

You can:
| Name | Alpha-2B adrenergic receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | ADRA2B |
| Synonym | alpha2-C2 alpha2B alpha2B-adrenoceptor Alpha-2BAR alpha-2B adrenoreceptor [ Show all ] |
| Disease | Neuropathic pain Alcohol use disorders |
| Length | 450 |
| Amino acid sequence | MDHQDPYSVQATAAIAAAITFLILFTIFGNALVILAVLTSRSLRAPQNLFLVSLAAADILVATLIIPFSLANELLGYWYFRRTWCEVYLALDVLFCTSSIVHLCAISLDRYWAVSRALEYNSKRTPRRIKCIILTVWLIAAVISLPPLIYKGDQGPQPRGRPQCKLNQEAWYILASSIGSFFAPCLIMILVYLRIYLIAKRSNRRGPRAKGGPGQGESKQPRPDHGGALASAKLPALASVASAREVNGHSKSTGEKEEGETPEDTGTRALPPSWAALPNSGQGQKEGVCGASPEDEAEEEEEEEEEEEECEPQAVPVSPASACSPPLQQPQGSRVLATLRGQVLLGRGVGAIGGQWWRRRAQLTREKRFTFVLAVVIGVFVLCWFPFFFSYSLGAICPKHCKVPHGLFQFFFWIGYCNSSLNPVIYTIFNQDFRRAFRRILCRPWTQTAW |
| UniProt | P18089 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P18089 |
| 3D structure model | This predicted structure model is from GPCR-EXP P18089. |
| BioLiP | N/A |
| Therapeutic Target Database | T41580 |
| ChEMBL | CHEMBL1942 |
| IUPHAR | 26 |
| DrugBank | BE0000572 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 369 | 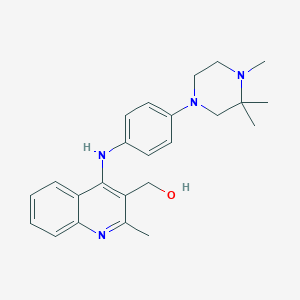 CHEMBL218675 CHEMBL218675 | C24H30N4O | 390.531 | 5 / 2 | 4.3 | Yes |
| 1494 | 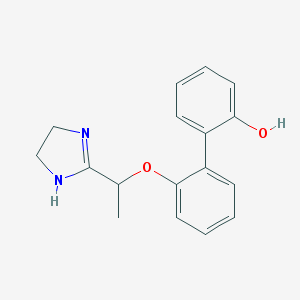 CHEMBL373830 CHEMBL373830 | C17H18N2O2 | 282.343 | 3 / 2 | 2.6 | Yes |
| 2271 | 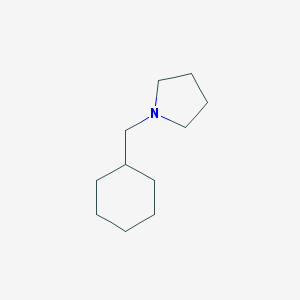 1-(cyclohexylmethyl)pyrrolidine 1-(cyclohexylmethyl)pyrrolidine | C11H21N | 167.296 | 1 / 0 | 3.1 | Yes |
| 3156 | 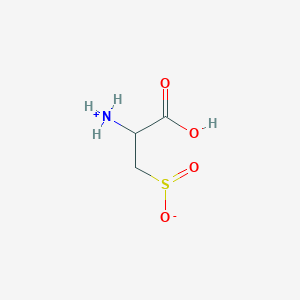 AC1N0YGA AC1N0YGA | C3H7NO4S | 153.152 | 5 / 2 | -4.7 | Yes |
| 4322 | 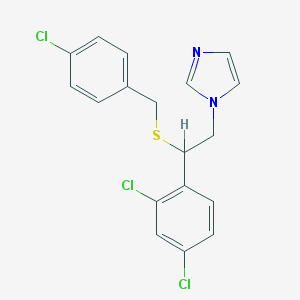 sulconazole sulconazole | C18H15Cl3N2S | 397.742 | 2 / 0 | 6.1 | No |
| 4416 | 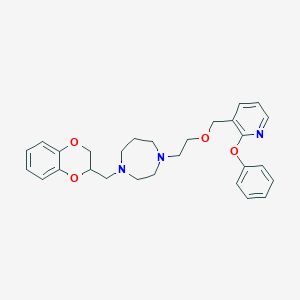 CHEMBL491419 CHEMBL491419 | C28H33N3O4 | 475.589 | 7 / 0 | 4.0 | Yes |
| 4569 | 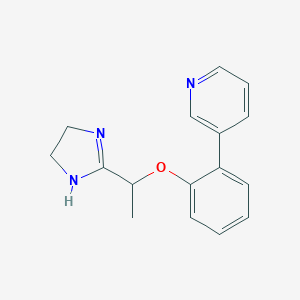 CHEMBL373535 CHEMBL373535 | C16H17N3O | 267.332 | 3 / 1 | 1.9 | Yes |
| 4667 | 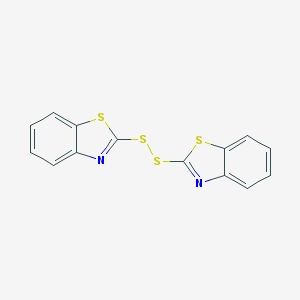 120-78-5 120-78-5 | C14H8N2S4 | 332.472 | 6 / 0 | 5.6 | No |
| 5064 | 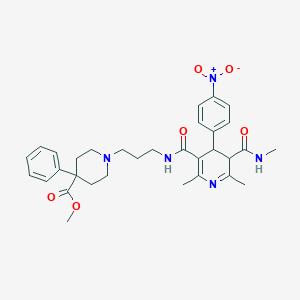 BDBM50068818 BDBM50068818 | C32H39N5O6 | 589.693 | 8 / 2 | 2.9 | No |
| 5125 | 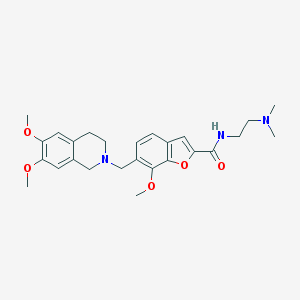 CHEMBL248272 CHEMBL248272 | C26H33N3O5 | 467.566 | 7 / 1 | 3.3 | Yes |
| 5668 | 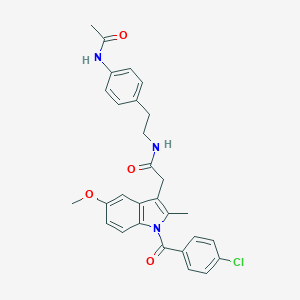 AC1L1HUK AC1L1HUK | C29H28ClN3O4 | 518.01 | 4 / 2 | 5.2 | No |
| 463707 | 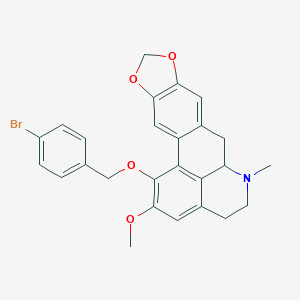 CHEMBL3633757 CHEMBL3633757 | C26H24BrNO4 | 494.385 | 5 / 0 | 5.4 | No |
| 7348 | 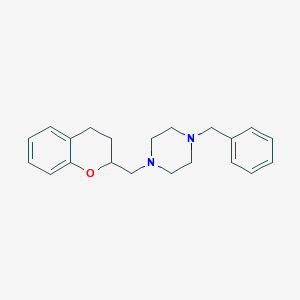 CHEMBL179648 CHEMBL179648 | C21H26N2O | 322.452 | 3 / 0 | 3.7 | Yes |
| 7405 | 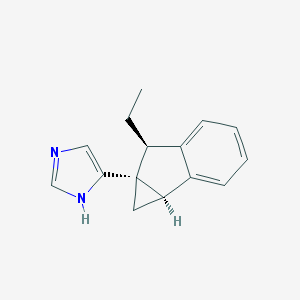 CHEMBL1256610 CHEMBL1256610 | C15H16N2 | 224.307 | 1 / 1 | 2.9 | Yes |
| 8053 | 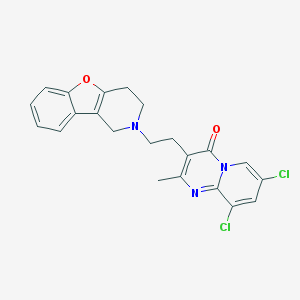 CHEMBL163247 CHEMBL163247 | C22H19Cl2N3O2 | 428.313 | 4 / 0 | 4.0 | Yes |
| 521681 | 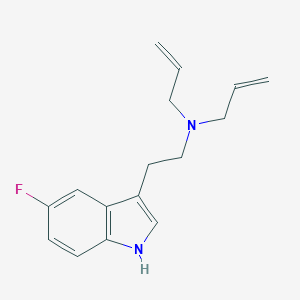 CHEMBL3754496 CHEMBL3754496 | C16H19FN2 | 258.34 | 2 / 1 | 2.9 | Yes |
| 9860 | 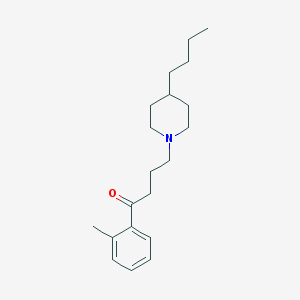 AC-42 AC-42 | C20H31NO | 301.474 | 2 / 0 | 5.2 | No |
| 10522 | 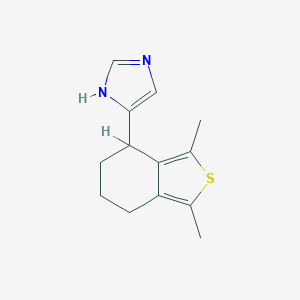 CHEMBL30739 CHEMBL30739 | C13H16N2S | 232.345 | 2 / 1 | 3.2 | Yes |
| 10723 | 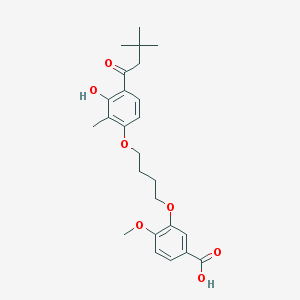 CHEMBL3287723 CHEMBL3287723 | C25H32O7 | 444.524 | 7 / 2 | 5.7 | No |
| 11151 | 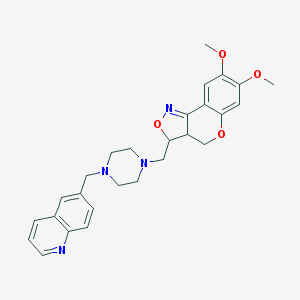 CHEMBL90966 CHEMBL90966 | C27H30N4O4 | 474.561 | 8 / 0 | 3.2 | Yes |
| 11587 | 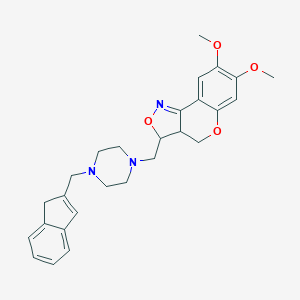 CHEMBL328377 CHEMBL328377 | C27H31N3O4 | 461.562 | 7 / 0 | 3.3 | Yes |
| 11718 | 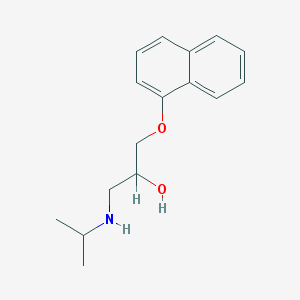 propranolol propranolol | C16H21NO2 | 259.349 | 3 / 2 | 3.0 | Yes |
| 12454 | 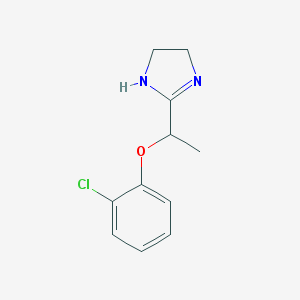 CHEMBL1956191 CHEMBL1956191 | C11H13ClN2O | 224.688 | 2 / 1 | 2.0 | Yes |
| 12492 | 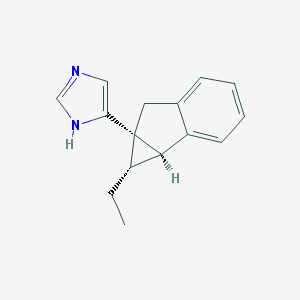 CHEMBL1255723 CHEMBL1255723 | C15H16N2 | 224.307 | 1 / 1 | 3.1 | Yes |
| 13155 | 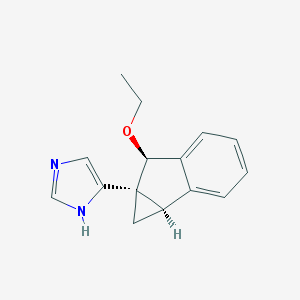 CHEMBL1256565 CHEMBL1256565 | C15H16N2O | 240.306 | 2 / 1 | 1.9 | Yes |
| 13847 | 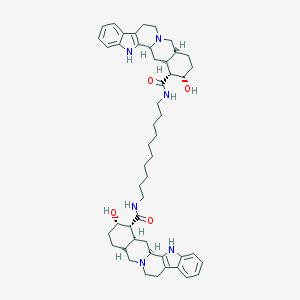 CHEMBL554019 CHEMBL554019 | C50H68N6O4 | 817.132 | 6 / 6 | 8.0 | No |
| 15089 | 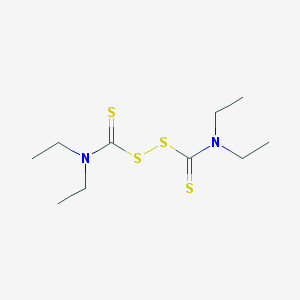 disulfiram disulfiram | C10H20N2S4 | 296.524 | 4 / 0 | 3.9 | Yes |
| 15680 | 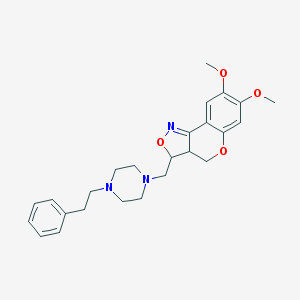 CHEMBL92496 CHEMBL92496 | C25H31N3O4 | 437.54 | 7 / 0 | 3.4 | Yes |
| 16311 | 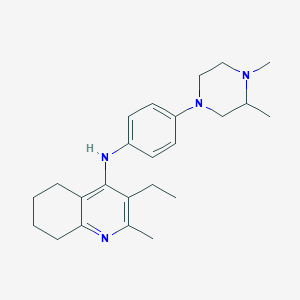 CHEMBL217768 CHEMBL217768 | C24H34N4 | 378.564 | 4 / 1 | 5.2 | No |
| 16771 | 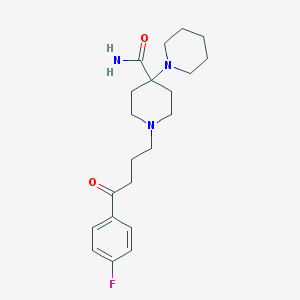 Pipamperone Pipamperone | C21H30FN3O2 | 375.488 | 5 / 1 | 2.0 | Yes |
| 18665 | 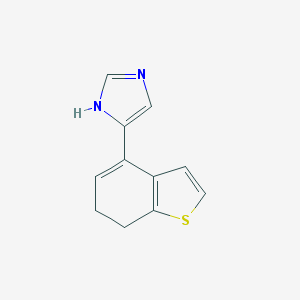 CHEMBL286246 CHEMBL286246 | C11H10N2S | 202.275 | 2 / 1 | 2.1 | Yes |
| 18690 | 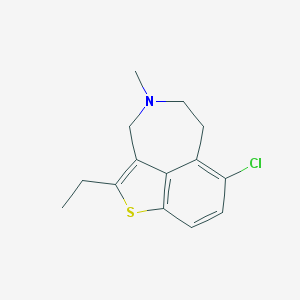 SK&F-106686 SK&F-106686 | C14H16ClNS | 265.799 | 2 / 0 | 4.3 | Yes |
| 20054 | 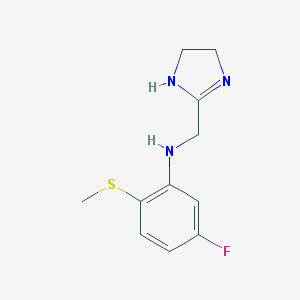 CHEMBL72768 CHEMBL72768 | C11H14FN3S | 239.312 | 4 / 2 | 1.5 | Yes |
| 21116 | 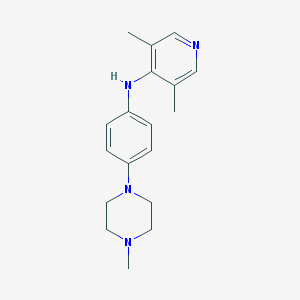 CHEMBL217180 CHEMBL217180 | C18H24N4 | 296.418 | 4 / 1 | 3.1 | Yes |
| 21890 | 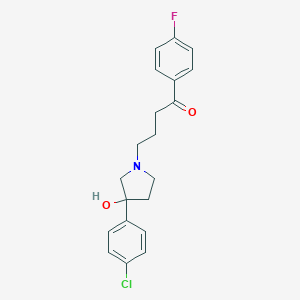 CHEMBL150161 CHEMBL150161 | C20H21ClFNO2 | 361.841 | 4 / 1 | 2.9 | Yes |
| 22389 | 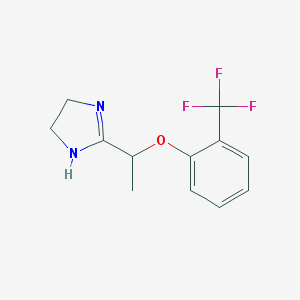 CHEMBL1956193 CHEMBL1956193 | C12H13F3N2O | 258.244 | 5 / 1 | 2.2 | Yes |
| 22606 | 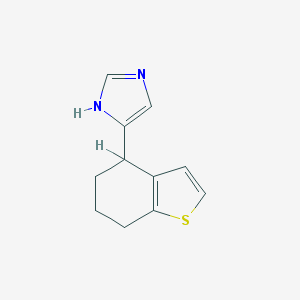 CHEMBL284213 CHEMBL284213 | C11H12N2S | 204.291 | 2 / 1 | 2.3 | Yes |
| 23272 | 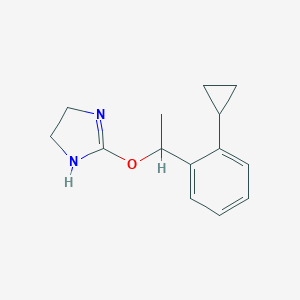 CHEMBL576623 CHEMBL576623 | C14H18N2O | 230.311 | 2 / 1 | 2.4 | Yes |
| 24245 | 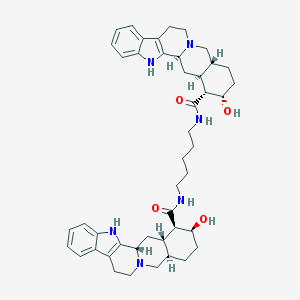 CHEMBL555190 CHEMBL555190 | C45H58N6O4 | 746.997 | 6 / 6 | 5.5 | No |
| 26025 | 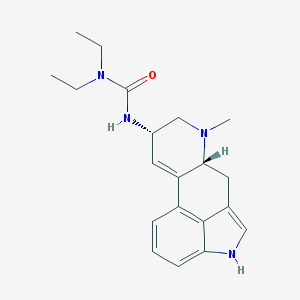 LISURIDE LISURIDE | C20H26N4O | 338.455 | 2 / 2 | 2.7 | Yes |
| 459443 | 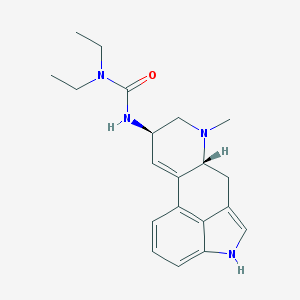 S-(-)-Lisuride S-(-)-Lisuride | C20H26N4O | 338.455 | 2 / 2 | 2.7 | Yes |
| 26091 | 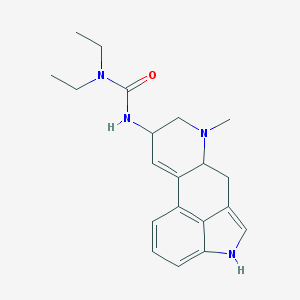 AC1L1H1T AC1L1H1T | C20H26N4O | 338.455 | 2 / 2 | 2.7 | Yes |
| 26304 | 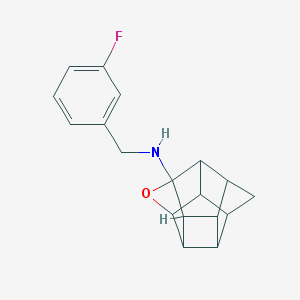 CHEMBL600053 CHEMBL600053 | C18H18FNO | 283.346 | 3 / 1 | 2.5 | Yes |
| 26546 | 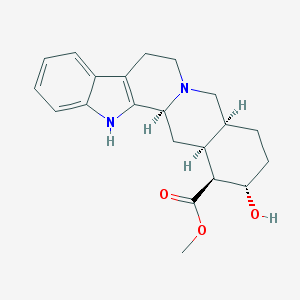 Rauwolscine Rauwolscine | C21H26N2O3 | 354.45 | 4 / 2 | 2.9 | Yes |
| 26567 | 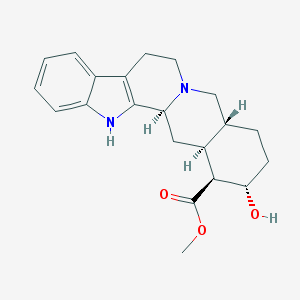 corynanthine corynanthine | C21H26N2O3 | 354.45 | 4 / 2 | 2.9 | Yes |
| 26582 | 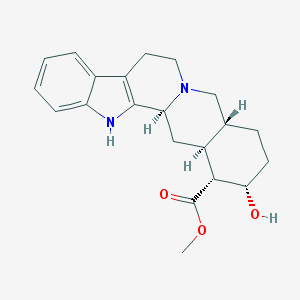 Yohimbine Yohimbine | C21H26N2O3 | 354.45 | 4 / 2 | 2.9 | Yes |
| 26634 | 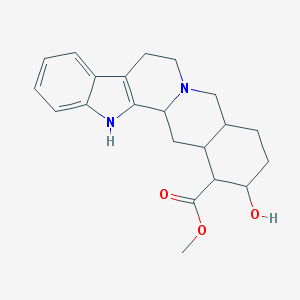 CHEBI:48565 CHEBI:48565 | C21H26N2O3 | 354.45 | 4 / 2 | 2.9 | Yes |
| 459448 | 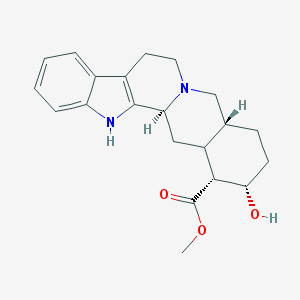 CHEMBL196400 CHEMBL196400 | C21H26N2O3 | 354.45 | 4 / 2 | 2.9 | Yes |
| 26937 | 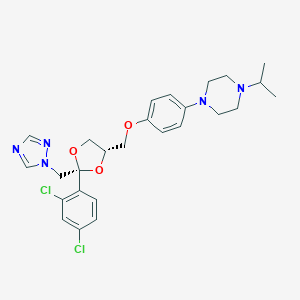 terconazole terconazole | C26H31Cl2N5O3 | 532.466 | 7 / 0 | 4.8 | No |
| 27215 | 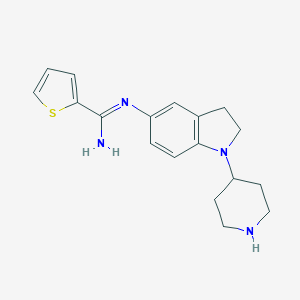 CHEMBL2203713 CHEMBL2203713 | C18H22N4S | 326.462 | 4 / 2 | 2.9 | Yes |
| 27509 | 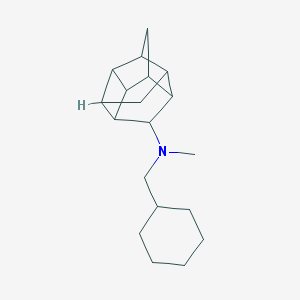 CHEMBL2432063 CHEMBL2432063 | C19H29N | 271.448 | 1 / 0 | 4.7 | Yes |
| 29583 | 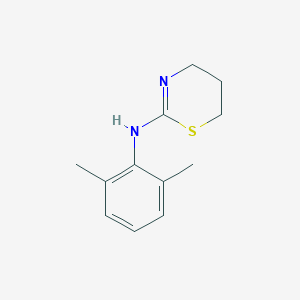 xylazine xylazine | C12H16N2S | 220.334 | 2 / 1 | 2.8 | Yes |
| 29663 | 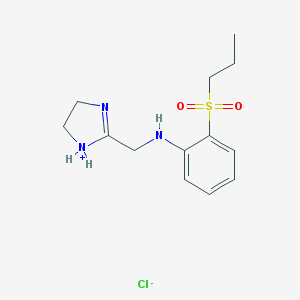 CID 10903225 CID 10903225 | C13H20ClN3O2S | 317.832 | 5 / 2 | N/A | N/A |
| 30204 | 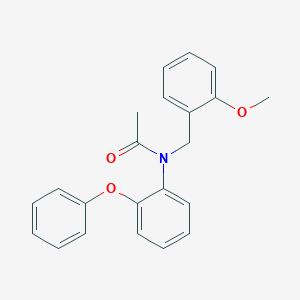 SCHEMBL14542291 SCHEMBL14542291 | C22H21NO3 | 347.414 | 3 / 0 | 4.3 | Yes |
| 31084 | 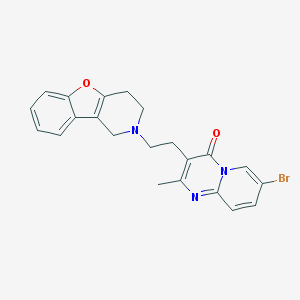 CHEMBL165350 CHEMBL165350 | C22H20BrN3O2 | 438.325 | 4 / 0 | 3.2 | Yes |
| 31597 | 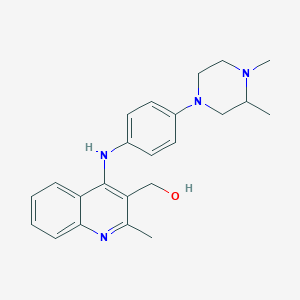 CHEMBL214996 CHEMBL214996 | C23H28N4O | 376.504 | 5 / 2 | 4.2 | Yes |
| 32110 | 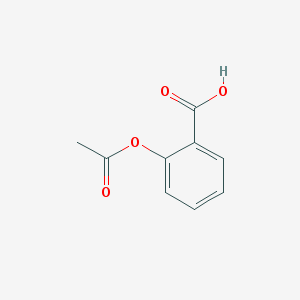 aspirin aspirin | C9H8O4 | 180.159 | 4 / 1 | 1.2 | Yes |
| 32539 | 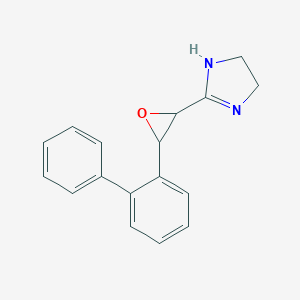 CHEMBL222933 CHEMBL222933 | C17H16N2O | 264.328 | 2 / 1 | 2.1 | Yes |
| 33060 | 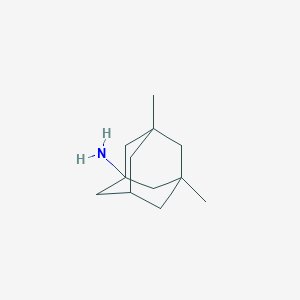 memantine memantine | C12H21N | 179.307 | 1 / 1 | 3.3 | Yes |
| 33395 | 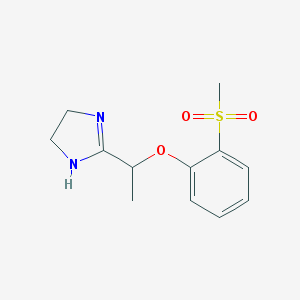 CHEMBL1956200 CHEMBL1956200 | C12H16N2O3S | 268.331 | 4 / 1 | 0.6 | Yes |
| 34138 | 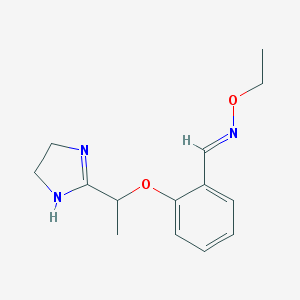 CHEMBL492639 CHEMBL492639 | C14H19N3O2 | 261.325 | 4 / 1 | 1.7 | Yes |
| 35013 | 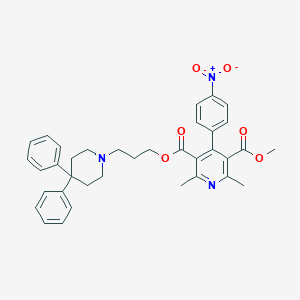 CHEMBL123769 CHEMBL123769 | C36H37N3O6 | 607.707 | 8 / 0 | 7.1 | No |
| 35253 | 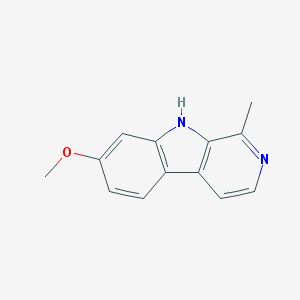 HARMINE HARMINE | C13H12N2O | 212.252 | 2 / 1 | 3.6 | Yes |
| 35645 | 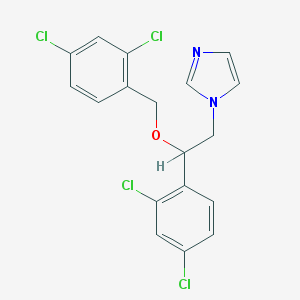 miconazole miconazole | C18H14Cl4N2O | 416.123 | 2 / 0 | 5.3 | No |
| 36993 | 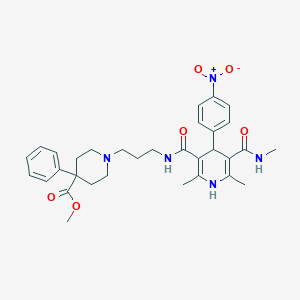 SNAP-5540 SNAP-5540 | C32H39N5O6 | 589.693 | 8 / 3 | 3.5 | No |
| 37877 | 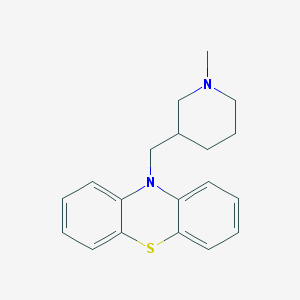 Pecazine Pecazine | C19H22N2S | 310.459 | 3 / 0 | 5.6 | No |
| 40067 | 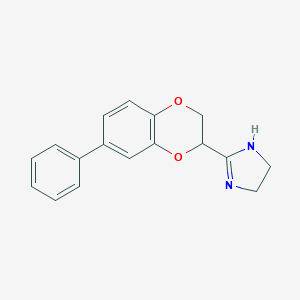 CHEMBL492445 CHEMBL492445 | C17H16N2O2 | 280.327 | 3 / 1 | 2.3 | Yes |
| 40249 | 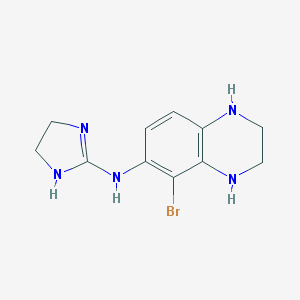 CHEMBL74283 CHEMBL74283 | C11H14BrN5 | 296.172 | 3 / 4 | 1.0 | Yes |
| 40286 | 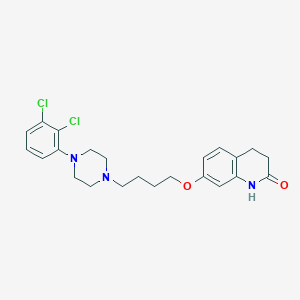 Aripiprazole Aripiprazole | C23H27Cl2N3O2 | 448.388 | 4 / 1 | 4.6 | Yes |
| 42466 | 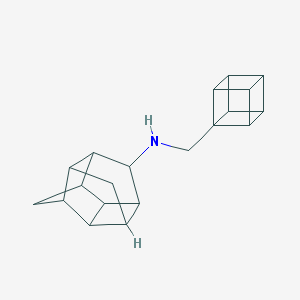 CHEMBL2432039 CHEMBL2432039 | C20H23N | 277.411 | 1 / 1 | 2.7 | Yes |
| 43019 | 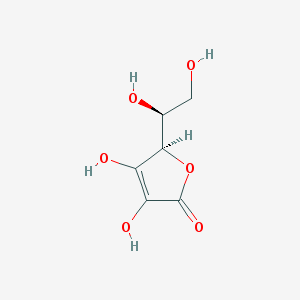 l-ascorbic acid l-ascorbic acid | C6H8O6 | 176.124 | 6 / 4 | -1.6 | Yes |
| 43276 | 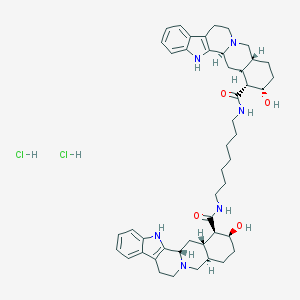 CHEMBL543232 CHEMBL543232 | C47H64Cl2N6O4 | 847.967 | 6 / 8 | N/A | No |
| 43331 | 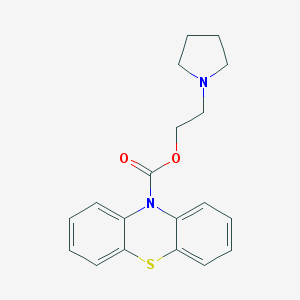 CHEMBL481153 CHEMBL481153 | C19H20N2O2S | 340.441 | 4 / 0 | 3.9 | Yes |
| 43992 | 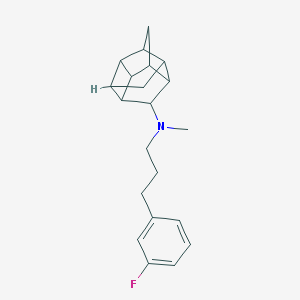 CHEMBL2432060 CHEMBL2432060 | C21H26FN | 311.444 | 2 / 0 | 4.8 | Yes |
| 44229 | 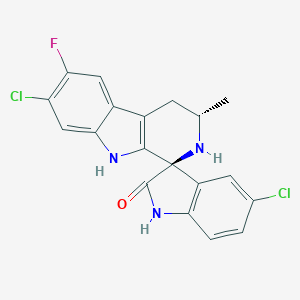 Cipargamin Cipargamin | C19H14Cl2FN3O | 390.239 | 3 / 3 | 3.9 | Yes |
| 44559 | 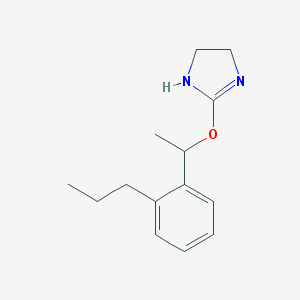 CHEMBL576078 CHEMBL576078 | C14H20N2O | 232.327 | 2 / 1 | 2.9 | Yes |
| 44793 | 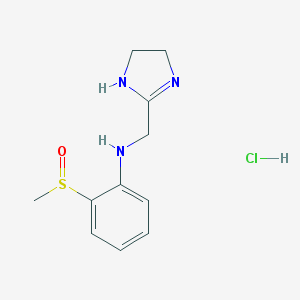 CHEMBL542402 CHEMBL542402 | C11H16ClN3OS | 273.779 | 4 / 3 | N/A | N/A |
| 45766 | 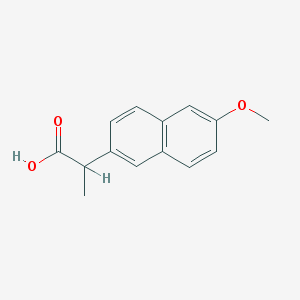 23981-80-8 23981-80-8 | C14H14O3 | 230.263 | 3 / 1 | 3.3 | Yes |
| 46057 | 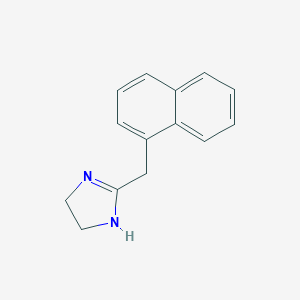 naphazoline naphazoline | C14H14N2 | 210.28 | 1 / 1 | 2.1 | Yes |
| 46217 | 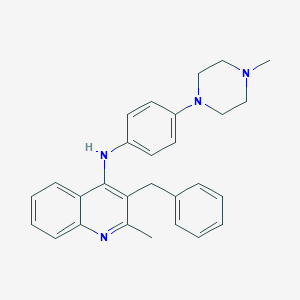 CHEMBL384925 CHEMBL384925 | C28H30N4 | 422.576 | 4 / 1 | 6.0 | No |
| 47498 | 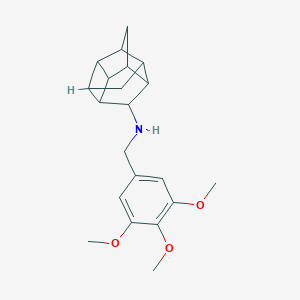 CHEMBL2432051 CHEMBL2432051 | C21H27NO3 | 341.451 | 4 / 1 | 3.3 | Yes |
| 47548 | 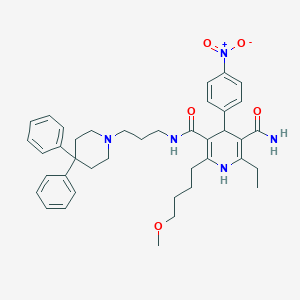 N-[3-(4,4-Diphenylpiperidino)propyl]-2-(4-methoxybutyl)-4-(4-nitrophenyl)-6-ethyl-1,4-dihydropyridine-3,5-dicarboxamide N-[3-(4,4-Diphenylpiperidino)propyl]-2-(4-methoxybutyl)-4-(4-nitrophenyl)-6-ethyl-1,4-dihydropyridine-3,5-dicarboxamide | C40H49N5O5 | 679.862 | 7 / 3 | 6.0 | No |
| 48553 | 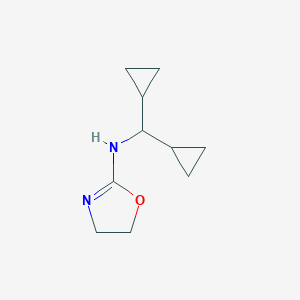 Rilmenidine Rilmenidine | C10H16N2O | 180.251 | 2 / 1 | 1.2 | Yes |
| 459657 | 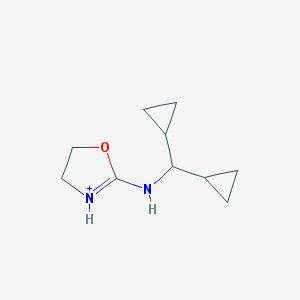 BDBM50262235 BDBM50262235 | C10H17N2O+ | 181.259 | 1 / 2 | 1.2 | Yes |
| 50010 | 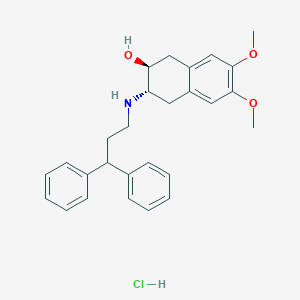 CHEMBL1203618 CHEMBL1203618 | C27H32ClNO3 | 454.007 | 4 / 3 | N/A | N/A |
| 50050 | 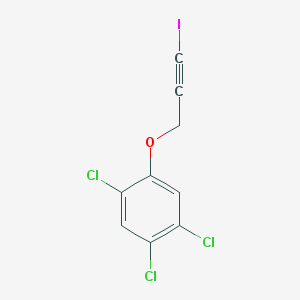 haloprogin haloprogin | C9H4Cl3IO | 361.384 | 1 / 0 | 4.6 | Yes |
| 459677 | 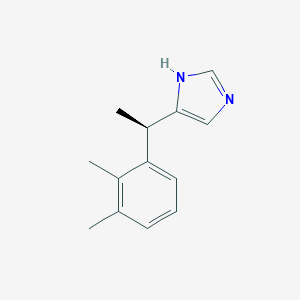 levomedetomidine levomedetomidine | C13H16N2 | 200.285 | 1 / 1 | 3.1 | Yes |
| 553490 | 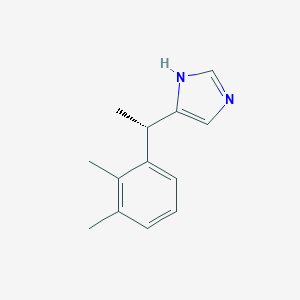 DEXMEDETOMIDINE DEXMEDETOMIDINE | C13H16N2 | 200.285 | 1 / 1 | 3.1 | Yes |
| 50759 | 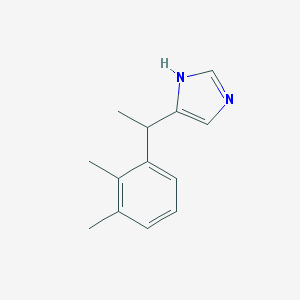 Medetomidine Medetomidine | C13H16N2 | 200.285 | 1 / 1 | 3.1 | Yes |
| 51149 | 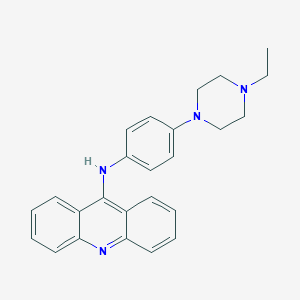 143069-08-3 143069-08-3 | C25H26N4 | 382.511 | 4 / 1 | 5.4 | No |
| 51549 | 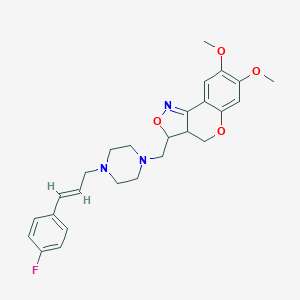 CHEMBL226636 CHEMBL226636 | C26H30FN3O4 | 467.541 | 8 / 0 | 3.7 | Yes |
| 52407 | 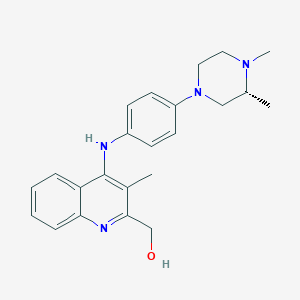 CHEMBL217254 CHEMBL217254 | C23H28N4O | 376.504 | 5 / 2 | 3.6 | Yes |
| 52578 | 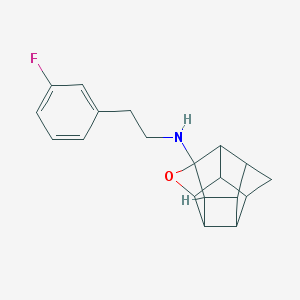 CHEMBL590615 CHEMBL590615 | C19H20FNO | 297.373 | 3 / 1 | 3.0 | Yes |
| 53043 | 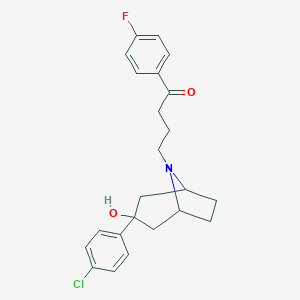 CHEMBL210578 CHEMBL210578 | C23H25ClFNO2 | 401.906 | 4 / 1 | 4.4 | Yes |
| 53784 | 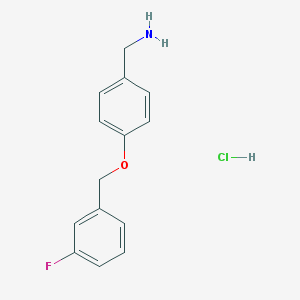 {4-[(3-fluorophenyl)methoxy]phenyl}methanamine hydrochloride {4-[(3-fluorophenyl)methoxy]phenyl}methanamine hydrochloride | C14H15ClFNO | 267.728 | 3 / 2 | N/A | N/A |
| 54926 | 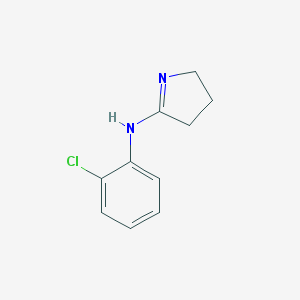 CHEMBL2058633 CHEMBL2058633 | C10H11ClN2 | 194.662 | 1 / 1 | 1.7 | Yes |
| 54938 | 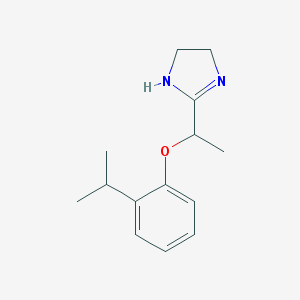 CHEMBL266613 CHEMBL266613 | C14H20N2O | 232.327 | 2 / 1 | 2.4 | Yes |
| 55052 | 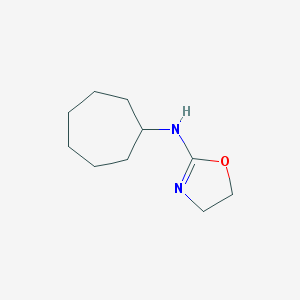 CHEMBL305928 CHEMBL305928 | C10H18N2O | 182.267 | 2 / 1 | 2.0 | Yes |
| 55636 | 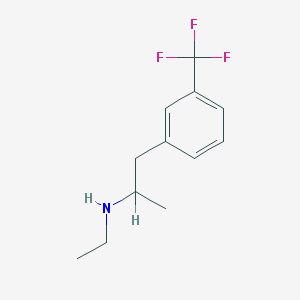 fenfluramine fenfluramine | C12H16F3N | 231.262 | 4 / 1 | 3.4 | Yes |
| 56318 | 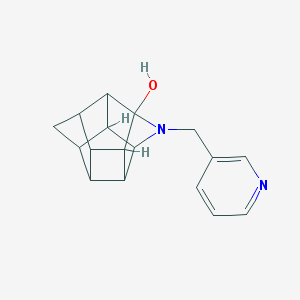 CHEMBL2205833 CHEMBL2205833 | C17H18N2O | 266.344 | 3 / 1 | 1.3 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218