O9 | 929.133 | 10 / 10 | 3.2 | No |
| 65556 | 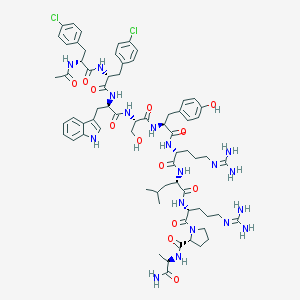 CHEMBL412984 CHEMBL412984 | C69H92Cl2N18O13 | 1452.51 | 15 / 17 | 2.2 | No |
| 66569 | 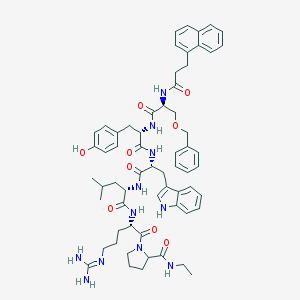 CHEMBL386525 CHEMBL386525 | C62H77N11O9 | 1120.37 | 10 / 10 | 5.9 | No |
| 72144 | 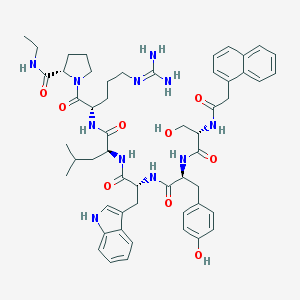 CHEMBL2369807 CHEMBL2369807 | C54H69N11O9 | 1016.21 | 10 / 11 | 3.6 | No |
| 76100 | 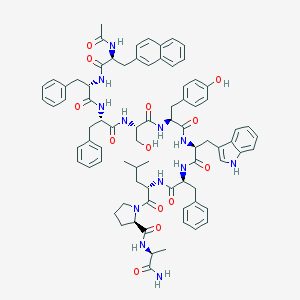 CHEMBL217123 CHEMBL217123 | C79H90N12O13 | 1415.66 | 13 / 13 | 7.7 | No |
| 77431 | 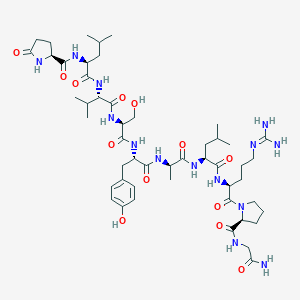 CHEMBL3272558 CHEMBL3272558 | C50H80N14O13 | 1085.28 | 14 / 14 | -1.2 | No |
| 79485 | 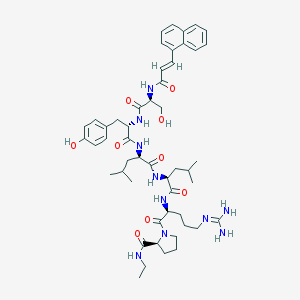 CHEMBL2369810 CHEMBL2369810 | C50H70N10O9 | 955.171 | 10 / 10 | 3.7 | No |
| 80453 | 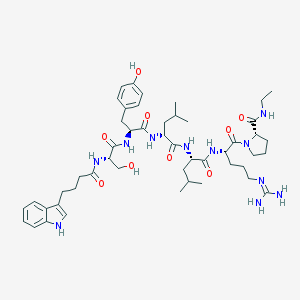 CHEMBL2369780 CHEMBL2369780 | C49H73N11O9 | 960.191 | 10 / 11 | 2.7 | No |
| 89345 | 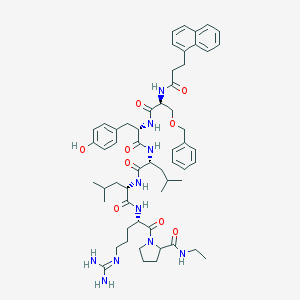 CHEMBL414382 CHEMBL414382 | C57H78N10O9 | 1047.31 | 10 / 9 | 5.5 | No |
| 91245 | 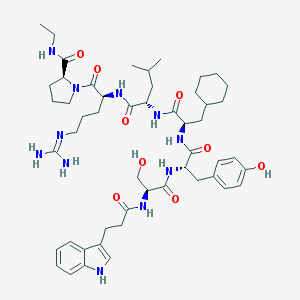 CHEMBL2369808 CHEMBL2369808 | C51H75N11O9 | 986.229 | 10 / 11 | 3.7 | No |
| 95098 | 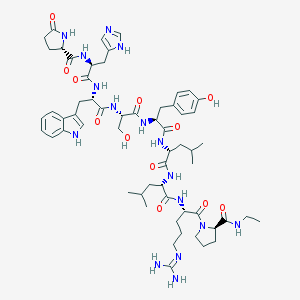 CHEMBL385468 CHEMBL385468 | C59H84N16O12 | 1209.42 | 14 / 15 | 0.7 | No |
| 98619 | 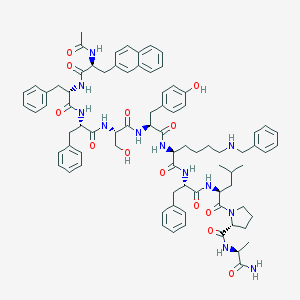 CHEMBL414329 CHEMBL414329 | C81H98N12O13 | 1447.75 | 14 / 13 | 7.8 | No |
| 105186 | 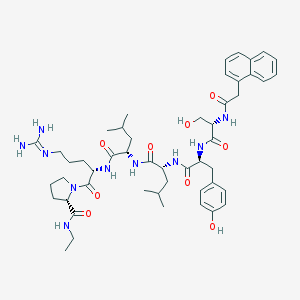 CHEMBL2369789 CHEMBL2369789 | C49H70N10O9 | 943.16 | 10 / 10 | 3.2 | No |
| 106677 | 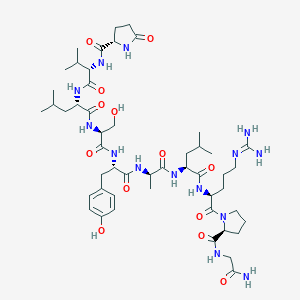 CHEMBL3272557 CHEMBL3272557 | C50H80N14O13 | 1085.28 | 14 / 14 | -1.2 | No |
| 107607 | 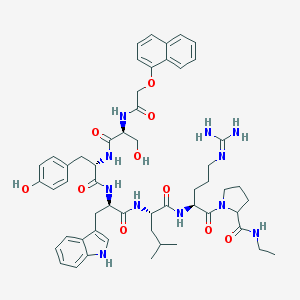 CHEMBL412573 CHEMBL412573 | C54H69N11O10 | 1032.21 | 11 / 11 | 3.7 | No |
| 113748 | 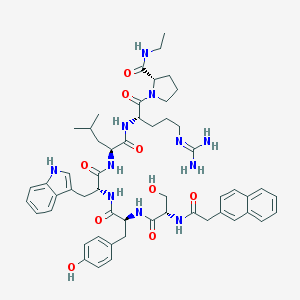 CHEMBL2369798 CHEMBL2369798 | C54H69N11O9 | 1016.21 | 10 / 11 | 3.6 | No |
| 126164 | 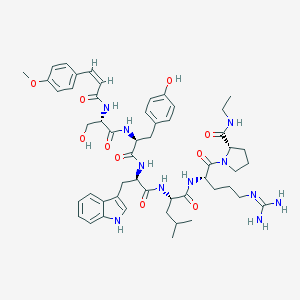 CHEMBL2369787 CHEMBL2369787 | C52H69N11O10 | 1008.19 | 11 / 11 | 2.8 | No |
| 126165 | 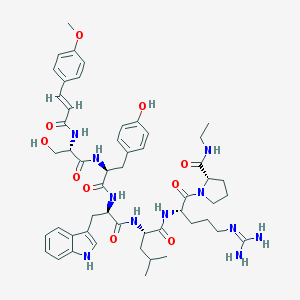 CHEMBL2369793 CHEMBL2369793 | C52H69N11O10 | 1008.19 | 11 / 11 | 2.8 | No |
| 127016 | 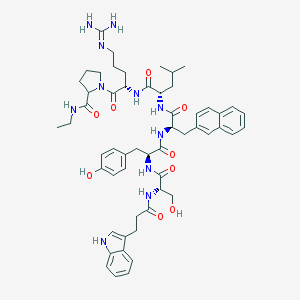 CHEMBL429543 CHEMBL429543 | C55H71N11O9 | 1030.24 | 10 / 11 | 3.9 | No |
| 127159 | 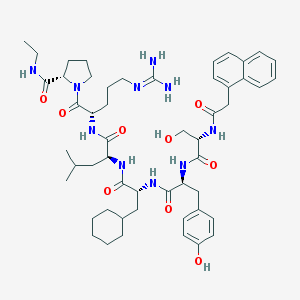 CHEMBL2369799 CHEMBL2369799 | C52H74N10O9 | 983.225 | 10 / 10 | 4.6 | No |
| 132353 | 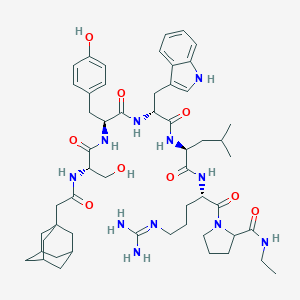 CHEMBL265413 CHEMBL265413 | C54H77N11O9 | 1024.28 | 10 / 11 | 3.8 | No |
| 139342 | 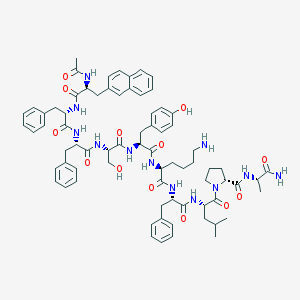 CHEMBL414753 CHEMBL414753 | C74H92N12O13 | 1357.62 | 14 / 13 | 5.8 | No |
| 148457 | 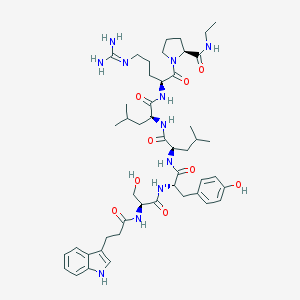 CHEMBL2369784 CHEMBL2369784 | C48H71N11O9 | 946.164 | 10 / 11 | 2.0 | No |
| 151130 | 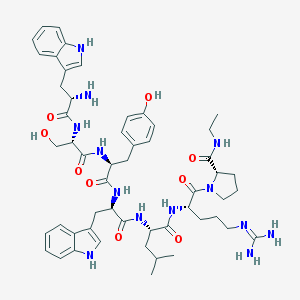 CHEMBL2369814 CHEMBL2369814 | C53H71N13O9 | 1034.23 | 11 / 13 | 1.9 | No |
| 151774 | 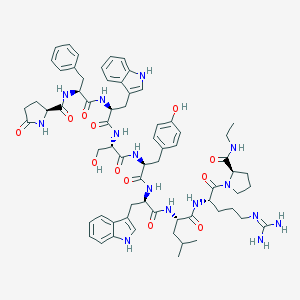 CHEMBL407606 CHEMBL407606 | C67H85N15O12 | 1292.51 | 13 / 15 | 3.0 | No |
| 154841 | 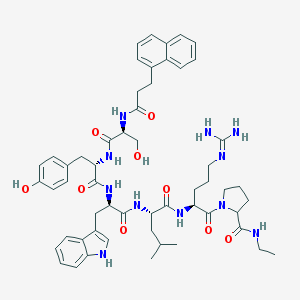 CHEMBL275694 CHEMBL275694 | C55H71N11O9 | 1030.24 | 10 / 11 | 3.9 | No |
| 161752 | 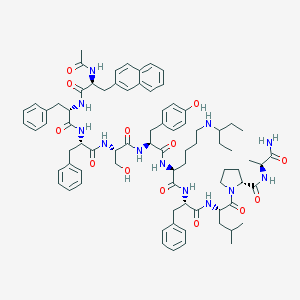 CHEMBL404944 CHEMBL404944 | C79H102N12O13 | 1427.76 | 14 / 13 | 8.2 | No |
| 169270 | 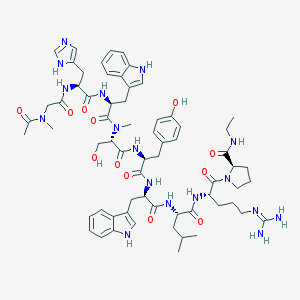 CHEMBL437798 CHEMBL437798 | C65H87N17O12 | 1298.52 | 14 / 14 | 1.4 | No |
| 171105 | 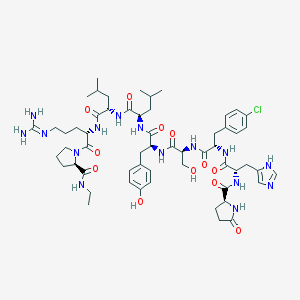 CHEMBL414071 CHEMBL414071 | C57H82ClN15O12 | 1204.83 | 14 / 14 | 0.6 | No |
| 173431 | 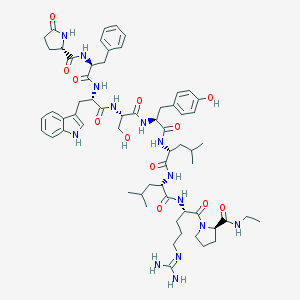 CHEMBL217405 CHEMBL217405 | C62H86N14O12 | 1219.46 | 13 / 14 | 2.6 | No |
| 178511 | 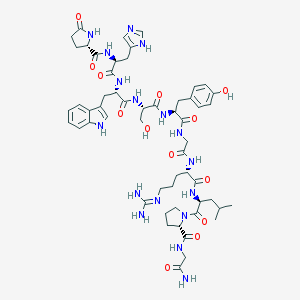 CHEMBL2371416 CHEMBL2371416 | C55H75N17O13 | 1182.31 | 15 / 16 | -2.4 | No |
| 181313 | 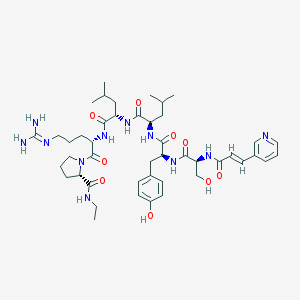 CHEMBL2369809 CHEMBL2369809 | C45H67N11O9 | 906.099 | 11 / 10 | 1.3 | No |
| 187404 | 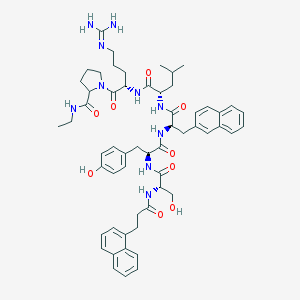 CHEMBL216617 CHEMBL216617 | C57H72N10O9 | 1041.26 | 10 / 10 | 5.0 | No |
| 199805 | 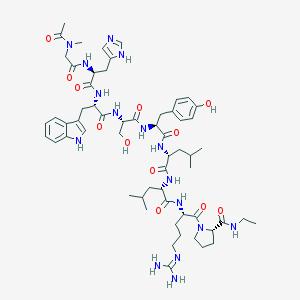 CHEMBL274682 CHEMBL274682 | C59H86N16O12 | 1211.44 | 14 / 14 | 0.8 | No |
| 200980 | 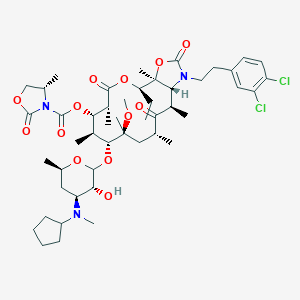 CHEMBL275935 CHEMBL275935 | C48H71Cl2N3O13 | 969.004 | 14 / 1 | 8.2 | No |
| 205302 | 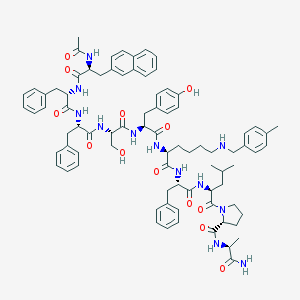 CHEMBL405264 CHEMBL405264 | C82H100N12O13 | 1461.77 | 14 / 13 | 8.2 | No |
| 225372 | 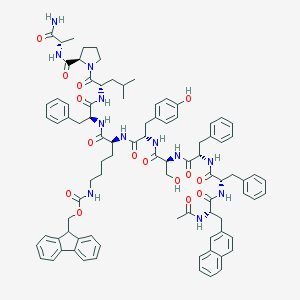 CHEMBL413879 CHEMBL413879 | C89H102N12O15 | 1579.86 | 15 / 13 | 9.6 | No |
| 227037 | 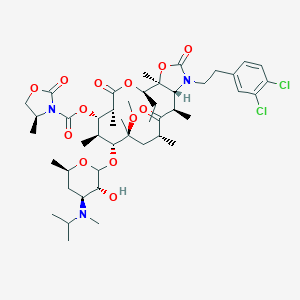 CHEMBL273440 CHEMBL273440 | C46H69Cl2N3O13 | 942.966 | 14 / 1 | 7.8 | No |
| 228806 | 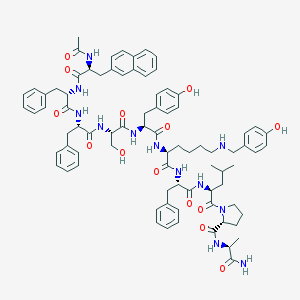 CHEMBL265231 CHEMBL265231 | C81H98N12O14 | 1463.74 | 15 / 14 | 7.4 | No |
| 243164 | 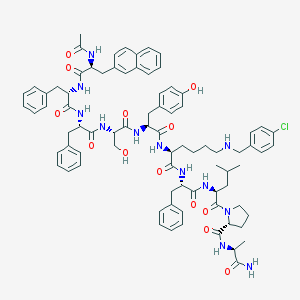 CHEMBL425278 CHEMBL425278 | C81H97ClN12O13 | 1482.19 | 14 / 13 | 8.4 | No |
| 244464 | 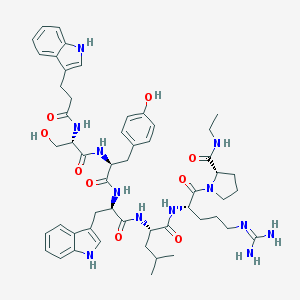 CHEMBL2369788 CHEMBL2369788 | C53H70N12O9 | 1019.22 | 10 / 12 | 2.7 | No |
| 251234 | 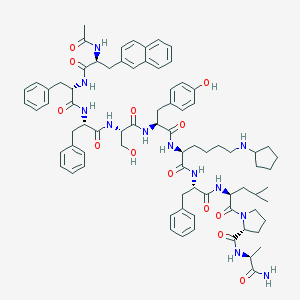 CHEMBL384049 CHEMBL384049 | C79H100N12O13 | 1425.74 | 14 / 13 | 7.6 | No |
| 251235 | 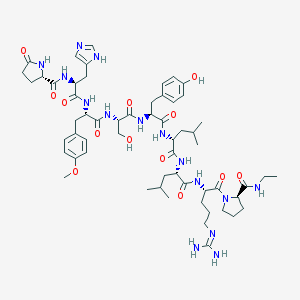 CHEMBL415571 CHEMBL415571 | C58H85N15O13 | 1200.41 | 15 / 14 | 0.0 | No |
| 495284 | 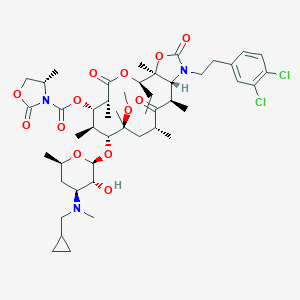 CHEMBL303274 CHEMBL303274 | C47H69Cl2N3O13 | 954.977 | 14 / 1 | 7.7 | No |
| 262830 | 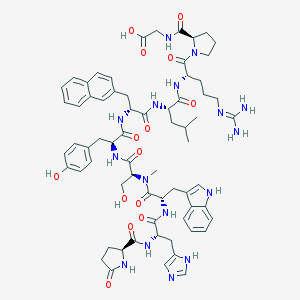 CHEMBL407123 CHEMBL407123 | C67H84N16O14 | 1337.51 | 16 / 15 | 1.7 | No |
| 263283 | 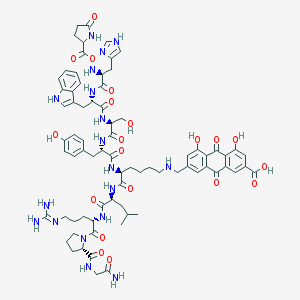 CID 44346805 CID 44346805 | C75H92N18O20 | 1565.67 | 24 / 20 | -1.9 | No |
| 265293 | 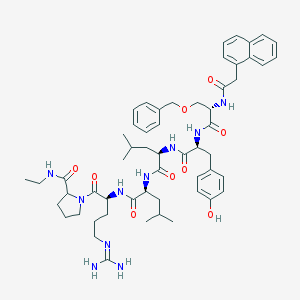 CHEMBL411175 CHEMBL411175 | C56H76N10O9 | 1033.29 | 10 / 9 | 5.2 | No |
| 275345 | 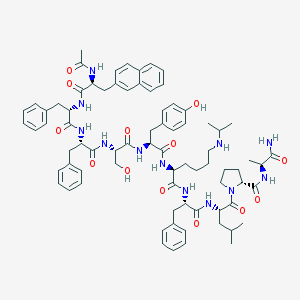 CHEMBL408991 CHEMBL408991 | C77H98N12O13 | 1399.7 | 14 / 13 | 7.1 | No |
| 287357 | 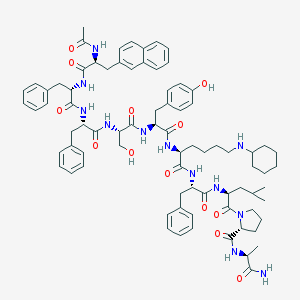 CHEMBL262162 CHEMBL262162 | C80H102N12O13 | 1439.77 | 14 / 13 | 8.1 | No |
| 313119 | 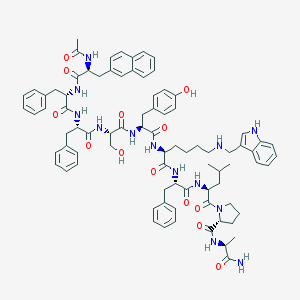 CHEMBL268106 CHEMBL268106 | C83H99N13O13 | 1486.78 | 14 / 14 | 7.9 | No |
| 316831 | 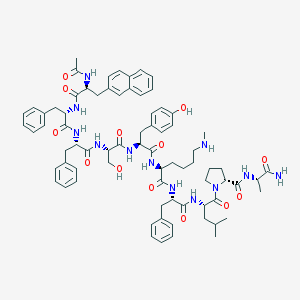 CHEMBL263167 CHEMBL263167 | C75H94N12O13 | 1371.65 | 14 / 13 | 6.3 | No |
| 328168 | 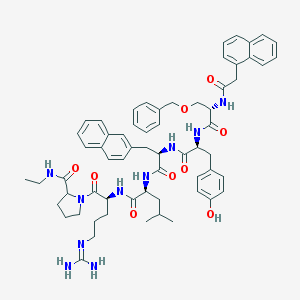 CHEMBL274609 CHEMBL274609 | C63H76N10O9 | 1117.36 | 10 / 9 | 6.7 | No |
| 330977 | 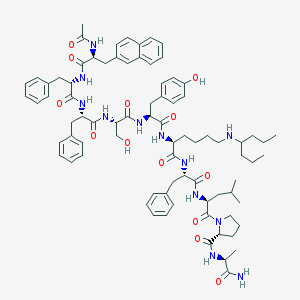 CHEMBL439880 CHEMBL439880 | C81H106N12O13 | 1455.81 | 14 / 13 | 8.9 | No |
| 338326 | 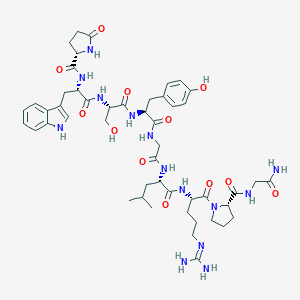 CHEMBL3272560 CHEMBL3272560 | C49H68N14O12 | 1045.17 | 13 / 14 | -2.3 | No |
| 344260 | 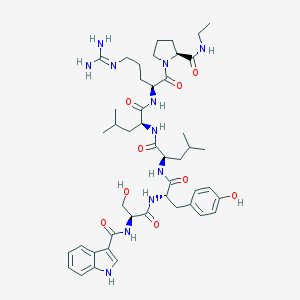 CHEMBL2369791 CHEMBL2369791 | C46H67N11O9 | 918.11 | 10 / 11 | 2.1 | No |
| 347116 | 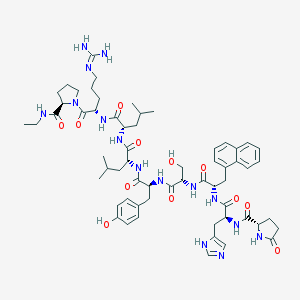 CHEMBL385042 CHEMBL385042 | C61H85N15O12 | 1220.44 | 14 / 14 | 1.9 | No |
| 358110 | 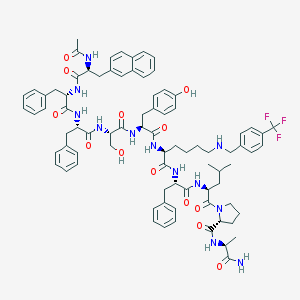 CHEMBL408640 CHEMBL408640 | C82H97F3N12O13 | 1515.74 | 17 / 13 | 8.7 | No |
| 368087 | 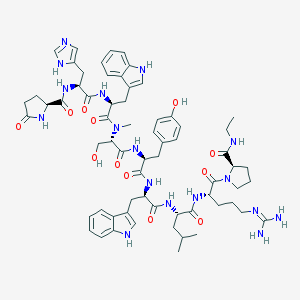 CHEMBL405737 CHEMBL405737 | C65H85N17O12 | 1296.5 | 14 / 15 | 1.3 | No |
| 368801 | 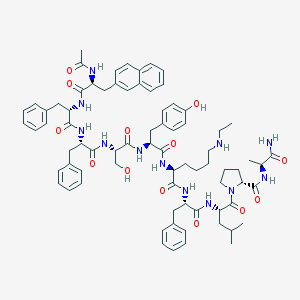 CHEMBL406607 CHEMBL406607 | C76H96N12O13 | 1385.68 | 14 / 13 | 6.7 | No |
| 388204 | 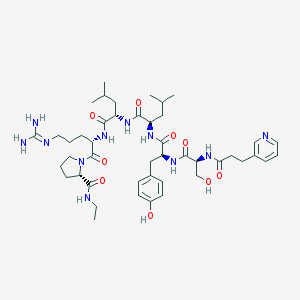 CHEMBL2369783 CHEMBL2369783 | C45H69N11O9 | 908.115 | 11 / 10 | 1.1 | No |
| 389540 | 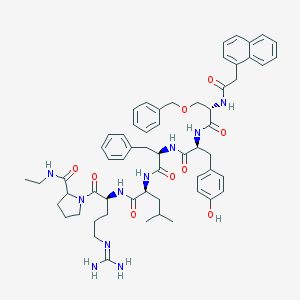 CHEMBL415109 CHEMBL415109 | C59H74N10O9 | 1067.3 | 10 / 9 | 5.5 | No |
| 394586 | 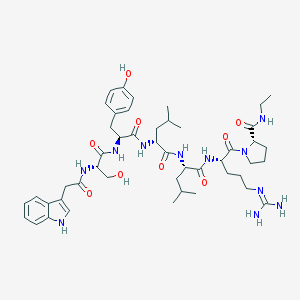 CHEMBL2369790 CHEMBL2369790 | C47H69N11O9 | 932.137 | 10 / 11 | 2.0 | No |
| 395307 | 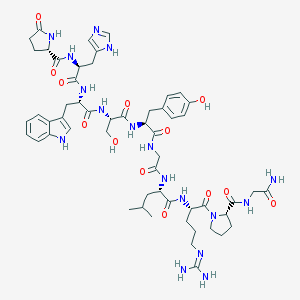 GONADORELIN GONADORELIN | C55H75N17O13 | 1182.31 | 15 / 16 | -2.4 | No |
| 397847 | 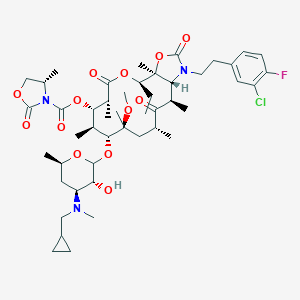 CHEMBL12282 CHEMBL12282 | C47H69ClFN3O13 | 938.525 | 15 / 1 | 7.2 | No |
| 398616 | 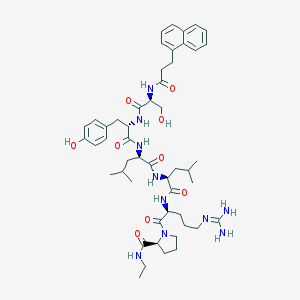 CHEMBL2369813 CHEMBL2369813 | C50H72N10O9 | 957.187 | 10 / 10 | 3.5 | No |
| 419295 | 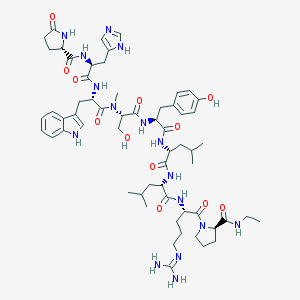 CHEMBL266205 CHEMBL266205 | C60H86N16O12 | 1223.45 | 14 / 14 | 0.9 | No |
| 423933 | 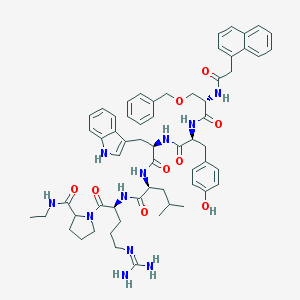 CHEMBL264298 CHEMBL264298 | C61H75N11O9 | 1106.34 | 10 / 10 | 5.6 | No |
| 435840 | 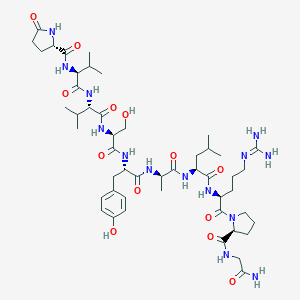 CHEMBL3272559 CHEMBL3272559 | C49H78N14O13 | 1071.25 | 14 / 14 | -1.6 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218