

You can:
| Name | D(4) dopamine receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | DRD4 |
| Synonym | D4 receptor D4R Dopamine D4 receptor dopamine receptor 4 d(2C) dopamine receptor |
| Disease | Erectile dysfunction Psychotic disorders Schizophrenia |
| Length | 467 |
| Amino acid sequence | MGNRSTADADGLLAGRGPAAGASAGASAGLAGQGAAALVGGVLLIGAVLAGNSLVCVSVATERALQTPTNSFIVSLAAADLLLALLVLPLFVYSEVQGGAWLLSPRLCDALMAMDVMLCTASIFNLCAISVDRFVAVAVPLRYNRQGGSRRQLLLIGATWLLSAAVAAPVLCGLNDVRGRDPAVCRLEDRDYVVYSSVCSFFLPCPLMLLLYWATFRGLQRWEVARRAKLHGRAPRRPSGPGPPSPTPPAPRLPQDPCGPDCAPPAPGLPRGPCGPDCAPAAPGLPPDPCGPDCAPPAPGLPQDPCGPDCAPPAPGLPRGPCGPDCAPPAPGLPQDPCGPDCAPPAPGLPPDPCGSNCAPPDAVRAAALPPQTPPQTRRRRRAKITGRERKAMRVLPVVVGAFLLCWTPFFVVHITQALCPACSVPPRLVSAVTWLGYVNSALNPVIYTVFNAEFRNVFRKALRACC |
| UniProt | P21917 |
| Protein Data Bank | 5wiv, 5wiu |
| GPCR-HGmod model | P21917 |
| 3D structure model | This structure is from PDB ID 5wiv. |
| BioLiP | BL0394824, BL0394826, BL0394825 |
| Therapeutic Target Database | T24983 |
| ChEMBL | CHEMBL219 |
| IUPHAR | 217 |
| DrugBank | BE0009376, BE0000389 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 82 | 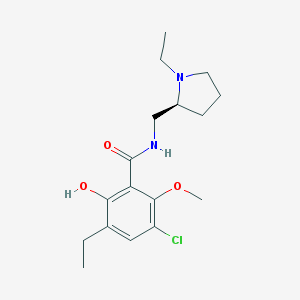 Eticlopride Eticlopride | C17H25ClN2O3 | 340.848 | 4 / 2 | 3.1 | Yes |
| 94 | 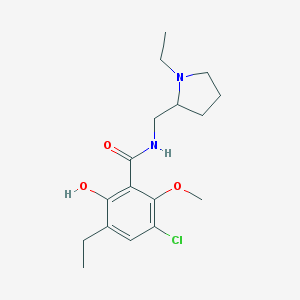 AC1O8PYY AC1O8PYY | C17H25ClN2O3 | 340.848 | 4 / 2 | 3.1 | Yes |
| 521443 | 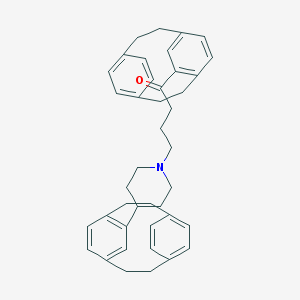 CHEMBL1256170 CHEMBL1256170 | C41H45NO | 567.817 | 2 / 0 | 9.2 | No |
| 535919 | 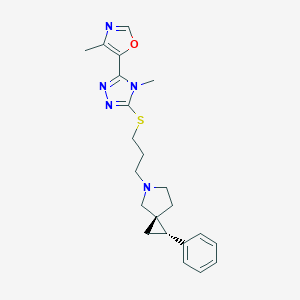 CHEMBL3983937 CHEMBL3983937 | C22H27N5OS | 409.552 | 6 / 0 | 3.3 | Yes |
| 443 | 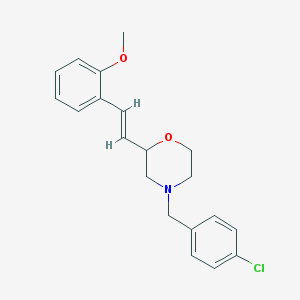 CHEMBL320071 CHEMBL320071 | C20H22ClNO2 | 343.851 | 3 / 0 | 4.2 | Yes |
| 472 | 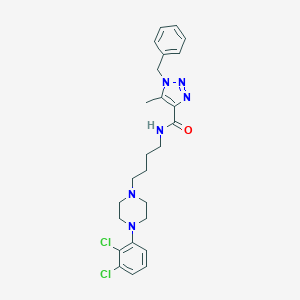 CHEMBL373873 CHEMBL373873 | C25H30Cl2N6O | 501.456 | 5 / 1 | 4.9 | No |
| 521453 | 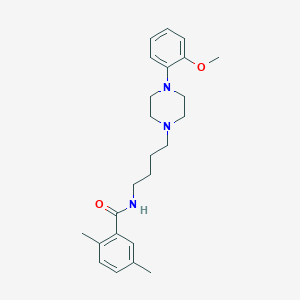 CHEMBL1258494 CHEMBL1258494 | C24H33N3O2 | 395.547 | 4 / 1 | 4.2 | Yes |
| 943 | 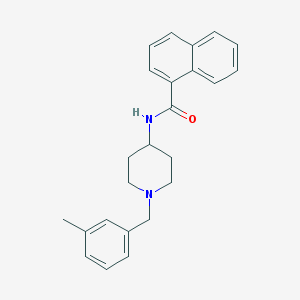 CHEMBL233404 CHEMBL233404 | C24H26N2O | 358.485 | 2 / 1 | 4.9 | Yes |
| 985 | 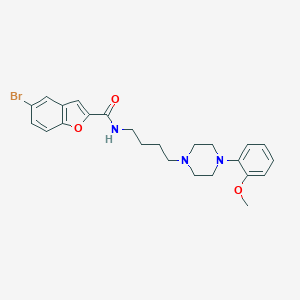 CHEMBL221681 CHEMBL221681 | C24H28BrN3O3 | 486.41 | 5 / 1 | 4.9 | Yes |
| 1019 | 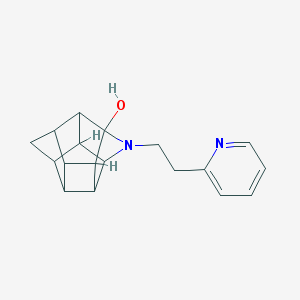 CHEMBL2205816 CHEMBL2205816 | C18H20N2O | 280.371 | 3 / 1 | 1.8 | Yes |
| 1059 | 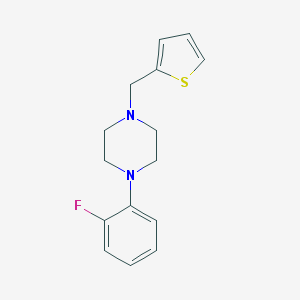 CHEMBL2420770 CHEMBL2420770 | C15H17FN2S | 276.373 | 4 / 0 | 3.2 | Yes |
| 1299 | 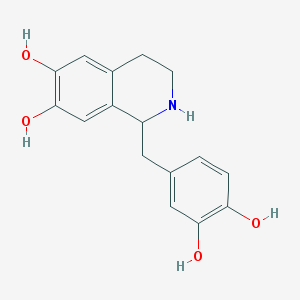 Tetrahydropapaveroline Tetrahydropapaveroline | C16H17NO4 | 287.315 | 5 / 5 | 1.9 | Yes |
| 1585 | 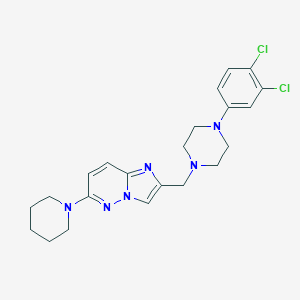 CHEMBL207543 CHEMBL207543 | C22H26Cl2N6 | 445.392 | 5 / 0 | 4.3 | Yes |
| 1655 | 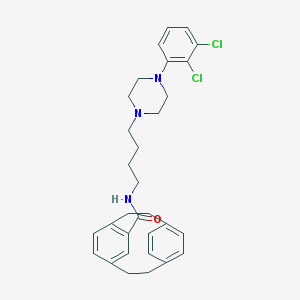 CHEMBL210461 CHEMBL210461 | C31H35Cl2N3O | 536.541 | 3 / 1 | 7.3 | No |
| 1980 | 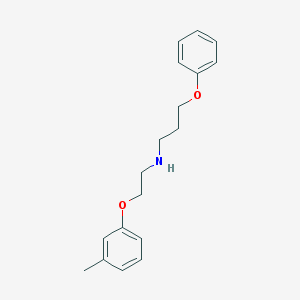 CHEMBL128232 CHEMBL128232 | C18H23NO2 | 285.387 | 3 / 1 | 3.9 | Yes |
| 521507 | 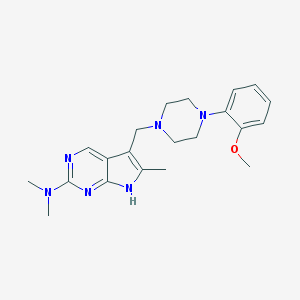 CHEMBL558838 CHEMBL558838 | C21H28N6O | 380.496 | 6 / 1 | 2.9 | Yes |
| 2269 | 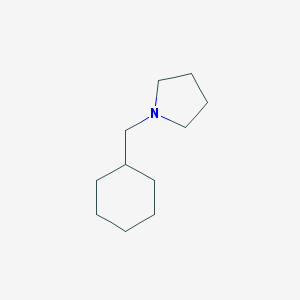 1-(cyclohexylmethyl)pyrrolidine 1-(cyclohexylmethyl)pyrrolidine | C11H21N | 167.296 | 1 / 0 | 3.1 | Yes |
| 521520 | 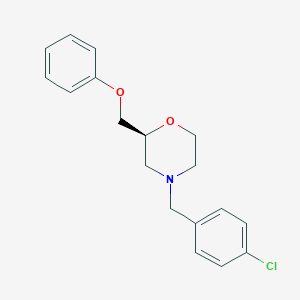 CHEMBL3793112 CHEMBL3793112 | C18H20ClNO2 | 317.813 | 3 / 0 | 3.7 | Yes |
| 2408 | 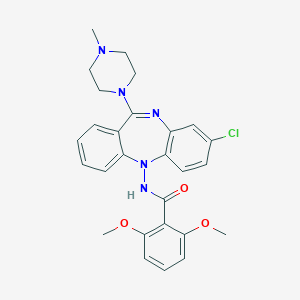 CHEMBL425069 CHEMBL425069 | C27H28ClN5O3 | 506.003 | 6 / 1 | 4.3 | No |
| 463259 | 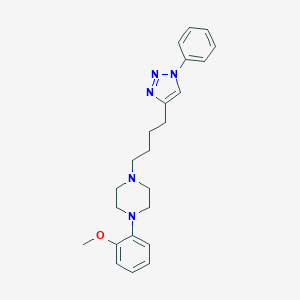 CHEMBL3353895 CHEMBL3353895 | C23H29N5O | 391.519 | 5 / 0 | 4.0 | Yes |
| 521535 | 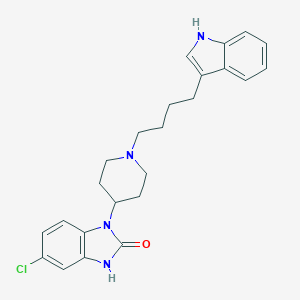 CHEMBL3818805 CHEMBL3818805 | C24H27ClN4O | 422.957 | 2 / 2 | 4.8 | Yes |
| 3351 | 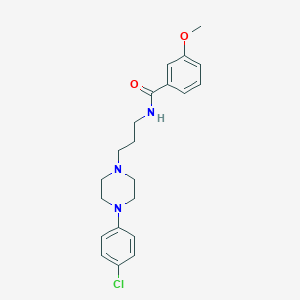 CHEMBL92713 CHEMBL92713 | C21H26ClN3O2 | 387.908 | 4 / 1 | 3.8 | Yes |
| 3466 | 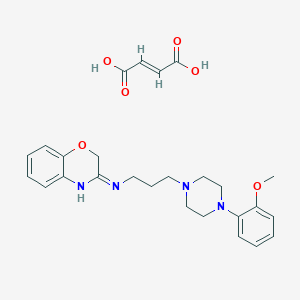 CHEMBL80711 CHEMBL80711 | C26H32N4O6 | 496.564 | 9 / 3 | N/A | N/A |
| 3651 | 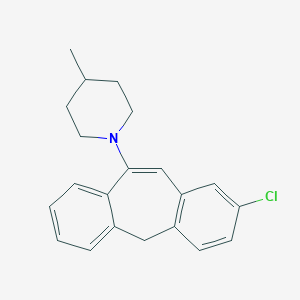 CHEMBL269396 CHEMBL269396 | C21H22ClN | 323.864 | 1 / 0 | 6.1 | No |
| 3795 | 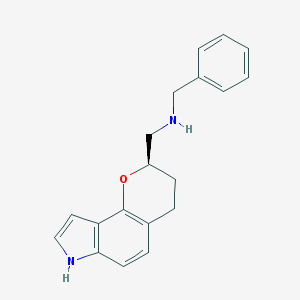 CHEMBL60185 CHEMBL60185 | C19H20N2O | 292.382 | 2 / 2 | 3.5 | Yes |
| 4478 | 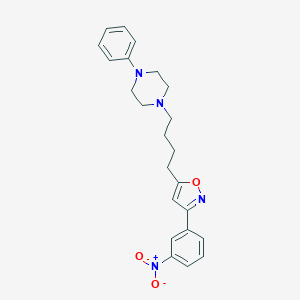 CHEMBL538977 CHEMBL538977 | C23H26N4O3 | 406.486 | 6 / 0 | 4.5 | Yes |
| 521579 | 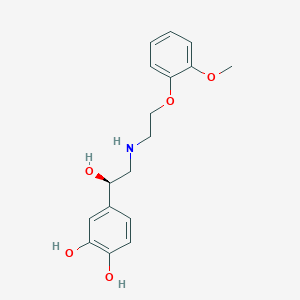 CHEMBL3797876 CHEMBL3797876 | C17H21NO5 | 319.357 | 6 / 4 | 1.6 | Yes |
| 521588 | 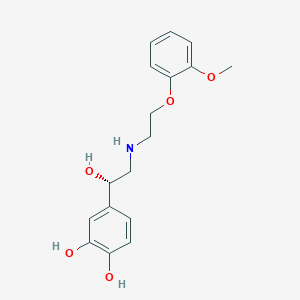 CHEMBL3799593 CHEMBL3799593 | C17H21NO5 | 319.357 | 6 / 4 | 1.6 | Yes |
| 4685 | 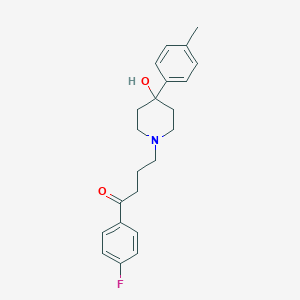 Moperone Moperone | C22H26FNO2 | 355.453 | 4 / 1 | 3.0 | Yes |
| 5030 | 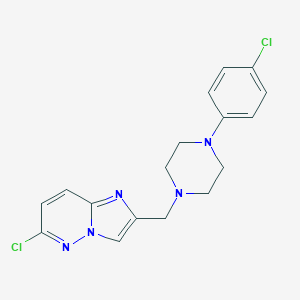 CHEMBL211135 CHEMBL211135 | C17H17Cl2N5 | 362.258 | 4 / 0 | 3.4 | Yes |
| 5236 | 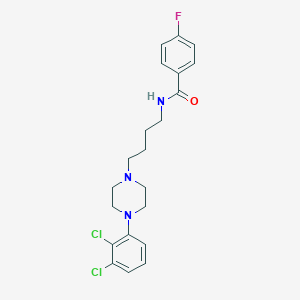 CHEMBL196744 CHEMBL196744 | C21H24Cl2FN3O | 424.341 | 4 / 1 | 4.9 | Yes |
| 521617 | 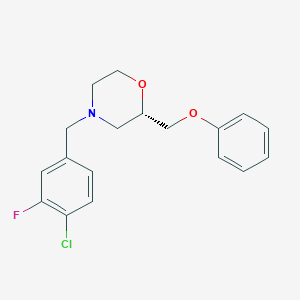 CHEMBL3794174 CHEMBL3794174 | C18H19ClFNO2 | 335.803 | 4 / 0 | 3.8 | Yes |
| 5924 | 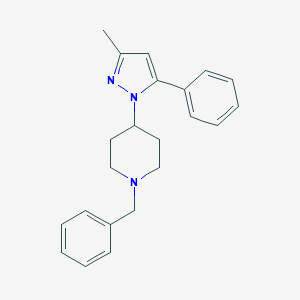 CHEMBL24809 CHEMBL24809 | C22H25N3 | 331.463 | 2 / 0 | 4.3 | Yes |
| 5995 | 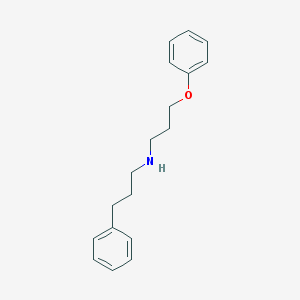 CHEMBL129931 CHEMBL129931 | C18H23NO | 269.388 | 2 / 1 | 4.1 | Yes |
| 6087 | 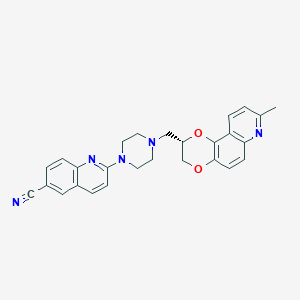 CHEMBL551964 CHEMBL551964 | C27H25N5O2 | 451.53 | 7 / 0 | 4.3 | Yes |
| 6091 | 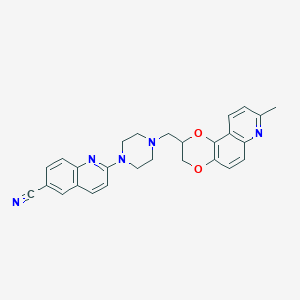 CID 10072510 CID 10072510 | C27H25N5O2 | 451.53 | 7 / 0 | 4.3 | Yes |
| 6262 | 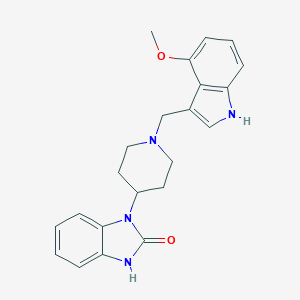 CHEMBL1173436 CHEMBL1173436 | C22H24N4O2 | 376.46 | 3 / 2 | 2.9 | Yes |
| 6271 | 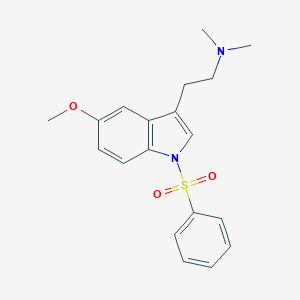 CHEMBL76237 CHEMBL76237 | C19H22N2O3S | 358.456 | 4 / 0 | 3.5 | Yes |
| 6300 | 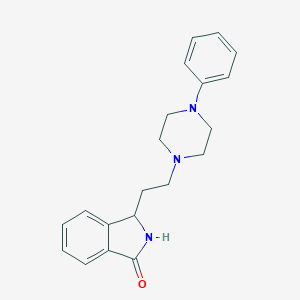 CHEMBL34239 CHEMBL34239 | C20H23N3O | 321.424 | 3 / 1 | 2.8 | Yes |
| 463728 | 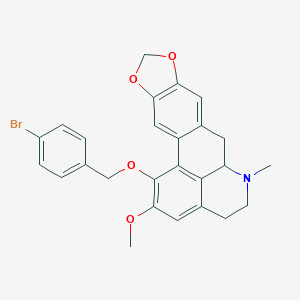 CHEMBL3633757 CHEMBL3633757 | C26H24BrNO4 | 494.385 | 5 / 0 | 5.4 | No |
| 6951 | 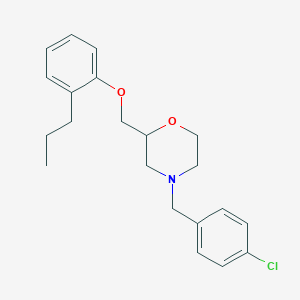 CHEMBL105234 CHEMBL105234 | C21H26ClNO2 | 359.894 | 3 / 0 | 5.1 | No |
| 6963 | 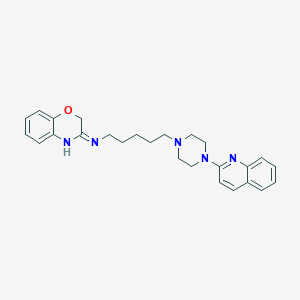 CHEMBL544296 CHEMBL544296 | C26H31N5O | 429.568 | 5 / 1 | 4.3 | Yes |
| 6981 | 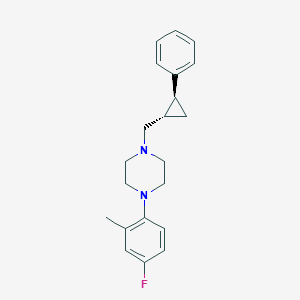 CHEMBL1202209 CHEMBL1202209 | C21H25FN2 | 324.443 | 3 / 0 | 4.5 | Yes |
| 7219 | 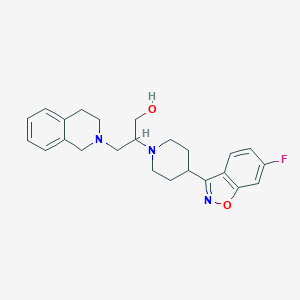 Isoindoline, 26 Isoindoline, 26 | C24H28FN3O2 | 409.505 | 6 / 1 | 3.4 | Yes |
| 7892 | 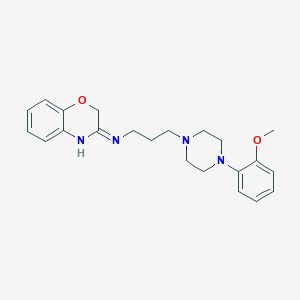 CHEMBL80711 CHEMBL80711 | C22H28N4O2 | 380.492 | 5 / 1 | 2.9 | Yes |
| 8114 | 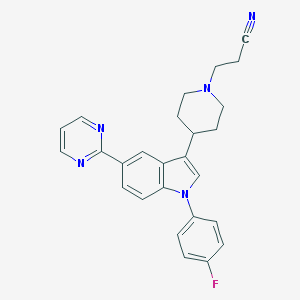 CHEMBL317333 CHEMBL317333 | C26H24FN5 | 425.511 | 5 / 0 | 4.0 | Yes |
| 555515 | 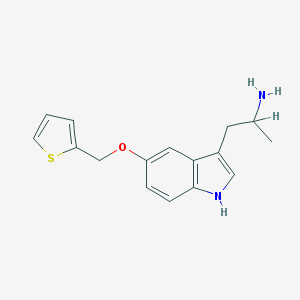 BW-723C86 BW-723C86 | C16H18N2OS | 286.393 | 3 / 2 | 3.0 | Yes |
| 8324 | 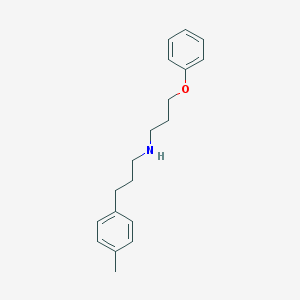 CHEMBL334989 CHEMBL334989 | C19H25NO | 283.415 | 2 / 1 | 4.5 | Yes |
| 8422 | 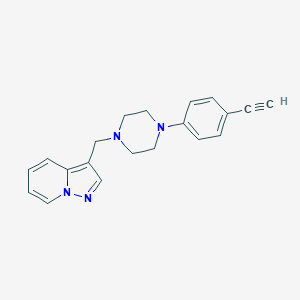 CHEMBL611793 CHEMBL611793 | C20H20N4 | 316.408 | 3 / 0 | 2.7 | Yes |
| 8457 | 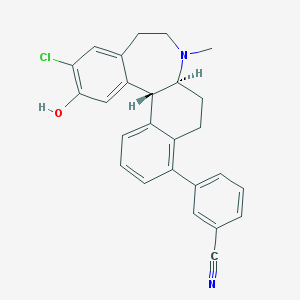 CHEMBL598105 CHEMBL598105 | C26H23ClN2O | 414.933 | 3 / 1 | 5.8 | No |
| 8678 | 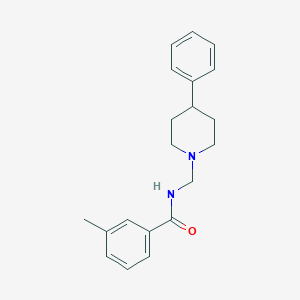 CHEMBL220189 CHEMBL220189 | C20H24N2O | 308.425 | 2 / 1 | 3.9 | Yes |
| 521710 | 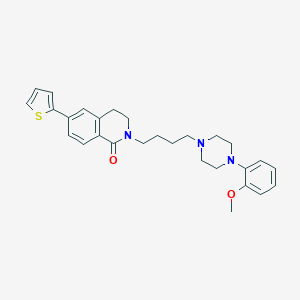 CHEMBL1771110 CHEMBL1771110 | C28H33N3O2S | 475.651 | 5 / 0 | 5.1 | No |
| 521711 | 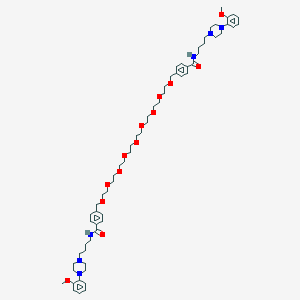 CHEMBL1928132 CHEMBL1928132 | C62H92N6O13 | 1129.45 | 17 / 2 | 5.1 | No |
| 9228 | 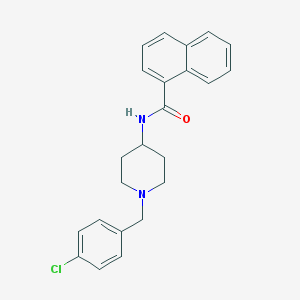 CHEMBL234641 CHEMBL234641 | C23H23ClN2O | 378.9 | 2 / 1 | 5.2 | No |
| 9237 | 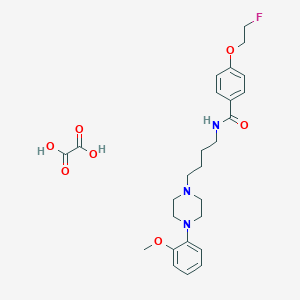 CHEMBL1688992 CHEMBL1688992 | C26H34FN3O7 | 519.57 | 10 / 3 | N/A | No |
| 442082 | 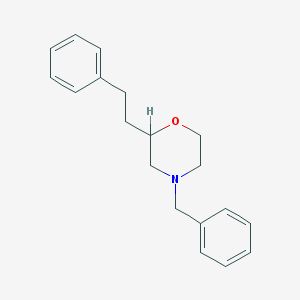 CHEMBL3335544 CHEMBL3335544 | C19H23NO | 281.399 | 2 / 0 | 3.8 | Yes |
| 521720 | 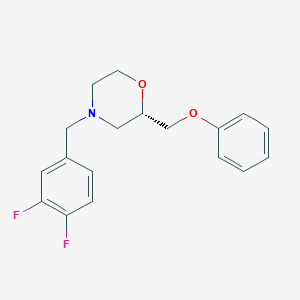 CHEMBL3793062 CHEMBL3793062 | C18H19F2NO2 | 319.352 | 5 / 0 | 3.3 | Yes |
| 521721 | 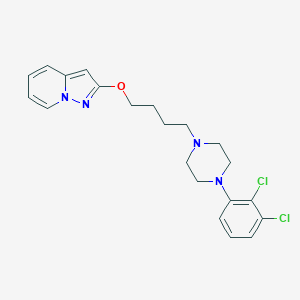 CHEMBL3287394 CHEMBL3287394 | C21H24Cl2N4O | 419.35 | 4 / 0 | 5.0 | Yes |
| 521723 | 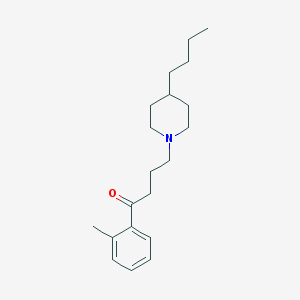 AC-42 AC-42 | C20H31NO | 301.474 | 2 / 0 | 5.2 | No |
| 10182 | 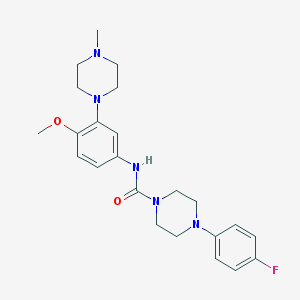 CHEMBL194206 CHEMBL194206 | C23H30FN5O2 | 427.524 | 6 / 1 | 2.7 | Yes |
| 10293 | 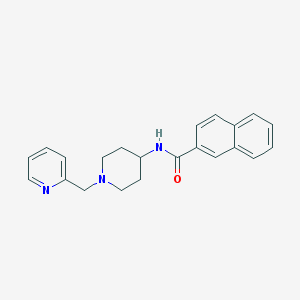 CHEMBL234827 CHEMBL234827 | C22H23N3O | 345.446 | 3 / 1 | 3.5 | Yes |
| 10549 | 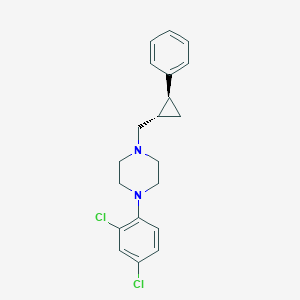 CHEMBL302075 CHEMBL302075 | C20H22Cl2N2 | 361.31 | 2 / 0 | 5.2 | No |
| 10550 | 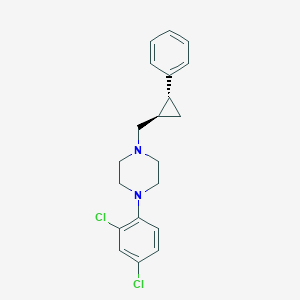 CHEMBL125455 CHEMBL125455 | C20H22Cl2N2 | 361.31 | 2 / 0 | 5.2 | No |
| 10735 | 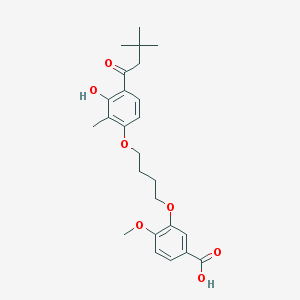 CHEMBL3287723 CHEMBL3287723 | C25H32O7 | 444.524 | 7 / 2 | 5.7 | No |
| 442130 | 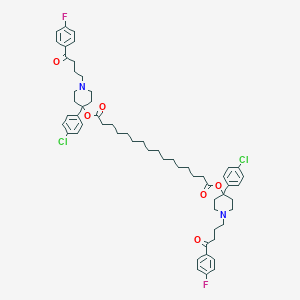 CHEMBL3318840 CHEMBL3318840 | C58H72Cl2F2N2O6 | 1002.12 | 10 / 0 | 14.6 | No |
| 11031 | 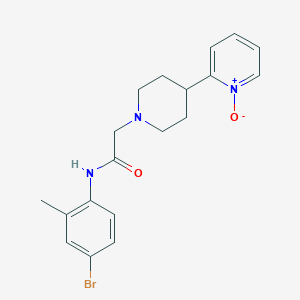 CHEMBL219211 CHEMBL219211 | C19H22BrN3O2 | 404.308 | 3 / 1 | 2.4 | Yes |
| 11078 | 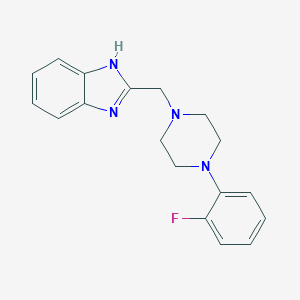 CHEMBL126510 CHEMBL126510 | C18H19FN4 | 310.376 | 4 / 1 | 3.1 | Yes |
| 11184 | 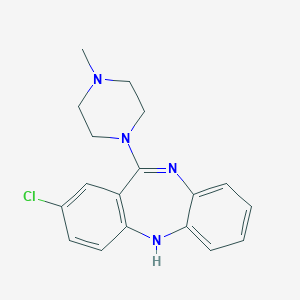 ISOCLOZAPINE ISOCLOZAPINE | C18H19ClN4 | 326.828 | 3 / 1 | 3.1 | Yes |
| 11194 | 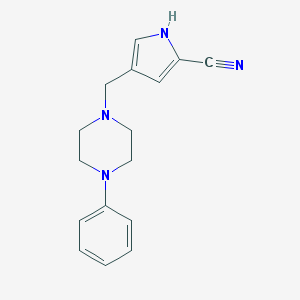 CHEMBL2113718 CHEMBL2113718 | C16H18N4 | 266.348 | 3 / 1 | 2.2 | Yes |
| 11334 | 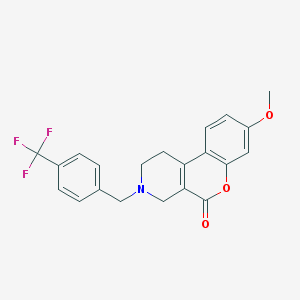 CHEMBL91938 CHEMBL91938 | C21H18F3NO3 | 389.374 | 7 / 0 | 3.7 | Yes |
| 11340 | 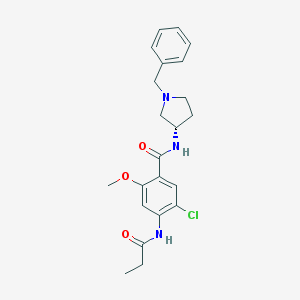 CHEMBL93198 CHEMBL93198 | C22H26ClN3O3 | 415.918 | 4 / 2 | 3.2 | Yes |
| 11555 | 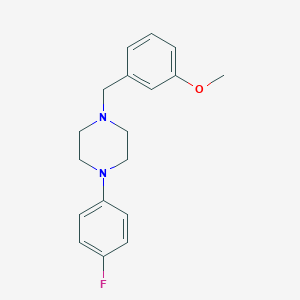 CBDivE_014929 CBDivE_014929 | C18H21FN2O | 300.377 | 4 / 0 | 3.4 | Yes |
| 521791 | 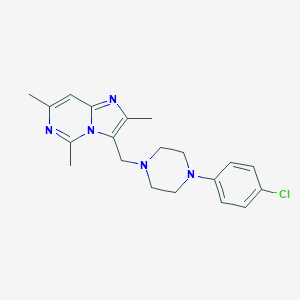 CHEMBL554692 CHEMBL554692 | C20H24ClN5 | 369.897 | 4 / 0 | 4.4 | Yes |
| 11814 | 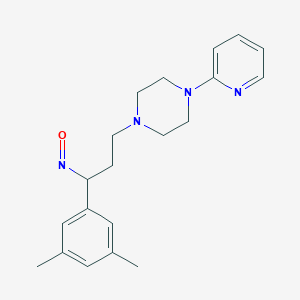 BDBM50193383 BDBM50193383 | C20H26N4O | 338.455 | 5 / 0 | 3.2 | Yes |
| 521820 | 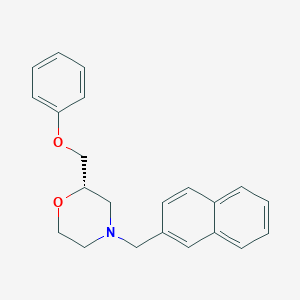 CHEMBL3792556 CHEMBL3792556 | C22H23NO2 | 333.431 | 3 / 0 | 4.4 | Yes |
| 13127 | 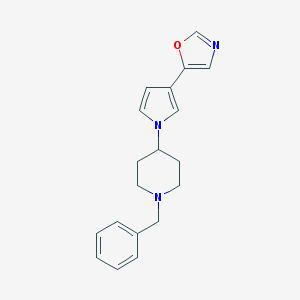 CHEMBL107529 CHEMBL107529 | C19H21N3O | 307.397 | 3 / 0 | 2.7 | Yes |
| 13233 | 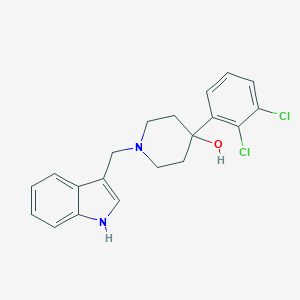 CHEMBL397770 CHEMBL397770 | C20H20Cl2N2O | 375.293 | 2 / 2 | 4.3 | Yes |
| 13350 | 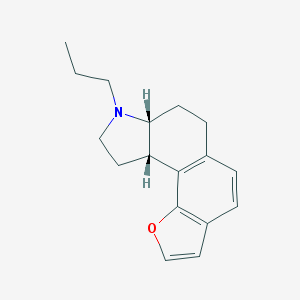 CHEMBL328107 CHEMBL328107 | C17H21NO | 255.361 | 2 / 0 | 3.9 | Yes |
| 13395 | 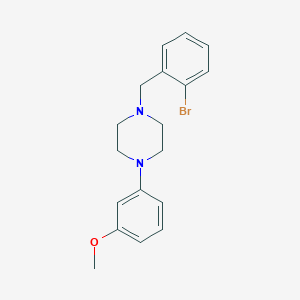 CHEMBL405970 CHEMBL405970 | C18H21BrN2O | 361.283 | 3 / 0 | 4.0 | Yes |
| 555535 | 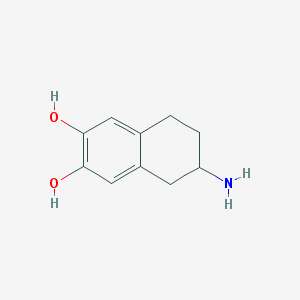 ADTN ADTN | C10H13NO2 | 179.219 | 3 / 3 | 1.4 | Yes |
| 13680 | 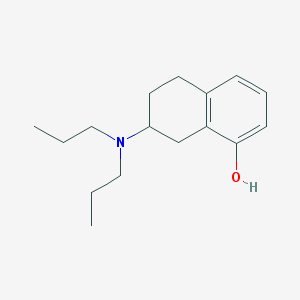 8-OH-Dpat 8-OH-Dpat | C16H25NO | 247.382 | 2 / 1 | 4.1 | Yes |
| 13726 | 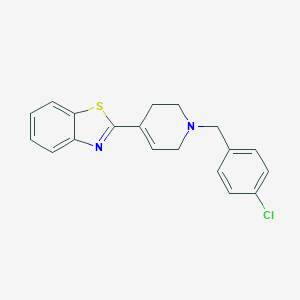 CHEMBL306265 CHEMBL306265 | C19H17ClN2S | 340.869 | 3 / 0 | 4.7 | Yes |
| 13816 | 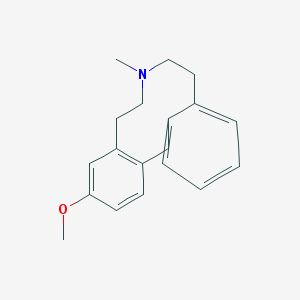 CHEMBL380330 CHEMBL380330 | C19H23NO | 281.399 | 2 / 0 | 4.1 | Yes |
| 464595 | 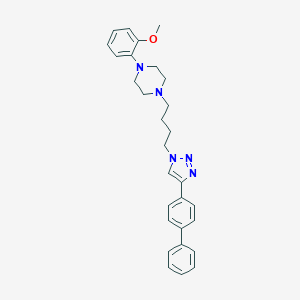 CHEMBL3588983 CHEMBL3588983 | C29H33N5O | 467.617 | 5 / 0 | 5.2 | No |
| 14182 | 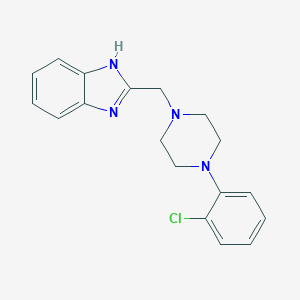 CHEMBL77563 CHEMBL77563 | C18H19ClN4 | 326.828 | 3 / 1 | 3.6 | Yes |
| 14188 | 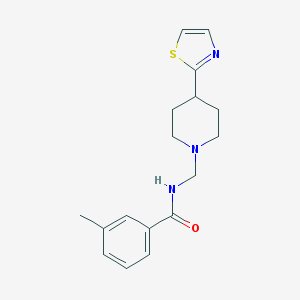 CHEMBL220083 CHEMBL220083 | C17H21N3OS | 315.435 | 4 / 1 | 3.0 | Yes |
| 14485 | 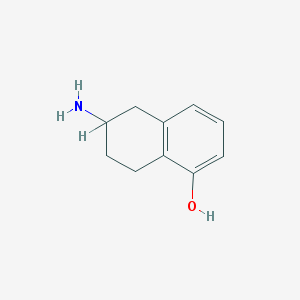 5-Hydroxy-2-aminotetralin 5-Hydroxy-2-aminotetralin | C10H13NO | 163.22 | 2 / 2 | 1.3 | Yes |
| 14497 | 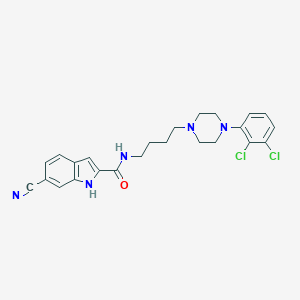 CHEMBL218151 CHEMBL218151 | C24H25Cl2N5O | 470.398 | 4 / 2 | 5.0 | Yes |
| 14518 | 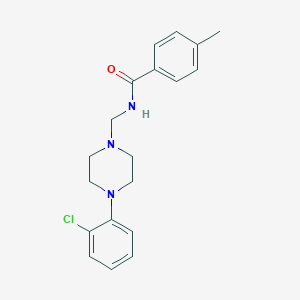 CHEMBL51536 CHEMBL51536 | C19H22ClN3O | 343.855 | 3 / 1 | 3.9 | Yes |
| 521886 | 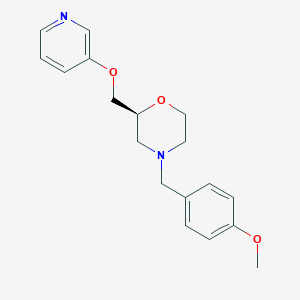 CHEMBL3793264 CHEMBL3793264 | C18H22N2O3 | 314.385 | 5 / 0 | 2.0 | Yes |
| 14728 | 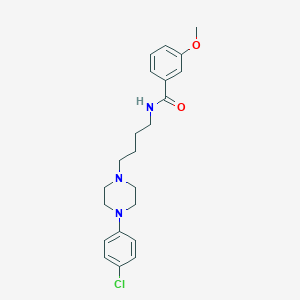 CHEMBL315879 CHEMBL315879 | C22H28ClN3O2 | 401.935 | 4 / 1 | 4.1 | Yes |
| 15000 | 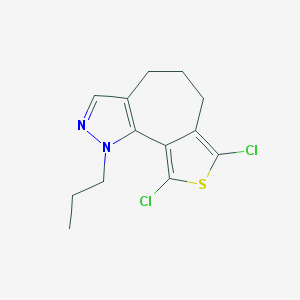 CHEMBL99085 CHEMBL99085 | C13H14Cl2N2S | 301.229 | 2 / 0 | 5.2 | No |
| 15101 | 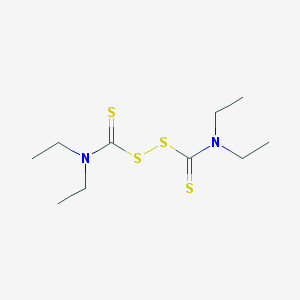 disulfiram disulfiram | C10H20N2S4 | 296.524 | 4 / 0 | 3.9 | Yes |
| 521906 | 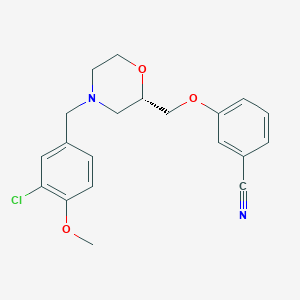 CHEMBL3793131 CHEMBL3793131 | C20H21ClN2O3 | 372.849 | 5 / 0 | 3.4 | Yes |
| 521907 | 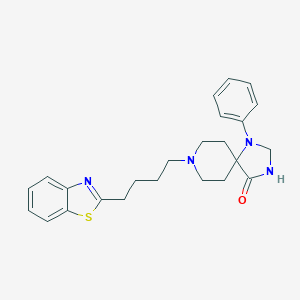 CHEMBL3819567 CHEMBL3819567 | C24H28N4OS | 420.575 | 5 / 1 | 4.8 | Yes |
| 15486 | 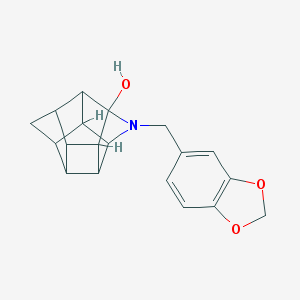 CHEMBL2205829 CHEMBL2205829 | C19H19NO3 | 309.365 | 4 / 1 | 2.2 | Yes |
| 15856 | 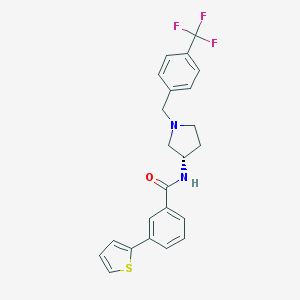 CHEMBL370652 CHEMBL370652 | C23H21F3N2OS | 430.489 | 6 / 1 | 5.2 | No |
| 15951 | 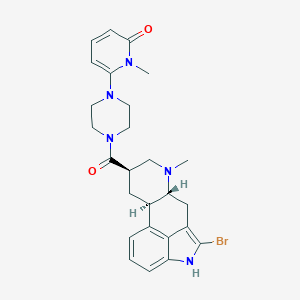 CHEMBL254500 CHEMBL254500 | C26H30BrN5O2 | 524.463 | 4 / 1 | 3.1 | No |
| 16036 | 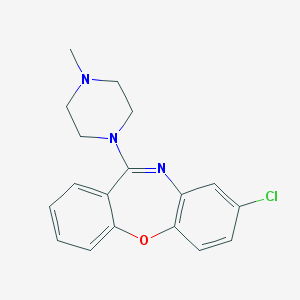 ISOLOXAPINE ISOLOXAPINE | C18H18ClN3O | 327.812 | 3 / 0 | 3.1 | Yes |
| 16114 | 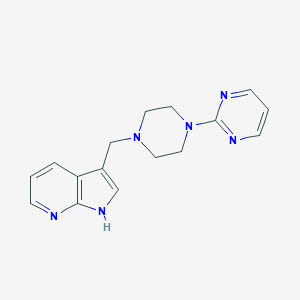 CHEMBL460644 CHEMBL460644 | C16H18N6 | 294.362 | 5 / 1 | 1.4 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218