

You can:
| Name | Endothelin receptor type B |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | EDNRB |
| Synonym | endothelin B receptor HSCR2 HSCR ETB receptor ET-BR [ Show all ] |
| Disease | Arrhythmia Hypertension Pulmonary arterial hypertension Solid tumours Cancer [ Show all ] |
| Length | 442 |
| Amino acid sequence | MQPPPSLCGRALVALVLACGLSRIWGEERGFPPDRATPLLQTAEIMTPPTKTLWPKGSNASLARSLAPAEVPKGDRTAGSPPRTISPPPCQGPIEIKETFKYINTVVSCLVFVLGIIGNSTLLRIIYKNKCMRNGPNILIASLALGDLLHIVIDIPINVYKLLAEDWPFGAEMCKLVPFIQKASVGITVLSLCALSIDRYRAVASWSRIKGIGVPKWTAVEIVLIWVVSVVLAVPEAIGFDIITMDYKGSYLRICLLHPVQKTAFMQFYKTAKDWWLFSFYFCLPLAITAFFYTLMTCEMLRKKSGMQIALNDHLKQRREVAKTVFCLVLVFALCWLPLHLSRILKLTLYNQNDPNRCELLSFLLVLDYIGINMASLNSCINPIALYLVSKRFKNCFKSCLCCWCQSFEEKQSLEEKQSCLKFKANDHGYDNFRSSNKYSSS |
| UniProt | P24530 |
| Protein Data Bank | 6igl, 6igk, 5xpr, 5x93 |
| GPCR-HGmod model | P24530 |
| 3D structure model | This structure is from PDB ID 6igl. |
| BioLiP | BL0388813, BL0433639, BL0433638, BL0388896, BL0388814 |
| Therapeutic Target Database | T92828 |
| ChEMBL | CHEMBL1785 |
| IUPHAR | 220 |
| DrugBank | BE0000043 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 267 | 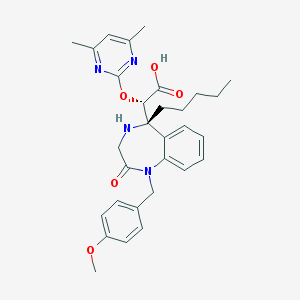 CHEMBL329744 CHEMBL329744 | C30H36N4O5 | 532.641 | 8 / 2 | 2.5 | No |
| 440 | 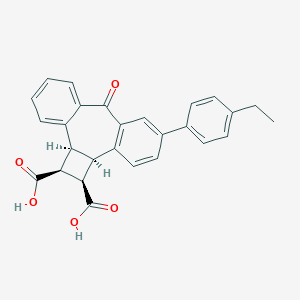 CHEMBL31662 CHEMBL31662 | C27H22O5 | 426.468 | 5 / 2 | 4.5 | Yes |
| 1075 | 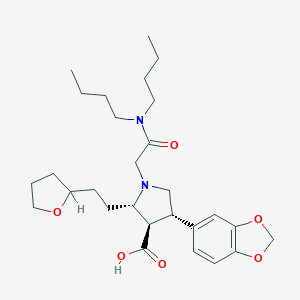 CHEMBL321831 CHEMBL321831 | C28H42N2O6 | 502.652 | 7 / 1 | 2.1 | No |
| 1161 | 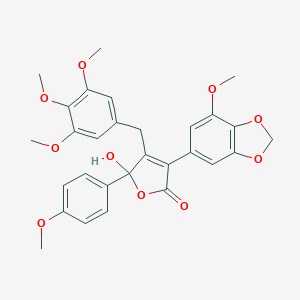 CHEMBL10781 CHEMBL10781 | C29H28O10 | 536.533 | 10 / 1 | 3.9 | No |
| 1702 | 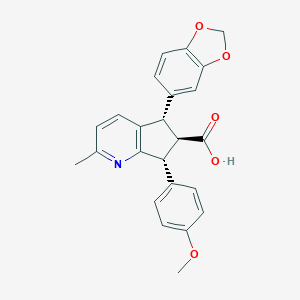 CHEMBL103001 CHEMBL103001 | C24H21NO5 | 403.434 | 6 / 1 | 3.8 | Yes |
| 1745 | 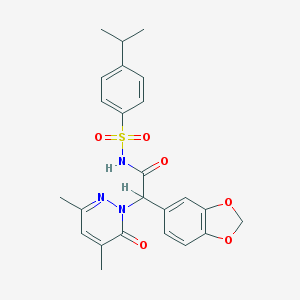 CHEMBL94918 CHEMBL94918 | C24H25N3O6S | 483.539 | 7 / 1 | 3.3 | Yes |
| 2388 | 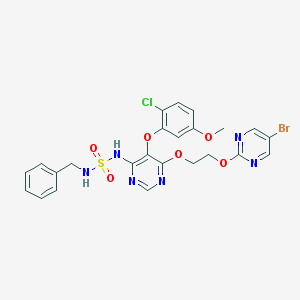 CHEMBL2165330 CHEMBL2165330 | C24H22BrClN6O6S | 637.89 | 12 / 2 | 4.1 | No |
| 2932 | 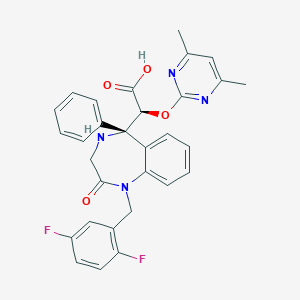 CHEMBL94438 CHEMBL94438 | C30H26F2N4O4 | 544.559 | 9 / 2 | 2.1 | No |
| 3291 | 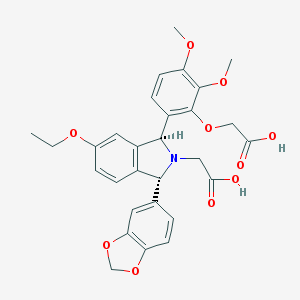 CHEMBL47868 CHEMBL47868 | C29H29NO10 | 551.548 | 11 / 2 | 1.7 | No |
| 3293 | 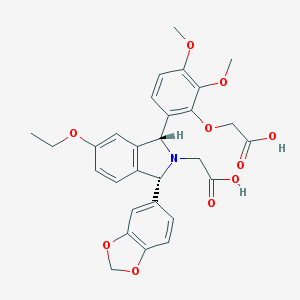 CHEMBL298664 CHEMBL298664 | C29H29NO10 | 551.548 | 11 / 2 | 1.7 | No |
| 3364 | 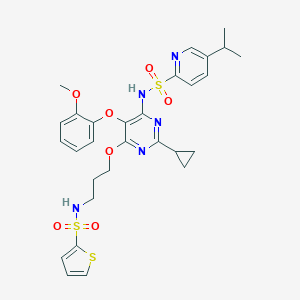 CHEMBL367444 CHEMBL367444 | C29H33N5O7S3 | 659.791 | 13 / 2 | 4.9 | No |
| 3374 | 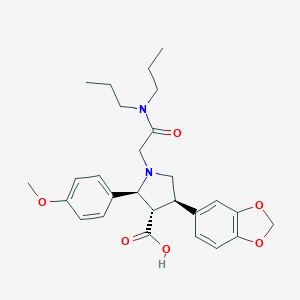 CHEMBL9139 CHEMBL9139 | C27H34N2O6 | 482.577 | 7 / 1 | 1.7 | Yes |
| 4114 | 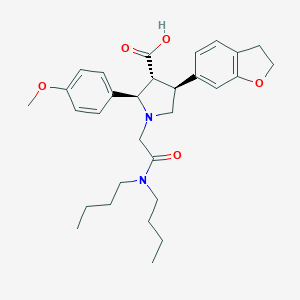 CHEMBL326191 CHEMBL326191 | C30H40N2O5 | 508.659 | 6 / 1 | 2.6 | No |
| 4181 | 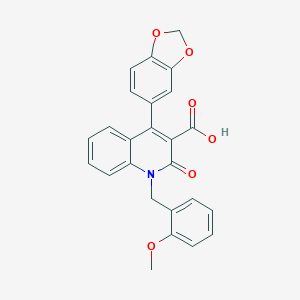 CHEMBL55853 CHEMBL55853 | C25H19NO6 | 429.428 | 6 / 1 | 4.3 | Yes |
| 4670 | 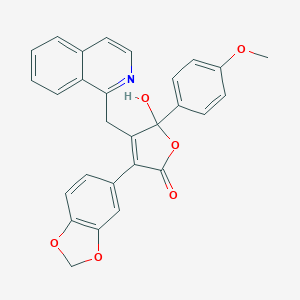 CHEMBL10860 CHEMBL10860 | C28H21NO6 | 467.477 | 7 / 1 | 4.3 | Yes |
| 5291 | 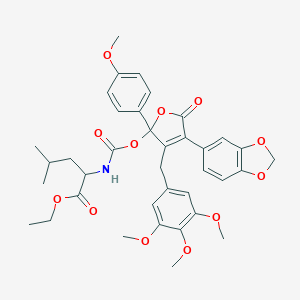 CHEMBL316523 CHEMBL316523 | C37H41NO12 | 691.73 | 12 / 1 | 6.4 | No |
| 5831 | 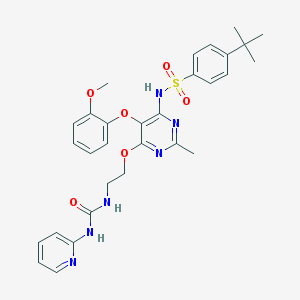 CHEMBL70924 CHEMBL70924 | C30H34N6O6S | 606.698 | 10 / 3 | 4.9 | No |
| 6224 | 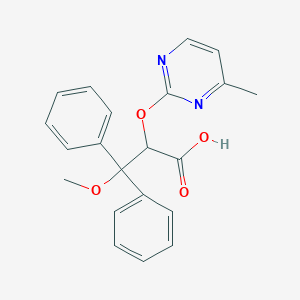 CHEMBL306218 CHEMBL306218 | C21H20N2O4 | 364.401 | 6 / 1 | 3.4 | Yes |
| 6281 | 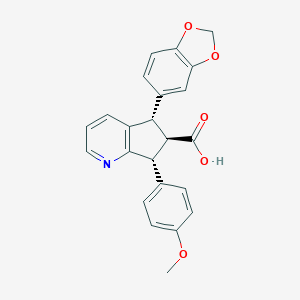 CHEMBL102825 CHEMBL102825 | C23H19NO5 | 389.407 | 6 / 1 | 3.4 | Yes |
| 6513 | 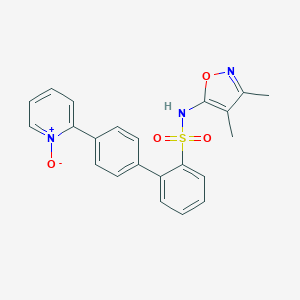 CHEMBL147238 CHEMBL147238 | C22H19N3O4S | 421.471 | 6 / 1 | 3.2 | Yes |
| 6579 | 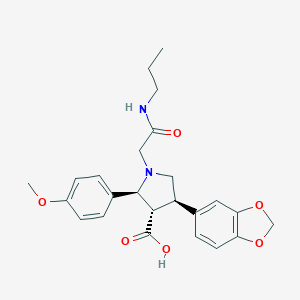 CHEMBL9141 CHEMBL9141 | C24H28N2O6 | 440.496 | 7 / 2 | 0.6 | Yes |
| 7105 | 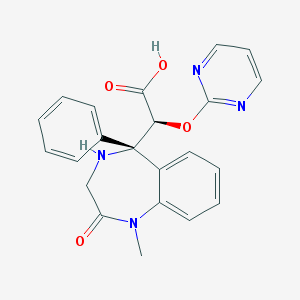 CHEMBL92612 CHEMBL92612 | C22H20N4O4 | 404.426 | 7 / 2 | -0.4 | Yes |
| 7556 | 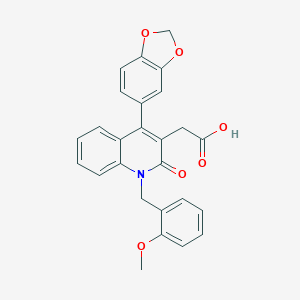 CHEMBL56349 CHEMBL56349 | C26H21NO6 | 443.455 | 6 / 1 | 3.4 | Yes |
| 8439 | 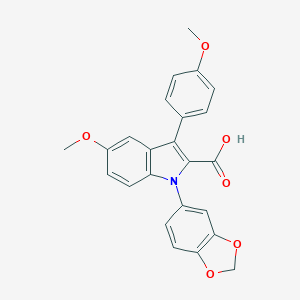 CHEMBL274370 CHEMBL274370 | C24H19NO6 | 417.417 | 6 / 1 | 4.9 | Yes |
| 8860 | 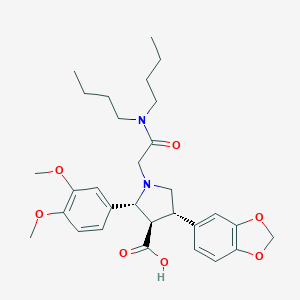 CHEMBL129220 CHEMBL129220 | C30H40N2O7 | 540.657 | 8 / 1 | 2.4 | No |
| 8909 | 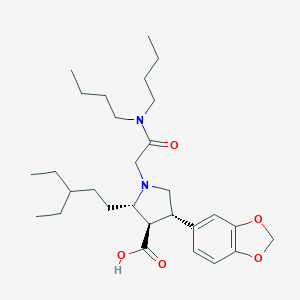 CHEMBL112770 CHEMBL112770 | C29H46N2O5 | 502.696 | 6 / 1 | 4.1 | No |
| 10368 | 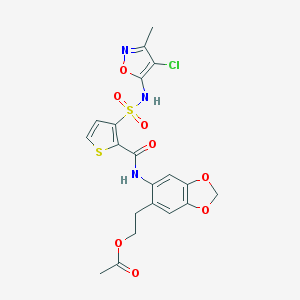 CHEMBL296927 CHEMBL296927 | C20H18ClN3O8S2 | 527.947 | 11 / 2 | 3.1 | No |
| 10419 | 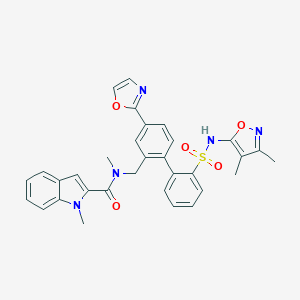 CHEMBL281977 CHEMBL281977 | C32H29N5O5S | 595.674 | 8 / 1 | 5.1 | No |
| 10754 | 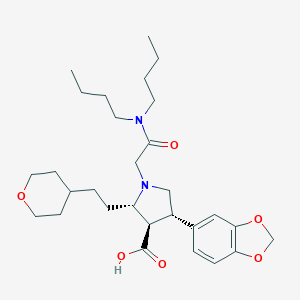 CHEMBL109474 CHEMBL109474 | C29H44N2O6 | 516.679 | 7 / 1 | 2.3 | No |
| 11424 | 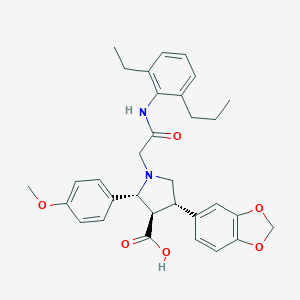 CHEMBL123997 CHEMBL123997 | C32H36N2O6 | 544.648 | 7 / 2 | 3.4 | No |
| 11625 | 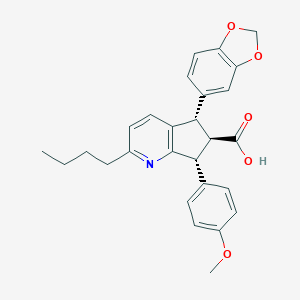 CHEMBL102969 CHEMBL102969 | C27H27NO5 | 445.515 | 6 / 1 | 5.1 | No |
| 12288 | 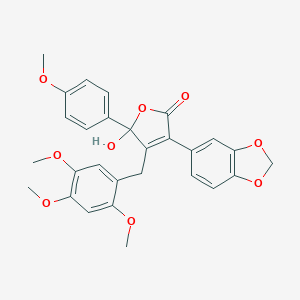 CHEMBL275701 CHEMBL275701 | C28H26O9 | 506.507 | 9 / 1 | 4.0 | No |
| 12373 | 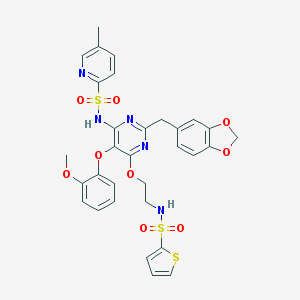 CHEMBL368945 CHEMBL368945 | C31H29N5O9S3 | 711.779 | 15 / 2 | 4.9 | No |
| 12416 | 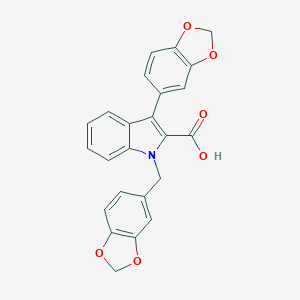 CHEMBL416518 CHEMBL416518 | C24H17NO6 | 415.401 | 6 / 1 | 4.7 | Yes |
| 12419 | 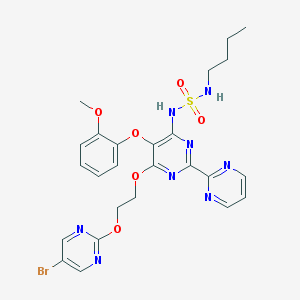 CHEMBL2163720 CHEMBL2163720 | C25H27BrN8O6S | 647.505 | 14 / 2 | 3.2 | No |
| 12715 | 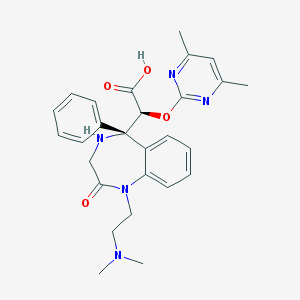 CHEMBL450628 CHEMBL450628 | C27H31N5O4 | 489.576 | 8 / 2 | 0.4 | Yes |
| 13540 | 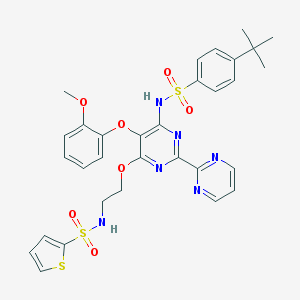 CHEMBL172030 CHEMBL172030 | C31H32N6O7S3 | 696.812 | 14 / 2 | 5.2 | No |
| 13717 | 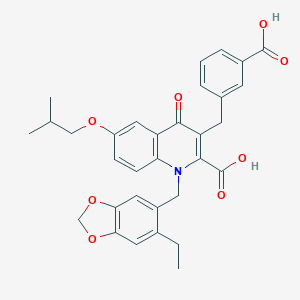 CHEMBL1271640 CHEMBL1271640 | C32H31NO8 | 557.599 | 9 / 2 | 6.3 | No |
| 14947 | 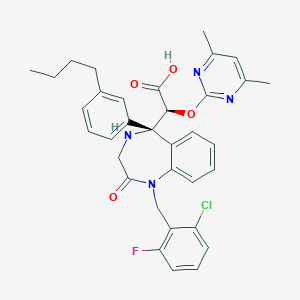 CHEMBL330135 CHEMBL330135 | C34H34ClFN4O4 | 617.118 | 8 / 2 | 4.5 | No |
| 15619 | 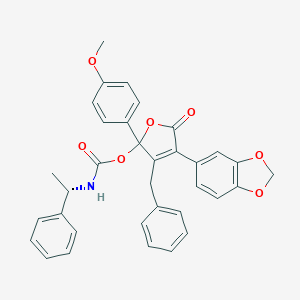 CHEMBL103501 CHEMBL103501 | C34H29NO7 | 563.606 | 7 / 1 | 6.3 | No |
| 15656 | 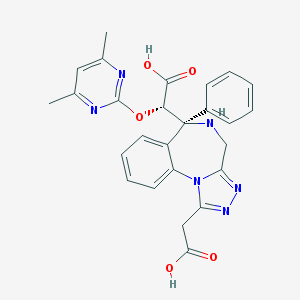 SCHEMBL5351769 SCHEMBL5351769 | C26H24N6O5 | 500.515 | 10 / 3 | -0.5 | No |
| 15769 | 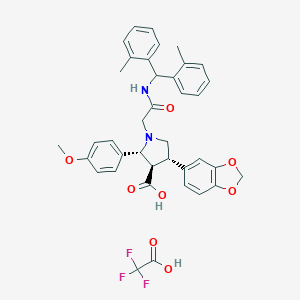 CHEMBL332109 CHEMBL332109 | C38H37F3N2O8 | 706.715 | 12 / 3 | N/A | No |
| 15861 | 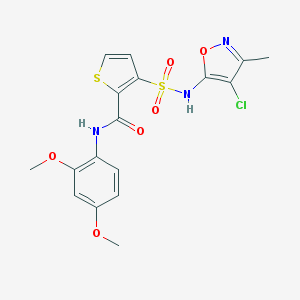 CHEMBL43788 CHEMBL43788 | C17H16ClN3O6S2 | 457.9 | 9 / 2 | 3.0 | Yes |
| 16366 | 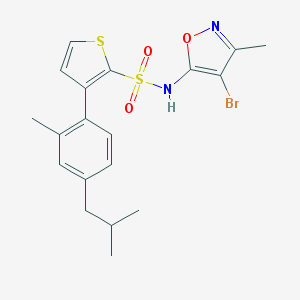 CHEMBL432815 CHEMBL432815 | C19H21BrN2O3S2 | 469.412 | 6 / 1 | 5.9 | No |
| 16661 | 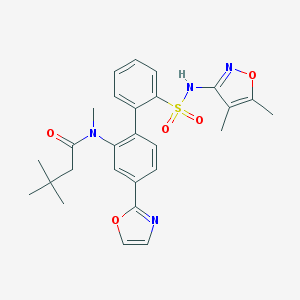 CHEMBL274489 CHEMBL274489 | C27H30N4O5S | 522.62 | 8 / 1 | 4.7 | No |
| 17104 | 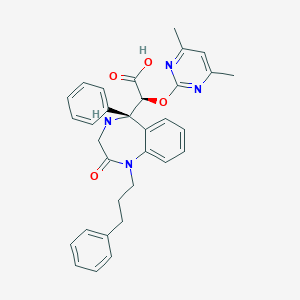 CHEMBL96114 CHEMBL96114 | C32H32N4O4 | 536.632 | 7 / 2 | 2.7 | No |
| 17152 | 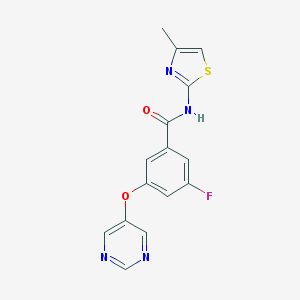 CHEMBL2440659 CHEMBL2440659 | C15H11FN4O2S | 330.337 | 7 / 1 | 2.4 | Yes |
| 17659 | 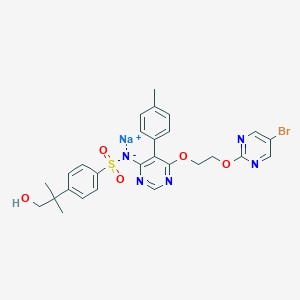 CHEMBL1627022 CHEMBL1627022 | C27H27BrN5NaO5S | 636.497 | 10 / 1 | N/A | No |
| 18581 | 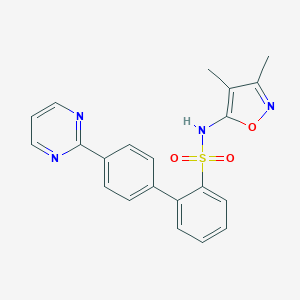 CHEMBL359273 CHEMBL359273 | C21H18N4O3S | 406.46 | 7 / 1 | 3.6 | Yes |
| 19174 | 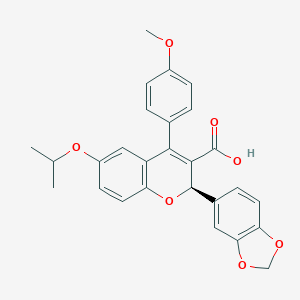 CHEMBL61211 CHEMBL61211 | C27H24O7 | 460.482 | 7 / 1 | 4.9 | Yes |
| 19765 | 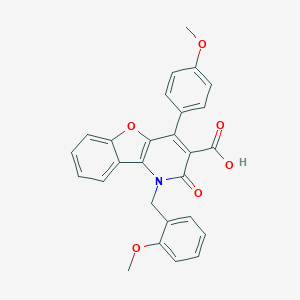 CHEMBL157742 CHEMBL157742 | C27H21NO6 | 455.466 | 6 / 1 | 4.9 | Yes |
| 20154 | 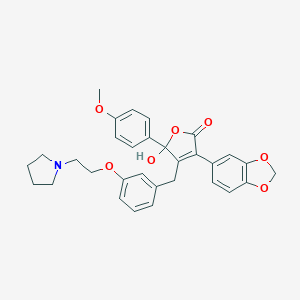 CHEMBL65939 CHEMBL65939 | C31H31NO7 | 529.589 | 8 / 1 | 4.5 | No |
| 20387 | 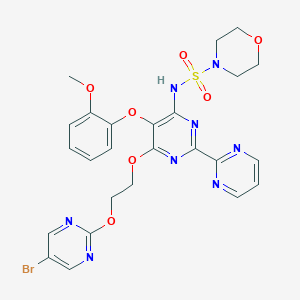 CHEMBL2163722 CHEMBL2163722 | C25H25BrN8O7S | 661.488 | 15 / 1 | 1.8 | No |
| 20391 | 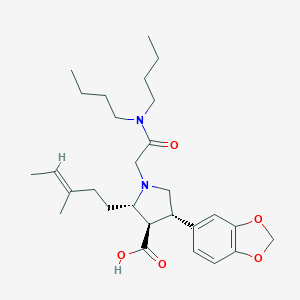 CHEMBL110904 CHEMBL110904 | C28H42N2O5 | 486.653 | 6 / 1 | 3.3 | Yes |
| 20441 | 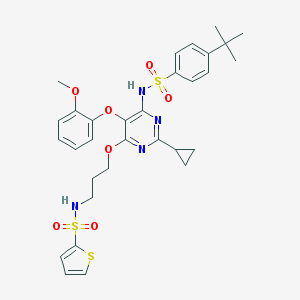 CHEMBL172297 CHEMBL172297 | C31H36N4O7S3 | 672.83 | 12 / 2 | 6.1 | No |
| 20893 | 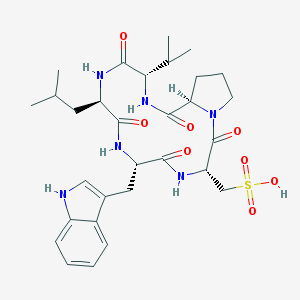 CHEMBL342615 CHEMBL342615 | C30H42N6O8S | 646.76 | 8 / 6 | 1.6 | No |
| 20905 | 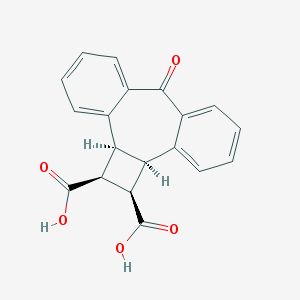 CHEMBL286808 CHEMBL286808 | C19H14O5 | 322.316 | 5 / 2 | 2.1 | Yes |
| 21072 | 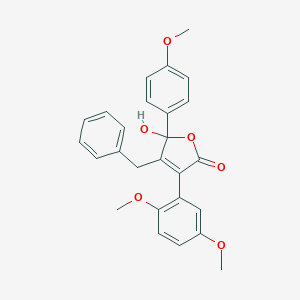 CHEMBL267096 CHEMBL267096 | C26H24O6 | 432.472 | 6 / 1 | 4.2 | Yes |
| 21536 | 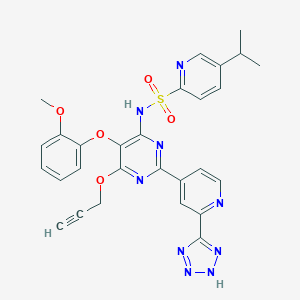 CHEMBL175057 CHEMBL175057 | C28H25N9O5S | 599.626 | 13 / 2 | 3.2 | No |
| 21766 | 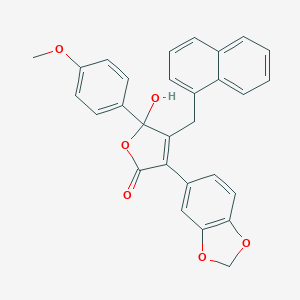 CHEMBL269409 CHEMBL269409 | C29H22O6 | 466.489 | 6 / 1 | 5.3 | No |
| 522155 | 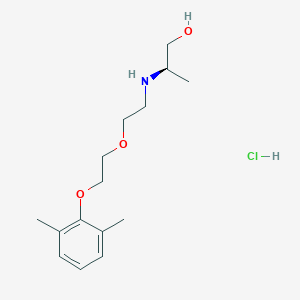 CHEMBL3775436 CHEMBL3775436 | C15H26ClNO3 | 303.827 | 4 / 3 | N/A | N/A |
| 522202 | 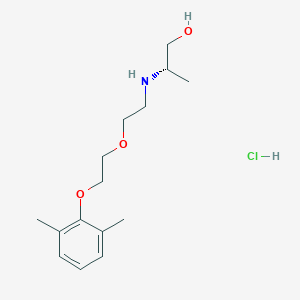 CHEMBL3775769 CHEMBL3775769 | C15H26ClNO3 | 303.827 | 4 / 3 | N/A | N/A |
| 22295 | 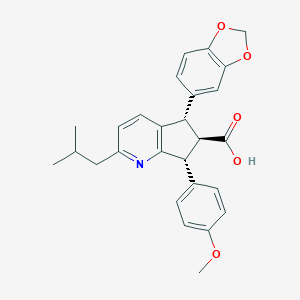 CHEMBL102314 CHEMBL102314 | C27H27NO5 | 445.515 | 6 / 1 | 5.0 | Yes |
| 22328 | 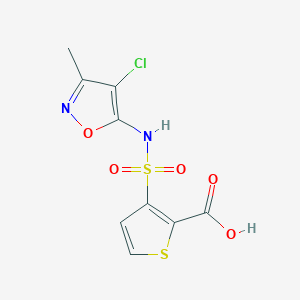 184040-74-2 184040-74-2 | C9H7ClN2O5S2 | 322.734 | 8 / 2 | 1.8 | Yes |
| 22722 | 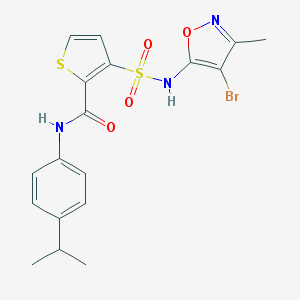 CHEMBL47417 CHEMBL47417 | C18H18BrN3O4S2 | 484.383 | 7 / 2 | 4.3 | Yes |
| 23457 | 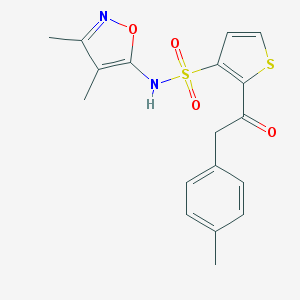 CHEMBL45875 CHEMBL45875 | C18H18N2O4S2 | 390.472 | 7 / 1 | 3.6 | Yes |
| 24317 | 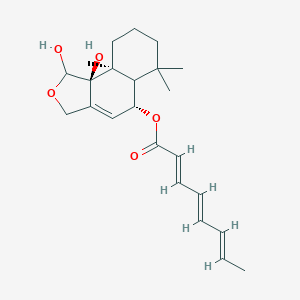 CHEMBL170335 CHEMBL170335 | C23H32O5 | 388.504 | 5 / 2 | 3.4 | Yes |
| 24457 | 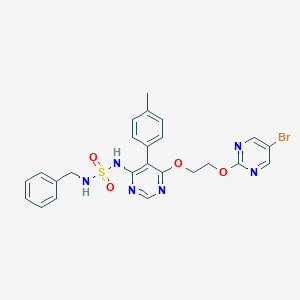 CHEMBL2165332 CHEMBL2165332 | C24H23BrN6O4S | 571.45 | 10 / 2 | 4.0 | No |
| 24823 | 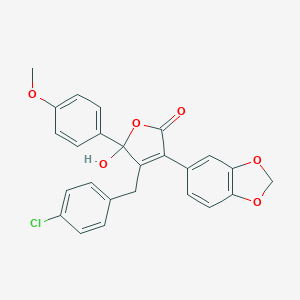 CHEMBL10733 CHEMBL10733 | C25H19ClO6 | 450.871 | 6 / 1 | 4.7 | Yes |
| 25210 | 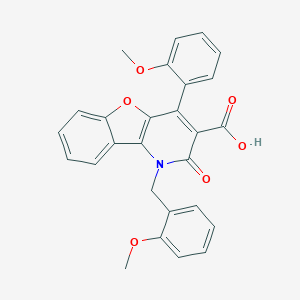 CHEMBL156269 CHEMBL156269 | C27H21NO6 | 455.466 | 6 / 1 | 4.9 | Yes |
| 25432 | 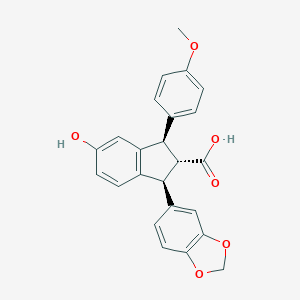 CHEMBL41087 CHEMBL41087 | C24H20O6 | 404.418 | 6 / 2 | 4.1 | Yes |
| 25452 | 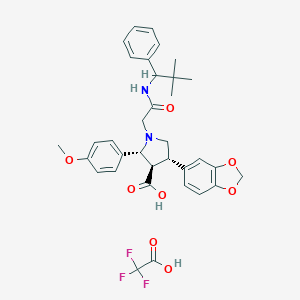 CHEMBL123316 CHEMBL123316 | C34H37F3N2O8 | 658.671 | 12 / 3 | N/A | No |
| 25766 | 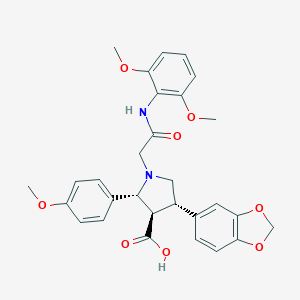 CHEMBL122832 CHEMBL122832 | C29H30N2O8 | 534.565 | 9 / 2 | 1.2 | No |
| 25974 | 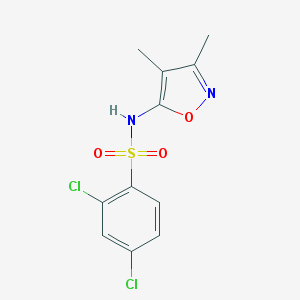 CHEMBL79054 CHEMBL79054 | C11H10Cl2N2O3S | 321.172 | 5 / 1 | 2.9 | Yes |
| 26175 | 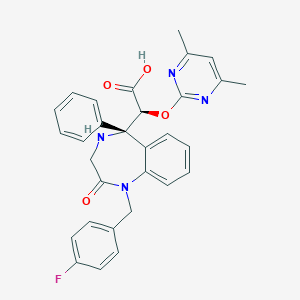 CHEMBL314755 CHEMBL314755 | C30H27FN4O4 | 526.568 | 8 / 2 | 2.0 | No |
| 26957 | 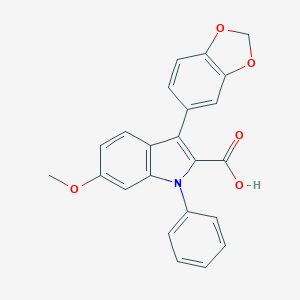 CHEMBL276615 CHEMBL276615 | C23H17NO5 | 387.391 | 5 / 1 | 4.9 | Yes |
| 27177 | 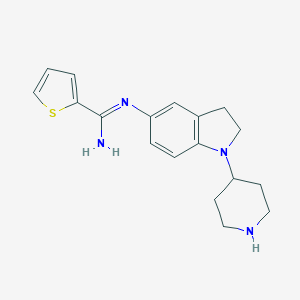 CHEMBL2203713 CHEMBL2203713 | C18H22N4S | 326.462 | 4 / 2 | 2.9 | Yes |
| 27640 | 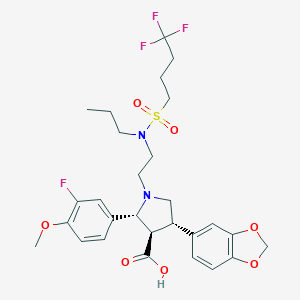 CHEMBL419367 CHEMBL419367 | C28H34F4N2O7S | 618.641 | 13 / 1 | 2.5 | No |
| 27713 | 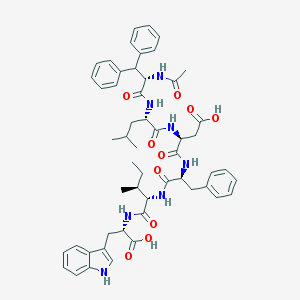 CHEMBL265116 CHEMBL265116 | C53H63N7O10 | 958.126 | 10 / 9 | 6.4 | No |
| 28769 | 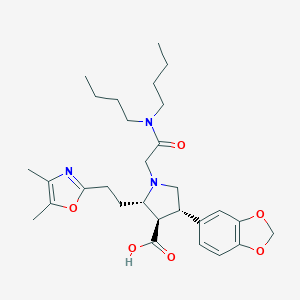 CHEMBL114074 CHEMBL114074 | C29H41N3O6 | 527.662 | 8 / 1 | 2.5 | No |
| 29036 | 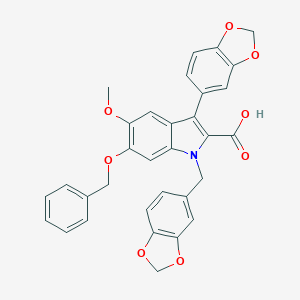 PD-159020 PD-159020 | C32H25NO8 | 551.551 | 8 / 1 | 6.1 | No |
| 29307 | 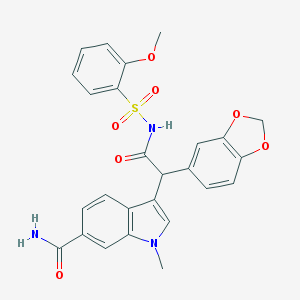 CHEMBL22367 CHEMBL22367 | C26H23N3O7S | 521.544 | 7 / 2 | 2.7 | No |
| 29555 | 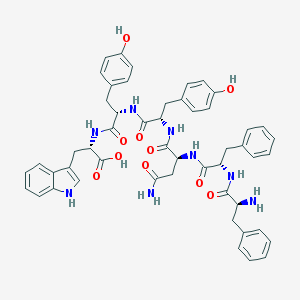 CHEMBL264253 CHEMBL264253 | C51H54N8O10 | 939.039 | 11 / 11 | 0.8 | No |
| 29890 | 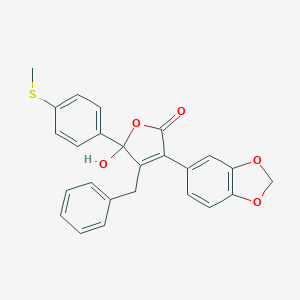 CHEMBL10911 CHEMBL10911 | C25H20O5S | 432.49 | 6 / 1 | 4.6 | Yes |
| 30235 | 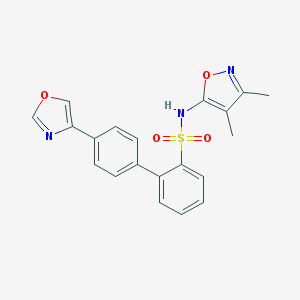 CHEMBL104848 CHEMBL104848 | C20H17N3O4S | 395.433 | 7 / 1 | 3.7 | Yes |
| 30259 | 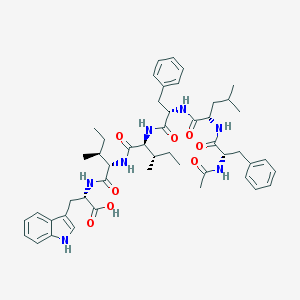 CHEMBL421190 CHEMBL421190 | C49H65N7O8 | 880.1 | 8 / 8 | 5.8 | No |
| 30319 | 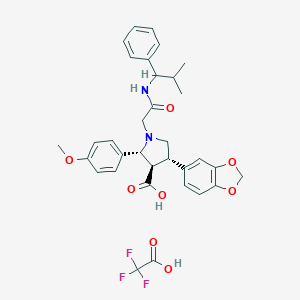 CHEMBL121647 CHEMBL121647 | C33H35F3N2O8 | 644.644 | 12 / 3 | N/A | No |
| 31106 | 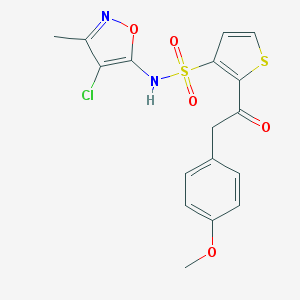 CHEMBL432109 CHEMBL432109 | C17H15ClN2O5S2 | 426.886 | 8 / 1 | 3.5 | Yes |
| 31305 | 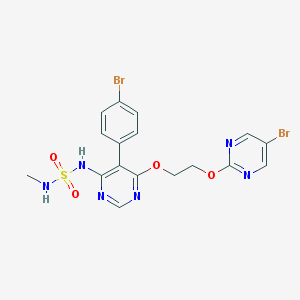 CHEMBL2165337 CHEMBL2165337 | C17H16Br2N6O4S | 560.221 | 10 / 2 | 2.8 | No |
| 31312 | 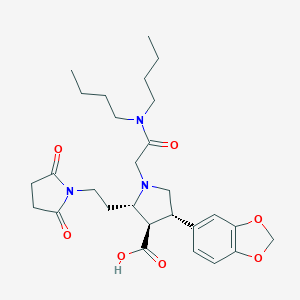 CHEMBL322799 CHEMBL322799 | C28H39N3O7 | 529.634 | 8 / 1 | 0.2 | No |
| 31766 | 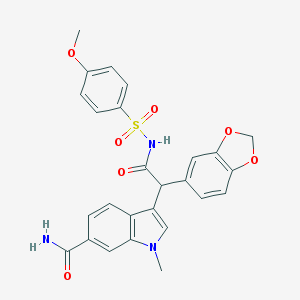 CHEMBL280379 CHEMBL280379 | C26H23N3O7S | 521.544 | 7 / 2 | 2.7 | No |
| 31790 | 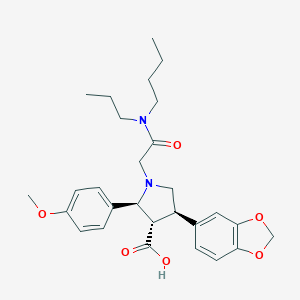 CHEMBL9334 CHEMBL9334 | C28H36N2O6 | 496.604 | 7 / 1 | 2.0 | Yes |
| 32022 | 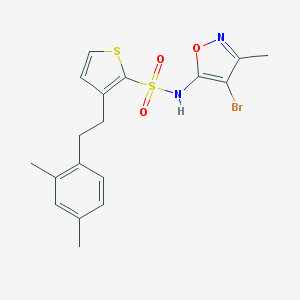 CHEMBL173189 CHEMBL173189 | C18H19BrN2O3S2 | 455.385 | 6 / 1 | 5.3 | No |
| 32027 | 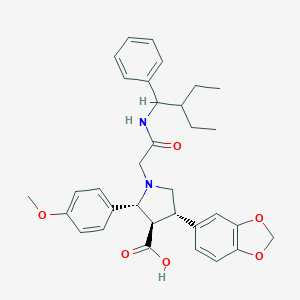 CHEMBL330895 CHEMBL330895 | C33H38N2O6 | 558.675 | 7 / 2 | 3.3 | No |
| 32175 | 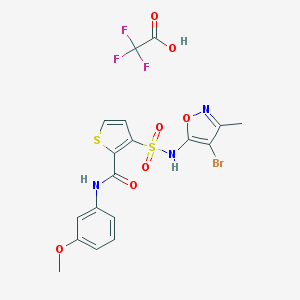 CHEMBL288967 CHEMBL288967 | C18H15BrF3N3O7S2 | 586.351 | 13 / 3 | N/A | No |
| 32537 | 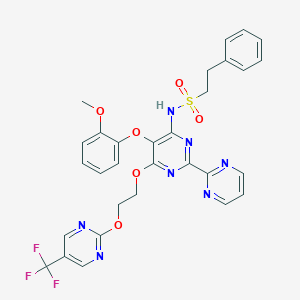 CHEMBL2163705 CHEMBL2163705 | C30H26F3N7O6S | 669.636 | 16 / 1 | 4.4 | No |
| 32950 | 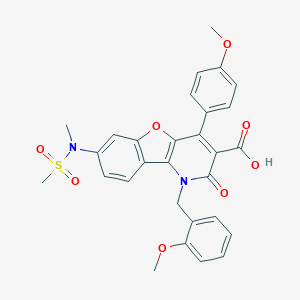 CHEMBL155708 CHEMBL155708 | C29H26N2O8S | 562.593 | 9 / 1 | 4.0 | No |
| 33272 | 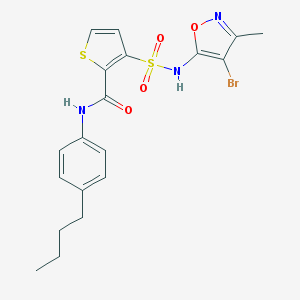 CHEMBL45760 CHEMBL45760 | C19H20BrN3O4S2 | 498.41 | 7 / 2 | 5.0 | Yes |
| 33531 | 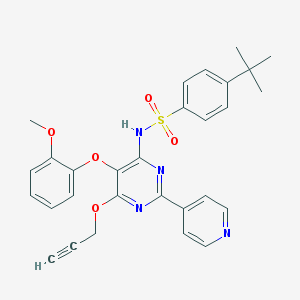 CHEMBL174094 CHEMBL174094 | C29H28N4O5S | 544.626 | 9 / 1 | 5.2 | No |
| 33550 | 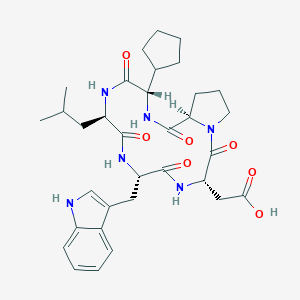 CHEMBL133257 CHEMBL133257 | C33H44N6O7 | 636.75 | 7 / 6 | 2.7 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218