

You can:
| Name | Beta-1 adrenergic receptor |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Adrb1 |
| Synonym | adrenergic receptor beta1-adrenoceptor Beta-1 adrenoreceptor Beta-1 adrenoceptor beta-1 adrenergic receptor [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 466 |
| Amino acid sequence | MGAGALALGASEPCNLSSAAPLPDGAATAARLLVLASPPASLLPPASEGSAPLSQQWTAGMGLLLALIVLLIVVGNVLVIVAIAKTPRLQTLTNLFIMSLASADLVMGLLVVPFGATIVVWGRWEYGSFFCELWTSVDVLCVTASIETLCVIALDRYLAITLPFRYQSLLTRARARALVCTVWAISALVSFLPILMHWWRAESDEARRCYNDPKCCDFVTNRAYAIASSVVSFYVPLCIMAFVYLRVFREAQKQVKKIDSCERRFLTGPPRPPSPAPSPSPGPPRPADSLANGRSSKRRPSRLVALREQKALKTLGIIMGVFTLCWLPFFLANVVKAFHRDLVPDRLFVFFNWLGYANSAFNPIIYCRSPDFRKAFQRLLCCARRAACRRRAAHGDRPRASGCLARAGPPPSPGAPSDDDDDDAGATPPARLLEPWAGCNGGTTTVDSDSSLDEPGRQGFSSESKV |
| UniProt | P18090 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3252 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 129 | 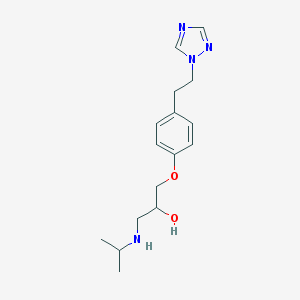 CHEMBL149540 CHEMBL149540 | C16H24N4O2 | 304.394 | 5 / 2 | 1.6 | Yes |
| 1262 | 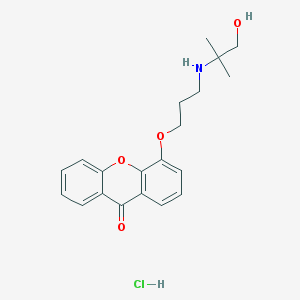 CHEMBL479644 CHEMBL479644 | C20H24ClNO4 | 377.865 | 5 / 3 | N/A | N/A |
| 2145 | 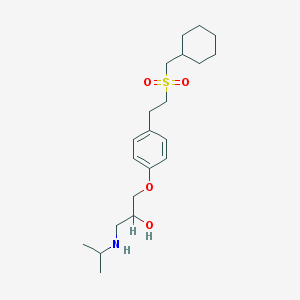 CHEMBL544661 CHEMBL544661 | C21H35NO4S | 397.574 | 5 / 2 | 3.8 | Yes |
| 3126 | 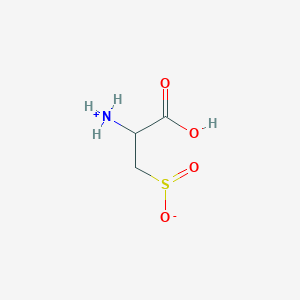 AC1N0YGA AC1N0YGA | C3H7NO4S | 153.152 | 5 / 2 | -4.7 | Yes |
| 5140 | 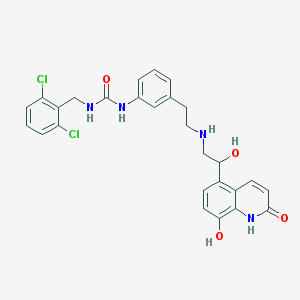 CHEMBL1682217 CHEMBL1682217 | C27H26Cl2N4O4 | 541.429 | 5 / 6 | 3.4 | No |
| 6723 | 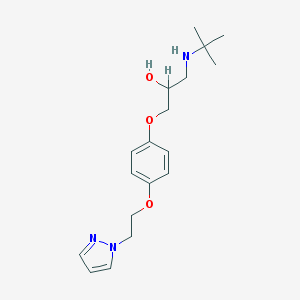 CHEMBL358360 CHEMBL358360 | C18H27N3O3 | 333.432 | 5 / 2 | 1.7 | Yes |
| 7239 | 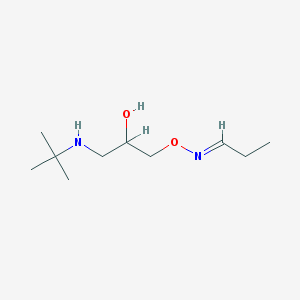 CHEMBL1178704 CHEMBL1178704 | C10H22N2O2 | 202.298 | 4 / 2 | 0.7 | Yes |
| 10166 | 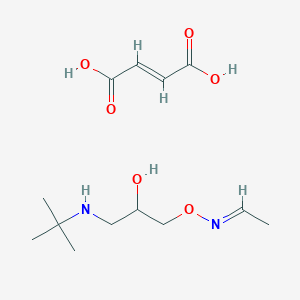 CID 44285269 CID 44285269 | C13H24N2O6 | 304.343 | 8 / 4 | N/A | N/A |
| 10581 | 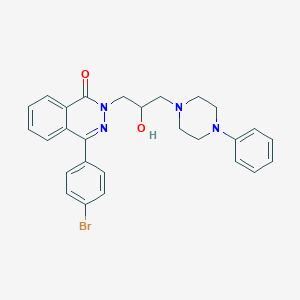 CHEMBL2283384 CHEMBL2283384 | C27H27BrN4O2 | 519.443 | 5 / 1 | 4.6 | No |
| 11720 | 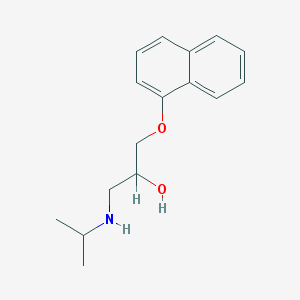 propranolol propranolol | C16H21NO2 | 259.349 | 3 / 2 | 3.0 | Yes |
| 12510 | 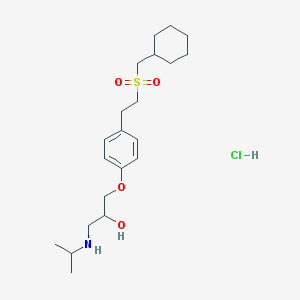 CHEMBL544661 CHEMBL544661 | C21H36ClNO4S | 434.032 | 5 / 3 | N/A | N/A |
| 13029 | 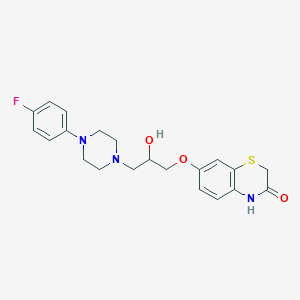 CHEMBL143944 CHEMBL143944 | C21H24FN3O3S | 417.499 | 7 / 2 | 2.5 | Yes |
| 13066 | 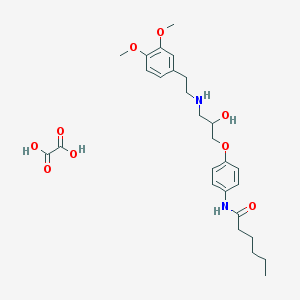 CHEMBL3230442 CHEMBL3230442 | C27H38N2O9 | 534.606 | 10 / 5 | N/A | No |
| 14579 | 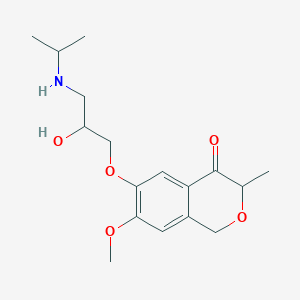 CHEMBL2375978 CHEMBL2375978 | C17H25NO5 | 323.389 | 6 / 2 | 1.2 | Yes |
| 15866 | 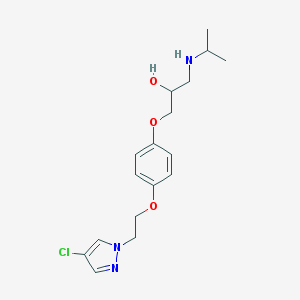 CHEMBL146767 CHEMBL146767 | C17H24ClN3O3 | 353.847 | 5 / 2 | 2.2 | Yes |
| 16559 | 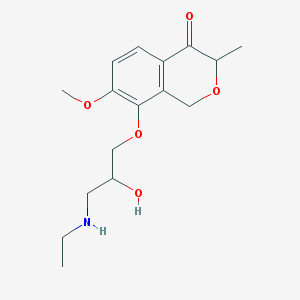 CHEMBL2375990 CHEMBL2375990 | C16H23NO5 | 309.362 | 6 / 2 | 0.8 | Yes |
| 19634 | 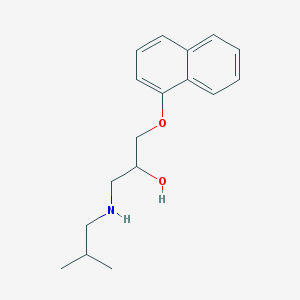 CHEMBL287651 CHEMBL287651 | C17H23NO2 | 273.376 | 3 / 2 | 3.5 | Yes |
| 20171 | 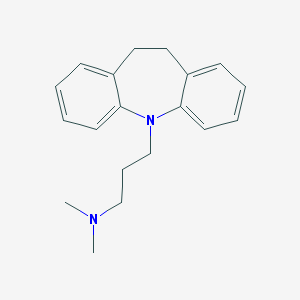 imipramine imipramine | C19H24N2 | 280.415 | 2 / 0 | 4.8 | Yes |
| 21602 | 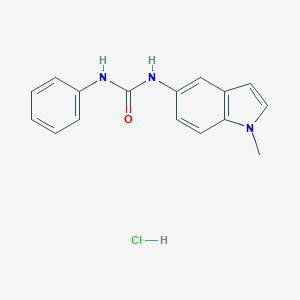 CHEMBL543964 CHEMBL543964 | C16H16ClN3O | 301.774 | 1 / 3 | N/A | N/A |
| 23769 | 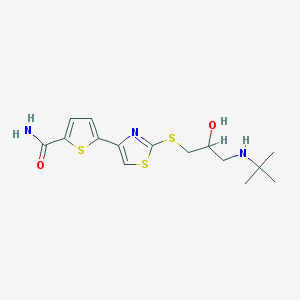 Arotinolol Arotinolol | C15H21N3O2S3 | 371.532 | 7 / 3 | 2.3 | Yes |
| 24668 | 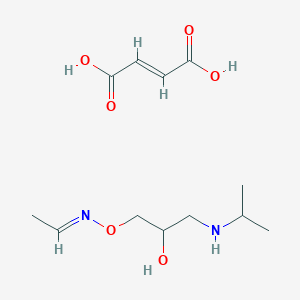 CID 44285288 CID 44285288 | C12H22N2O6 | 290.316 | 8 / 4 | N/A | N/A |
| 26185 | 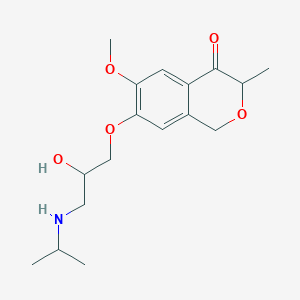 CHEMBL2375985 CHEMBL2375985 | C17H25NO5 | 323.389 | 6 / 2 | 1.2 | Yes |
| 26701 | 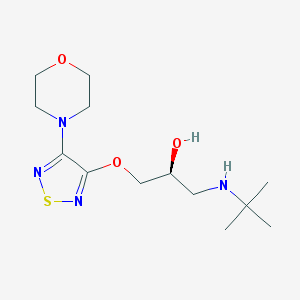 timolol timolol | C13H24N4O3S | 316.42 | 8 / 2 | 1.8 | Yes |
| 27787 | 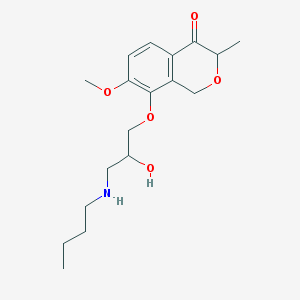 CHEMBL2375993 CHEMBL2375993 | C18H27NO5 | 337.416 | 6 / 2 | 1.7 | Yes |
| 28861 | 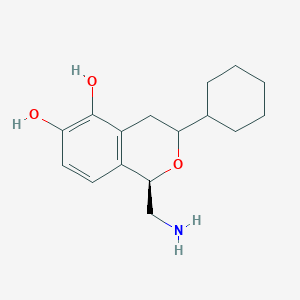 CHEMBL314459 CHEMBL314459 | C16H23NO3 | 277.364 | 4 / 3 | 2.4 | Yes |
| 35855 | 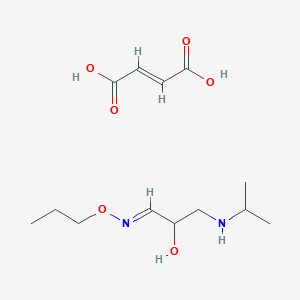 CID 44285228 CID 44285228 | C13H24N2O6 | 304.343 | 8 / 4 | N/A | N/A |
| 36960 | 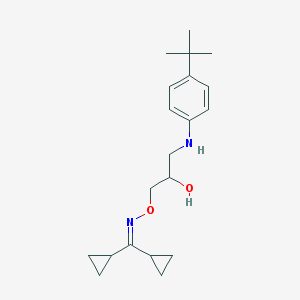 CHEMBL42117 CHEMBL42117 | C20H30N2O2 | 330.472 | 4 / 2 | 4.2 | Yes |
| 39788 | 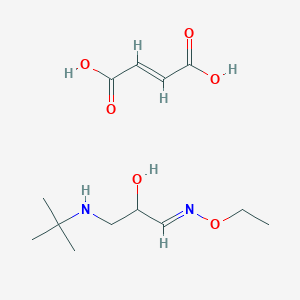 CID 44285227 CID 44285227 | C13H24N2O6 | 304.343 | 8 / 4 | N/A | N/A |
| 40032 | 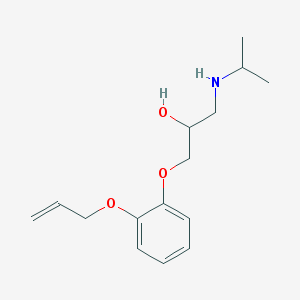 oxprenolol oxprenolol | C15H23NO3 | 265.353 | 4 / 2 | 2.1 | Yes |
| 40792 | 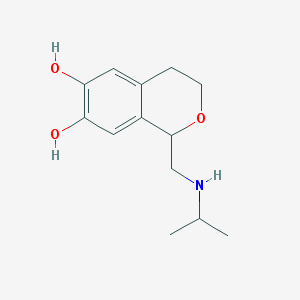 CHEMBL322842 CHEMBL322842 | C13H19NO3 | 237.299 | 4 / 3 | 1.3 | Yes |
| 41416 | 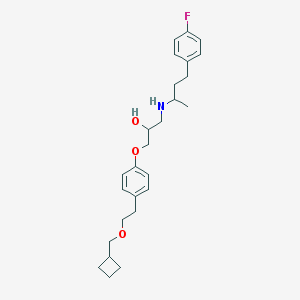 CHEMBL6749 CHEMBL6749 | C26H36FNO3 | 429.576 | 5 / 2 | 5.4 | No |
| 41889 | 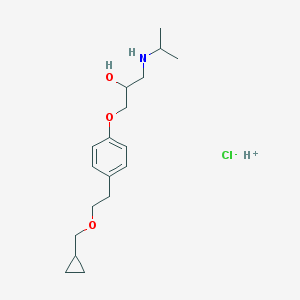 AC1LCWB4 AC1LCWB4 | C18H30ClNO3 | 343.892 | 5 / 3 | N/A | N/A |
| 43230 | 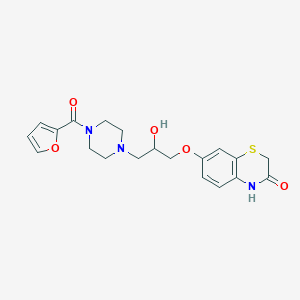 CHEMBL143733 CHEMBL143733 | C20H23N3O5S | 417.48 | 7 / 2 | 1.1 | Yes |
| 45548 | 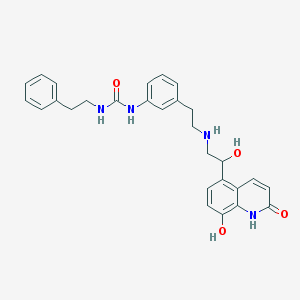 CHEMBL1682212 CHEMBL1682212 | C28H30N4O4 | 486.572 | 5 / 6 | 2.6 | No |
| 45798 | 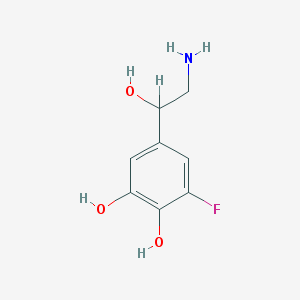 CHEMBL129190 CHEMBL129190 | C8H10FNO3 | 187.17 | 5 / 4 | -0.5 | Yes |
| 46536 | 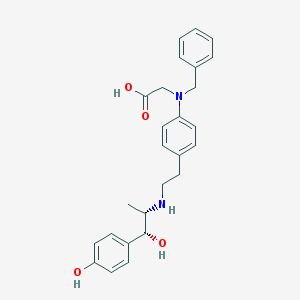 CHEMBL32470 CHEMBL32470 | C26H30N2O4 | 434.536 | 6 / 4 | 1.6 | Yes |
| 47016 | 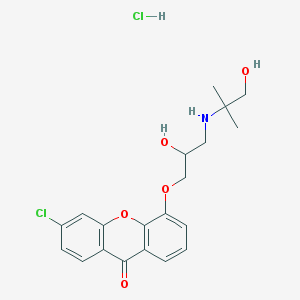 CHEMBL520600 CHEMBL520600 | C20H23Cl2NO5 | 428.306 | 6 / 4 | N/A | N/A |
| 51286 | 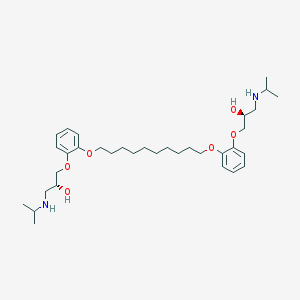 CHEMBL41696 CHEMBL41696 | C34H56N2O6 | 588.83 | 8 / 4 | 6.6 | No |
| 53365 | 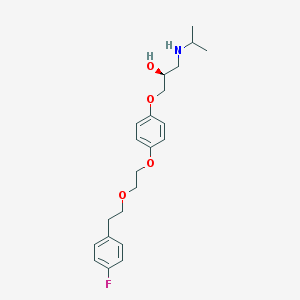 Flusoxolol Flusoxolol | C22H30FNO4 | 391.483 | 6 / 2 | 3.7 | Yes |
| 53366 | 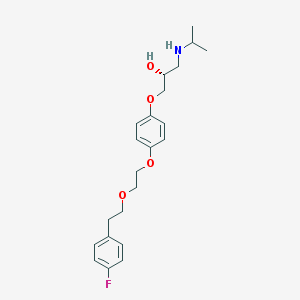 AC1OCD6K AC1OCD6K | C22H30FNO4 | 391.483 | 6 / 2 | 3.7 | Yes |
| 53368 | 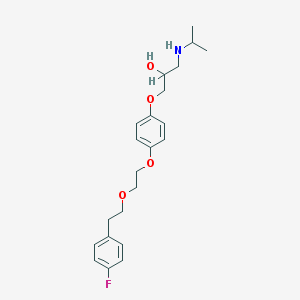 CHEMBL43405 CHEMBL43405 | C22H30FNO4 | 391.483 | 6 / 2 | 3.7 | Yes |
| 559092 | 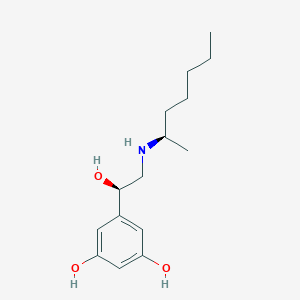 BDBM206775 BDBM206775 | C15H25NO3 | 267.369 | 4 / 4 | 2.7 | Yes |
| 559094 | 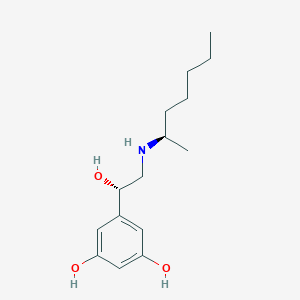 BDBM206777 BDBM206777 | C15H25NO3 | 267.369 | 4 / 4 | 2.7 | Yes |
| 537423 | 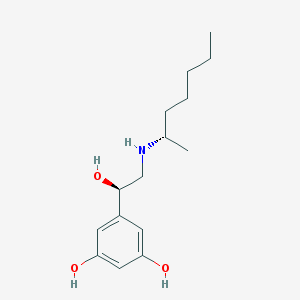 CHEMBL3930835 CHEMBL3930835 | C15H25NO3 | 267.369 | 4 / 4 | 2.7 | Yes |
| 537426 | 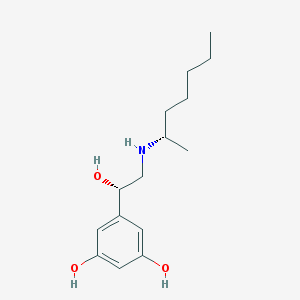 SCHEMBL2488461 SCHEMBL2488461 | C15H25NO3 | 267.369 | 4 / 4 | 2.7 | Yes |
| 59743 | 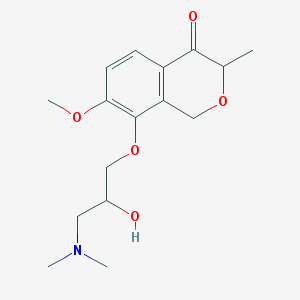 CHEMBL2375995 CHEMBL2375995 | C16H23NO5 | 309.362 | 6 / 1 | 0.9 | Yes |
| 61595 | 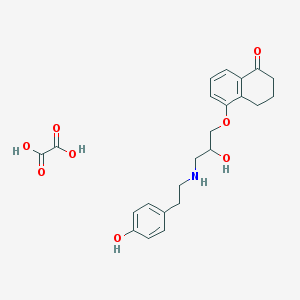 CHEMBL3230447 CHEMBL3230447 | C23H27NO8 | 445.468 | 9 / 5 | N/A | N/A |
| 62064 | 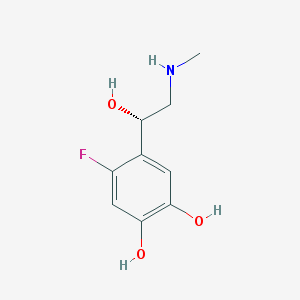 CHEMBL42216 CHEMBL42216 | C9H12FNO3 | 201.197 | 5 / 4 | 0.0 | Yes |
| 62066 | 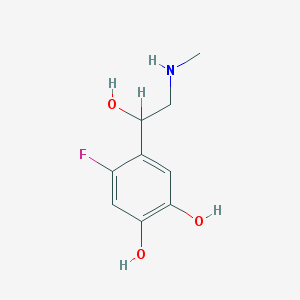 CHEMBL42359 CHEMBL42359 | C9H12FNO3 | 201.197 | 5 / 4 | 0.0 | Yes |
| 62068 | 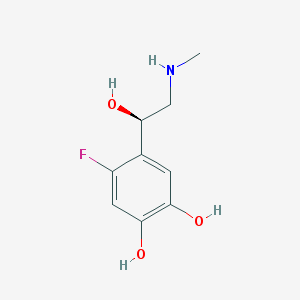 CHEMBL40986 CHEMBL40986 | C9H12FNO3 | 201.197 | 5 / 4 | 0.0 | Yes |
| 63923 | 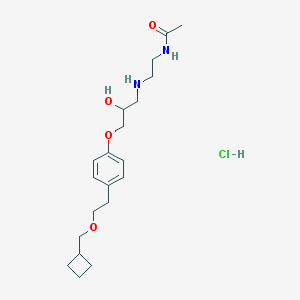 CHEMBL543484 CHEMBL543484 | C20H33ClN2O4 | 400.944 | 5 / 4 | N/A | N/A |
| 66146 | 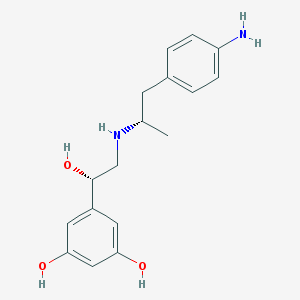 CHEMBL229400 CHEMBL229400 | C17H22N2O3 | 302.374 | 5 / 5 | 1.6 | Yes |
| 66148 | 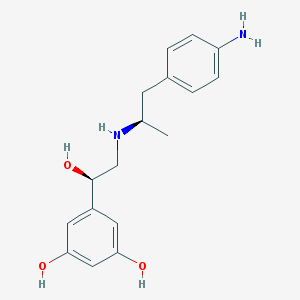 CHEMBL229391 CHEMBL229391 | C17H22N2O3 | 302.374 | 5 / 5 | 1.6 | Yes |
| 66150 | 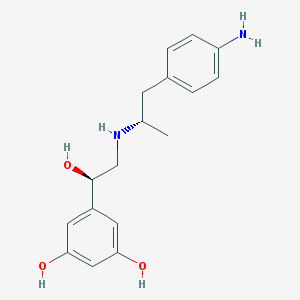 CHEMBL229442 CHEMBL229442 | C17H22N2O3 | 302.374 | 5 / 5 | 1.6 | Yes |
| 66151 | 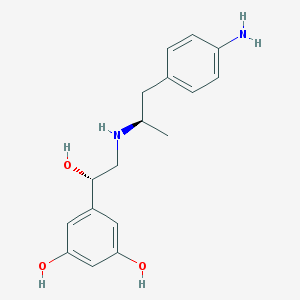 CHEMBL229443 CHEMBL229443 | C17H22N2O3 | 302.374 | 5 / 5 | 1.6 | Yes |
| 68400 | 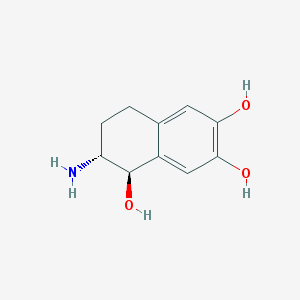 CHEMBL7185 CHEMBL7185 | C10H13NO3 | 195.218 | 4 / 4 | 0.1 | Yes |
| 68705 | 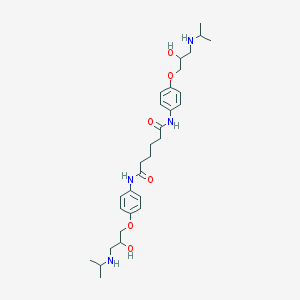 CHEMBL164145 CHEMBL164145 | C30H46N4O6 | 558.72 | 8 / 6 | 2.0 | No |
| 69361 | 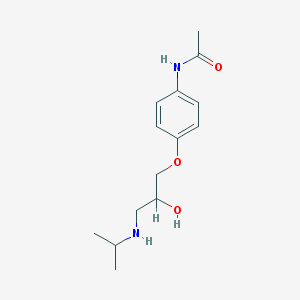 practolol practolol | C14H22N2O3 | 266.341 | 4 / 3 | 0.8 | Yes |
| 73161 | 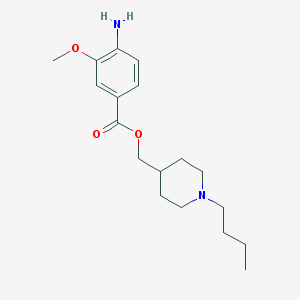 CHEMBL1258452 CHEMBL1258452 | C18H28N2O3 | 320.433 | 5 / 1 | 3.1 | Yes |
| 75793 | 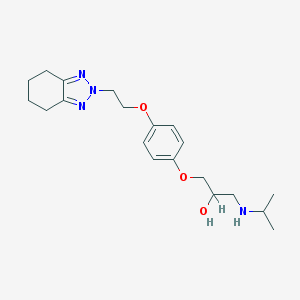 CHEMBL444003 CHEMBL444003 | C20H30N4O3 | 374.485 | 6 / 2 | 2.6 | Yes |
| 76053 | 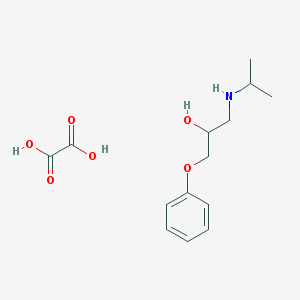 CHEMBL3230436 CHEMBL3230436 | C14H21NO6 | 299.323 | 7 / 4 | N/A | N/A |
| 76929 | 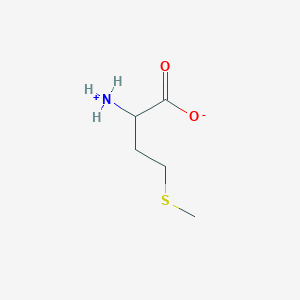 methionine zwitterion methionine zwitterion | C5H11NO2S | 149.208 | 3 / 1 | -1.2 | Yes |
| 77002 | 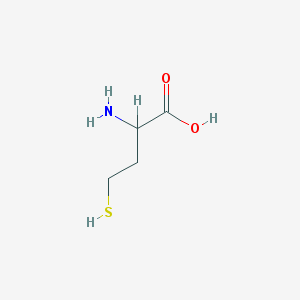 DL-Homocysteine DL-Homocysteine | C4H9NO2S | 135.181 | 4 / 3 | -3.4 | Yes |
| 77132 | 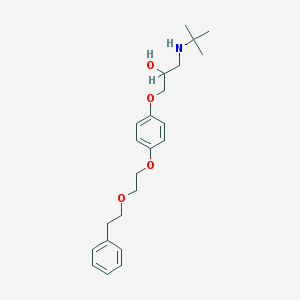 CHEMBL36211 CHEMBL36211 | C23H33NO4 | 387.52 | 5 / 2 | 3.8 | Yes |
| 80685 | 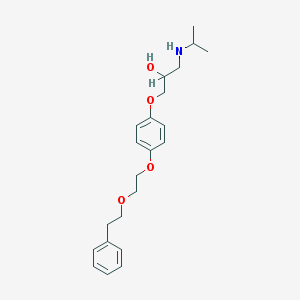 CHEMBL41360 CHEMBL41360 | C22H31NO4 | 373.493 | 5 / 2 | 3.6 | Yes |
| 82821 | 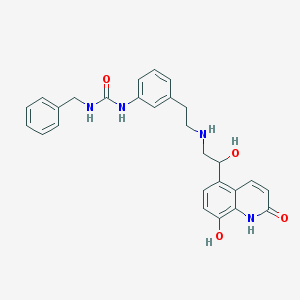 CHEMBL1682211 CHEMBL1682211 | C27H28N4O4 | 472.545 | 5 / 6 | 2.2 | No |
| 83598 | 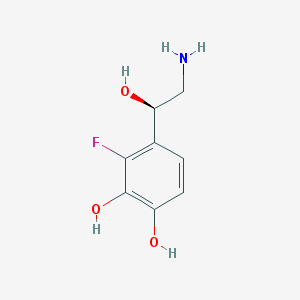 6-fluoronorepinehprine 6-fluoronorepinehprine | C8H10FNO3 | 187.17 | 5 / 4 | -0.5 | Yes |
| 83601 | 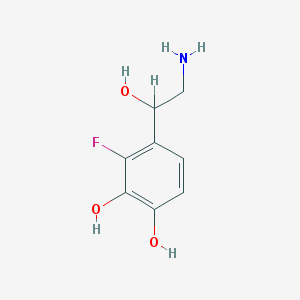 CHEMBL287587 CHEMBL287587 | C8H10FNO3 | 187.17 | 5 / 4 | -0.5 | Yes |
| 83607 | 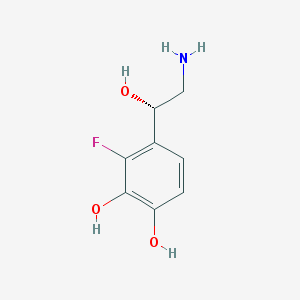 CHEMBL40796 CHEMBL40796 | C8H10FNO3 | 187.17 | 5 / 4 | -0.5 | Yes |
| 83965 | 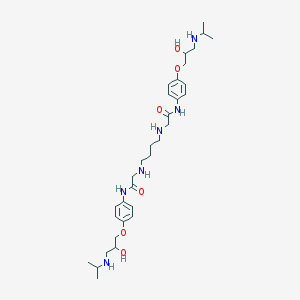 CHEMBL349175 CHEMBL349175 | C32H52N6O6 | 616.804 | 10 / 8 | 1.3 | No |
| 85681 | 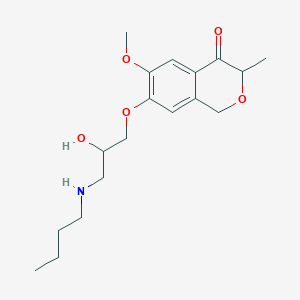 CHEMBL2375986 CHEMBL2375986 | C18H27NO5 | 337.416 | 6 / 2 | 1.7 | Yes |
| 87869 | 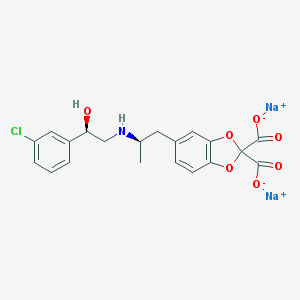 CL-316243 CL-316243 | C20H18ClNNa2O7 | 465.794 | 8 / 2 | N/A | N/A |
| 91383 | 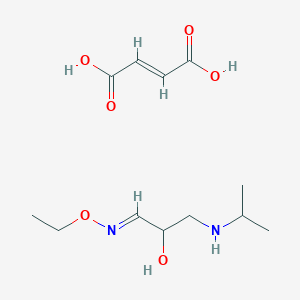 CID 44285235 CID 44285235 | C12H22N2O6 | 290.316 | 8 / 4 | N/A | N/A |
| 91410 | 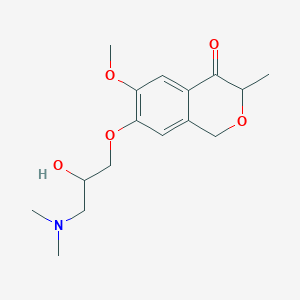 CHEMBL2375988 CHEMBL2375988 | C16H23NO5 | 309.362 | 6 / 1 | 0.9 | Yes |
| 91628 | 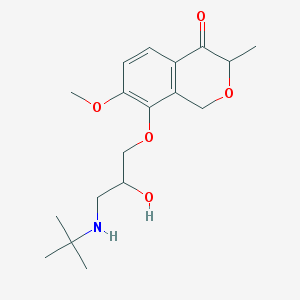 CHEMBL2375994 CHEMBL2375994 | C18H27NO5 | 337.416 | 6 / 2 | 1.4 | Yes |
| 94770 | 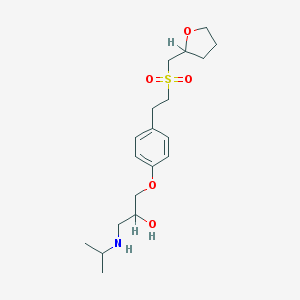 CHEMBL542777 CHEMBL542777 | C19H31NO5S | 385.519 | 6 / 2 | 1.7 | Yes |
| 94805 | 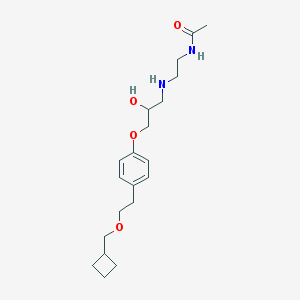 CHEMBL543484 CHEMBL543484 | C20H32N2O4 | 364.486 | 5 / 3 | 1.9 | Yes |
| 95568 | 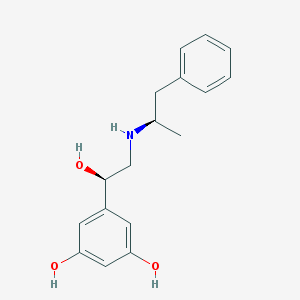 CHEMBL389629 CHEMBL389629 | C17H21NO3 | 287.359 | 4 / 4 | 2.3 | Yes |
| 95569 | 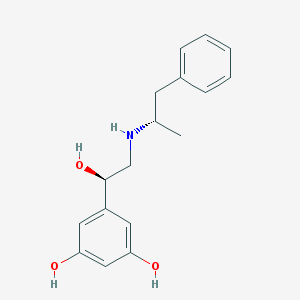 CHEMBL229388 CHEMBL229388 | C17H21NO3 | 287.359 | 4 / 4 | 2.3 | Yes |
| 95572 | 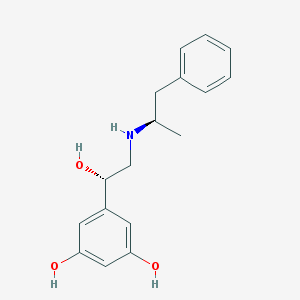 CHEMBL387825 CHEMBL387825 | C17H21NO3 | 287.359 | 4 / 4 | 2.3 | Yes |
| 95574 | 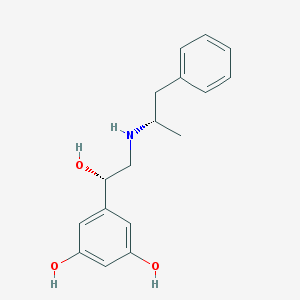 CHEMBL229390 CHEMBL229390 | C17H21NO3 | 287.359 | 4 / 4 | 2.3 | Yes |
| 96383 | 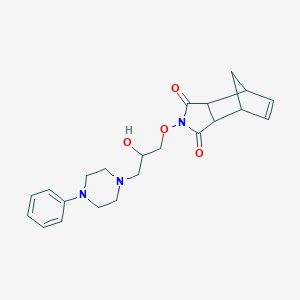 CHEMBL2261429 CHEMBL2261429 | C22H27N3O4 | 397.475 | 6 / 1 | 1.3 | Yes |
| 96654 | 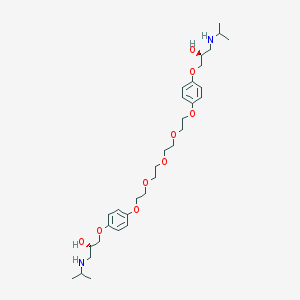 CHEMBL41607 CHEMBL41607 | C32H52N2O9 | 608.773 | 11 / 4 | 2.6 | No |
| 98503 | 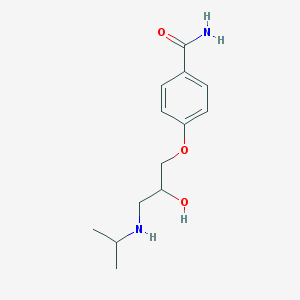 BRN 2116323 BRN 2116323 | C13H20N2O3 | 252.314 | 4 / 3 | 0.6 | Yes |
| 98627 | 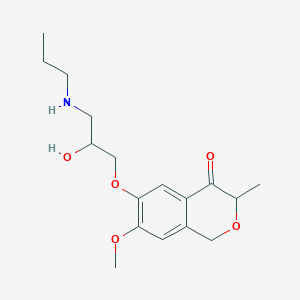 CHEMBL2375998 CHEMBL2375998 | C17H25NO5 | 323.389 | 6 / 2 | 1.3 | Yes |
| 98727 | 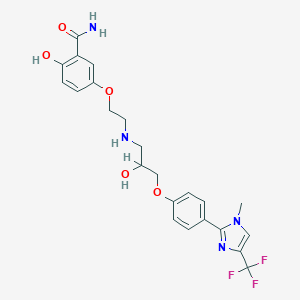 CGP 20712A CGP 20712A | C23H25F3N4O5 | 494.471 | 10 / 4 | 2.3 | Yes |
| 99466 | 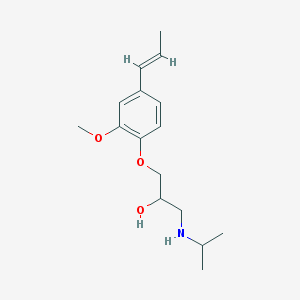 CHEMBL3247308 CHEMBL3247308 | C16H25NO3 | 279.38 | 4 / 2 | 2.7 | Yes |
| 99468 | 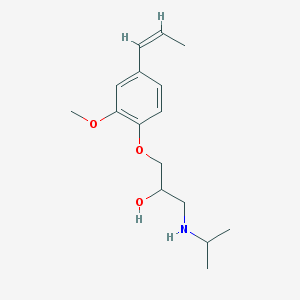 CHEMBL3247309 CHEMBL3247309 | C16H25NO3 | 279.38 | 4 / 2 | 2.7 | Yes |
| 99701 | 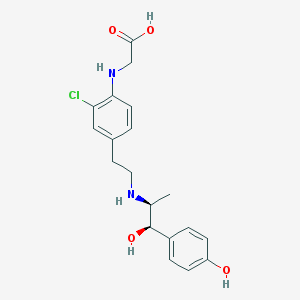 CHEMBL32670 CHEMBL32670 | C19H23ClN2O4 | 378.853 | 6 / 5 | 0.6 | Yes |
| 100102 | 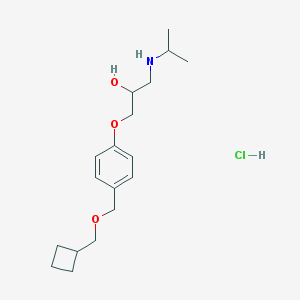 CHEMBL552898 CHEMBL552898 | C18H30ClNO3 | 343.892 | 4 / 3 | N/A | N/A |
| 100424 | 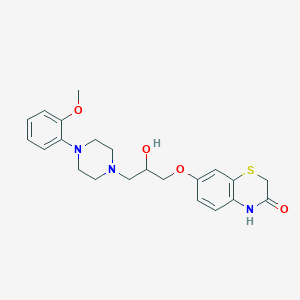 CHEMBL142948 CHEMBL142948 | C22H27N3O4S | 429.535 | 7 / 2 | 2.4 | Yes |
| 100676 | 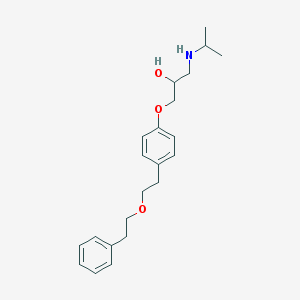 CHEMBL287886 CHEMBL287886 | C22H31NO3 | 357.494 | 4 / 2 | 3.8 | Yes |
| 101137 | 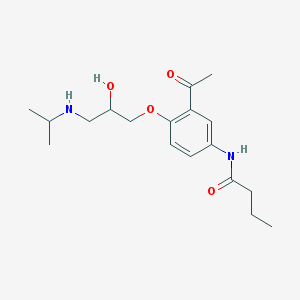 acebutolol acebutolol | C18H28N2O4 | 336.432 | 5 / 3 | 1.7 | Yes |
| 102493 | 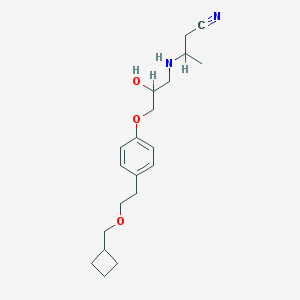 CHEMBL543717 CHEMBL543717 | C20H30N2O3 | 346.471 | 5 / 2 | 2.7 | Yes |
| 102907 | 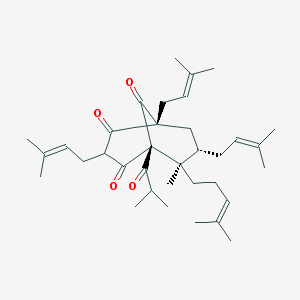 CHEMBL516641 CHEMBL516641 | C35H52O4 | 536.797 | 4 / 0 | 10.0 | No |
| 107343 | 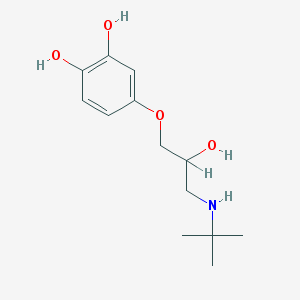 CHEMBL266619 CHEMBL266619 | C13H21NO4 | 255.314 | 5 / 4 | 1.1 | Yes |
| 108334 | 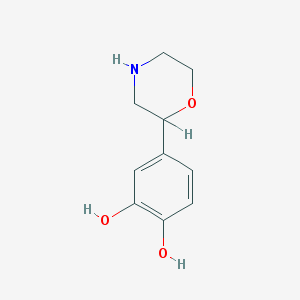 4-(2-Morpholinyl)pyrocatechol 4-(2-Morpholinyl)pyrocatechol | C10H13NO3 | 195.218 | 4 / 3 | 0.6 | Yes |
| 108874 | 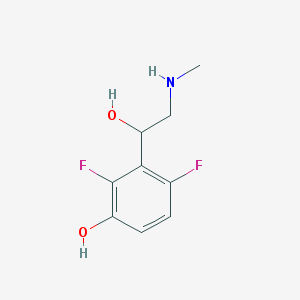 CHEMBL340189 CHEMBL340189 | C9H11F2NO2 | 203.189 | 5 / 3 | 0.5 | Yes |
| 113532 | 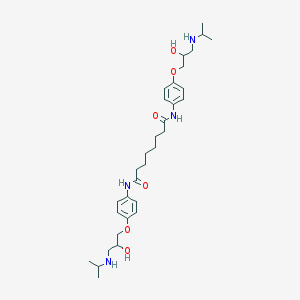 CHEMBL164146 CHEMBL164146 | C32H50N4O6 | 586.774 | 8 / 6 | 3.1 | No |
| 114197 | 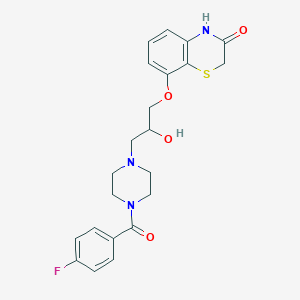 CHEMBL142496 CHEMBL142496 | C22H24FN3O4S | 445.509 | 7 / 2 | 1.8 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218