

You can:
| Name | Endothelin-1 receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | EDNRA |
| Synonym | ET-A ETA-R hET-AR ETA receptor ENDOR [ Show all ] |
| Disease | Vasospasm following subarachnoid hemorrhage Hormone refractory prostate cancer Hormone resistant prostate cancer Hypertension Hypotension [ Show all ] |
| Length | 427 |
| Amino acid sequence | METLCLRASFWLALVGCVISDNPERYSTNLSNHVDDFTTFRGTELSFLVTTHQPTNLVLPSNGSMHNYCPQQTKITSAFKYINTVISCTIFIVGMVGNATLLRIIYQNKCMRNGPNALIASLALGDLIYVVIDLPINVFKLLAGRWPFDHNDFGVFLCKLFPFLQKSSVGITVLNLCALSVDRYRAVASWSRVQGIGIPLVTAIEIVSIWILSFILAIPEAIGFVMVPFEYRGEQHKTCMLNATSKFMEFYQDVKDWWLFGFYFCMPLVCTAIFYTLMTCEMLNRRNGSLRIALSEHLKQRREVAKTVFCLVVIFALCWFPLHLSRILKKTVYNEMDKNRCELLSFLLLMDYIGINLATMNSCINPIALYFVSKKFKNCFQSCLCCCCYQSKSLMTSVPMNGTSIQWKNHDQNNHNTDRSSHKDSMN |
| UniProt | P25101 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P25101 |
| 3D structure model | This predicted structure model is from GPCR-EXP P25101. |
| BioLiP | N/A |
| Therapeutic Target Database | T23499 |
| ChEMBL | CHEMBL252 |
| IUPHAR | 219 |
| DrugBank | BE0000521 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 266 | 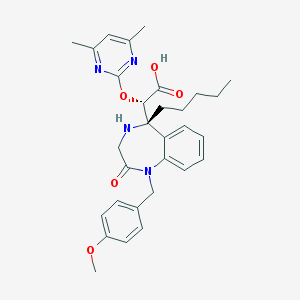 CHEMBL329744 CHEMBL329744 | C30H36N4O5 | 532.641 | 8 / 2 | 2.5 | No |
| 439 | 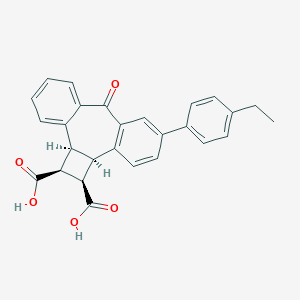 CHEMBL31662 CHEMBL31662 | C27H22O5 | 426.468 | 5 / 2 | 4.5 | Yes |
| 1162 | 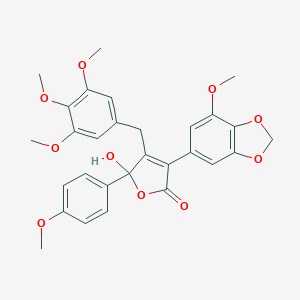 CHEMBL10781 CHEMBL10781 | C29H28O10 | 536.533 | 10 / 1 | 3.9 | No |
| 1701 | 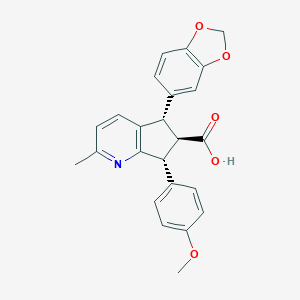 CHEMBL103001 CHEMBL103001 | C24H21NO5 | 403.434 | 6 / 1 | 3.8 | Yes |
| 2389 | 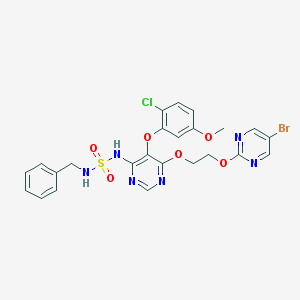 CHEMBL2165330 CHEMBL2165330 | C24H22BrClN6O6S | 637.89 | 12 / 2 | 4.1 | No |
| 2923 | 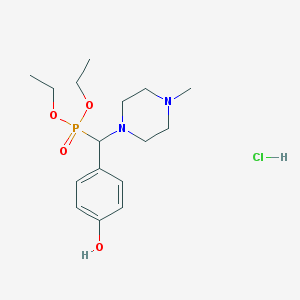 CHEMBL540049 CHEMBL540049 | C16H28ClN2O4P | 378.834 | 6 / 2 | N/A | N/A |
| 2933 | 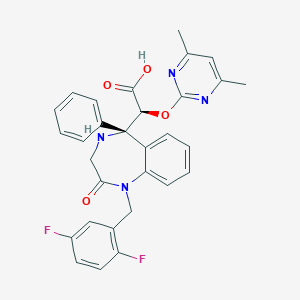 CHEMBL94438 CHEMBL94438 | C30H26F2N4O4 | 544.559 | 9 / 2 | 2.1 | No |
| 3292 | 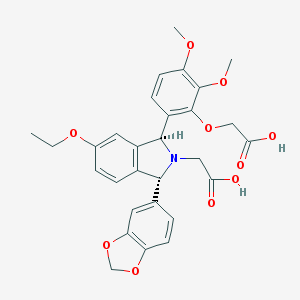 CHEMBL47868 CHEMBL47868 | C29H29NO10 | 551.548 | 11 / 2 | 1.7 | No |
| 3294 | 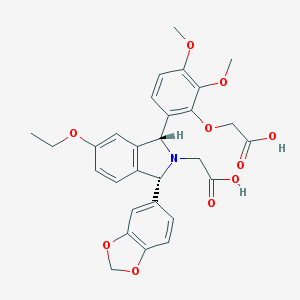 CHEMBL298664 CHEMBL298664 | C29H29NO10 | 551.548 | 11 / 2 | 1.7 | No |
| 3362 | 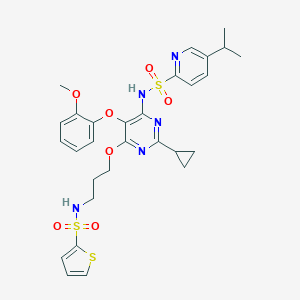 CHEMBL367444 CHEMBL367444 | C29H33N5O7S3 | 659.791 | 13 / 2 | 4.9 | No |
| 3377 | 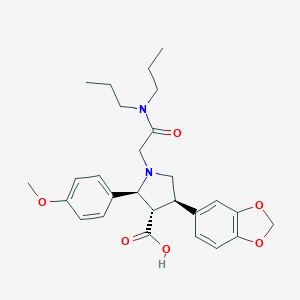 CHEMBL9139 CHEMBL9139 | C27H34N2O6 | 482.577 | 7 / 1 | 1.7 | Yes |
| 3572 | 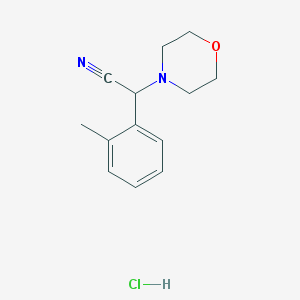 CHEMBL545490 CHEMBL545490 | C13H17ClN2O | 252.742 | 3 / 1 | N/A | N/A |
| 4671 | 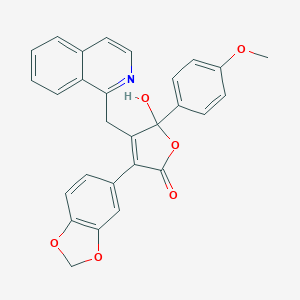 CHEMBL10860 CHEMBL10860 | C28H21NO6 | 467.477 | 7 / 1 | 4.3 | Yes |
| 5290 | 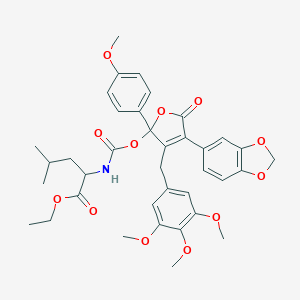 CHEMBL316523 CHEMBL316523 | C37H41NO12 | 691.73 | 12 / 1 | 6.4 | No |
| 5467 | 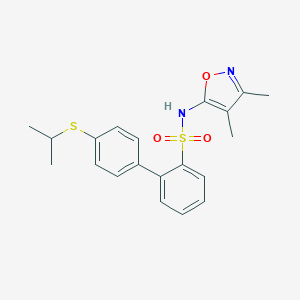 CHEMBL348685 CHEMBL348685 | C20H22N2O3S2 | 402.527 | 6 / 1 | 4.6 | Yes |
| 5830 | 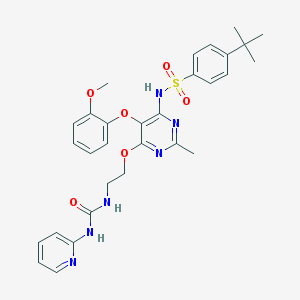 CHEMBL70924 CHEMBL70924 | C30H34N6O6S | 606.698 | 10 / 3 | 4.9 | No |
| 6226 | 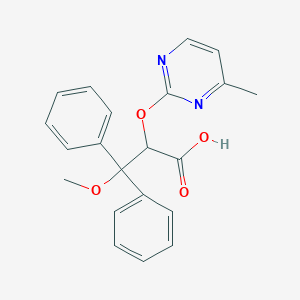 CHEMBL306218 CHEMBL306218 | C21H20N2O4 | 364.401 | 6 / 1 | 3.4 | Yes |
| 6282 | 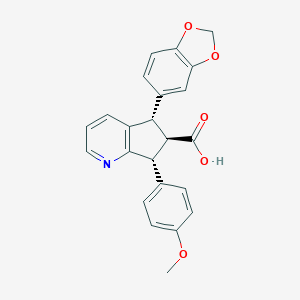 CHEMBL102825 CHEMBL102825 | C23H19NO5 | 389.407 | 6 / 1 | 3.4 | Yes |
| 6514 | 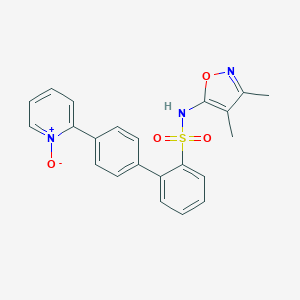 CHEMBL147238 CHEMBL147238 | C22H19N3O4S | 421.471 | 6 / 1 | 3.2 | Yes |
| 6578 | 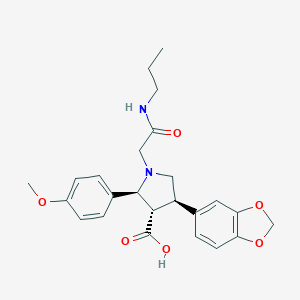 CHEMBL9141 CHEMBL9141 | C24H28N2O6 | 440.496 | 7 / 2 | 0.6 | Yes |
| 6617 | 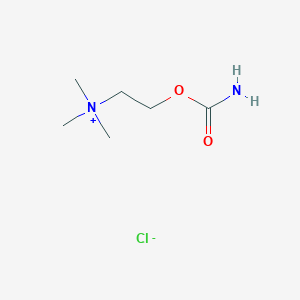 carbachol carbachol | C6H15ClN2O2 | 182.648 | 3 / 1 | N/A | N/A |
| 6824 | 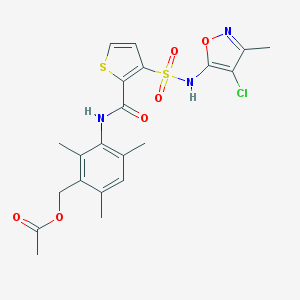 CHEMBL303743 CHEMBL303743 | C21H22ClN3O6S2 | 511.992 | 9 / 2 | 3.9 | No |
| 7104 | 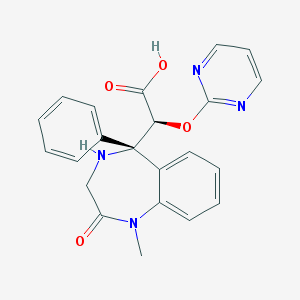 CHEMBL92612 CHEMBL92612 | C22H20N4O4 | 404.426 | 7 / 2 | -0.4 | Yes |
| 7802 | 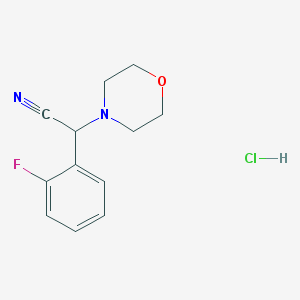 CHEMBL545738 CHEMBL545738 | C12H14ClFN2O | 256.705 | 4 / 1 | N/A | N/A |
| 8293 | 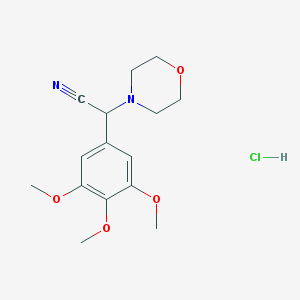 CHEMBL544331 CHEMBL544331 | C15H21ClN2O4 | 328.793 | 6 / 1 | N/A | N/A |
| 8440 | 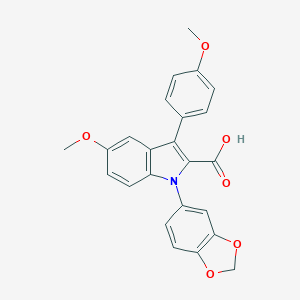 CHEMBL274370 CHEMBL274370 | C24H19NO6 | 417.417 | 6 / 1 | 4.9 | Yes |
| 8624 | 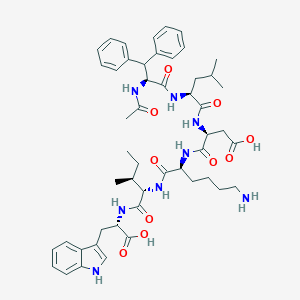 CHEMBL94525 CHEMBL94525 | C50H66N8O10 | 939.124 | 11 / 10 | 2.3 | No |
| 9307 | 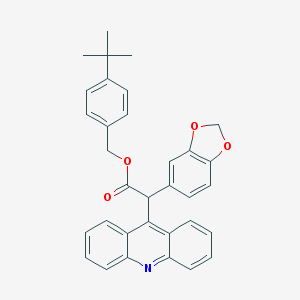 CHEMBL319679 CHEMBL319679 | C33H29NO4 | 503.598 | 5 / 0 | 8.0 | No |
| 9403 | 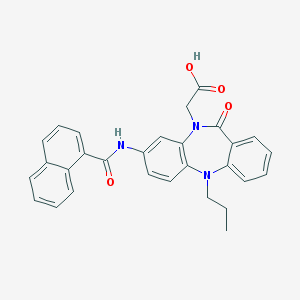 CHEMBL316247 CHEMBL316247 | C29H25N3O4 | 479.536 | 5 / 2 | 5.3 | No |
| 9864 | 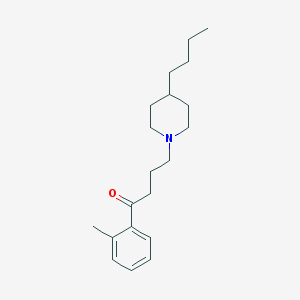 AC-42 AC-42 | C20H31NO | 301.474 | 2 / 0 | 5.2 | No |
| 9914 | 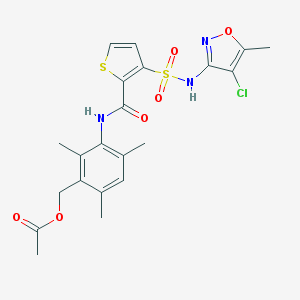 CHEMBL343785 CHEMBL343785 | C21H22ClN3O6S2 | 511.992 | 9 / 2 | 3.9 | No |
| 10238 | 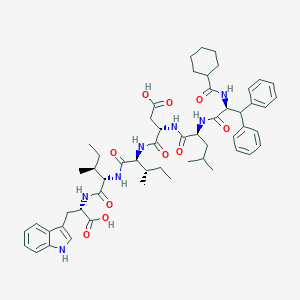 CHEMBL414082 CHEMBL414082 | C55H73N7O10 | 992.228 | 10 / 9 | 8.2 | No |
| 10367 | 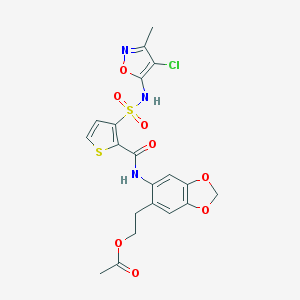 CHEMBL296927 CHEMBL296927 | C20H18ClN3O8S2 | 527.947 | 11 / 2 | 3.1 | No |
| 10418 | 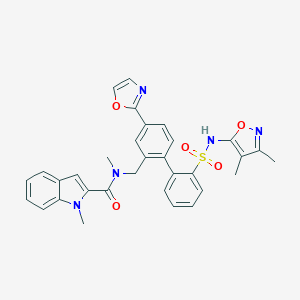 CHEMBL281977 CHEMBL281977 | C32H29N5O5S | 595.674 | 8 / 1 | 5.1 | No |
| 11425 | 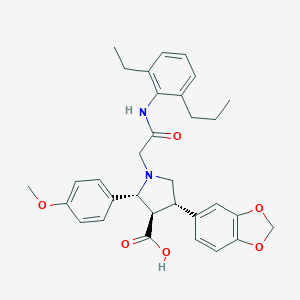 CHEMBL123997 CHEMBL123997 | C32H36N2O6 | 544.648 | 7 / 2 | 3.4 | No |
| 11624 | 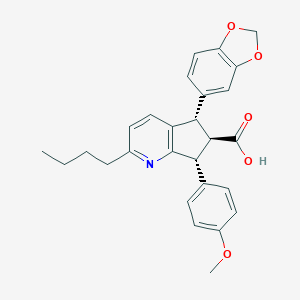 CHEMBL102969 CHEMBL102969 | C27H27NO5 | 445.515 | 6 / 1 | 5.1 | No |
| 521787 | 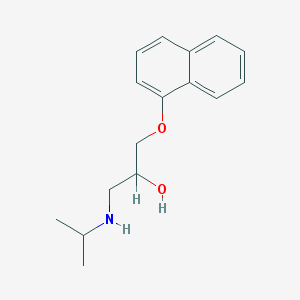 propranolol propranolol | C16H21NO2 | 259.349 | 3 / 2 | 3.0 | Yes |
| 11838 | 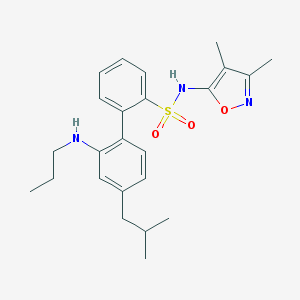 CHEMBL146845 CHEMBL146845 | C24H31N3O3S | 441.59 | 6 / 2 | 6.1 | No |
| 12289 | 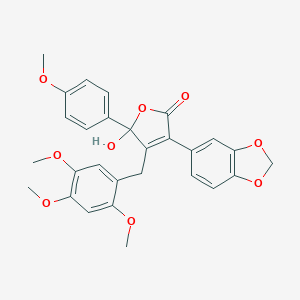 CHEMBL275701 CHEMBL275701 | C28H26O9 | 506.507 | 9 / 1 | 4.0 | No |
| 12372 | 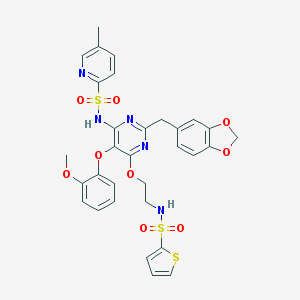 CHEMBL368945 CHEMBL368945 | C31H29N5O9S3 | 711.779 | 15 / 2 | 4.9 | No |
| 12415 | 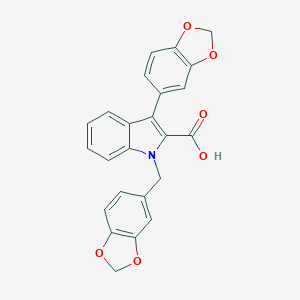 CHEMBL416518 CHEMBL416518 | C24H17NO6 | 415.401 | 6 / 1 | 4.7 | Yes |
| 12418 | 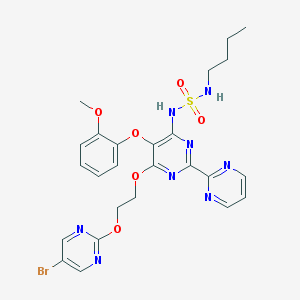 CHEMBL2163720 CHEMBL2163720 | C25H27BrN8O6S | 647.505 | 14 / 2 | 3.2 | No |
| 12714 | 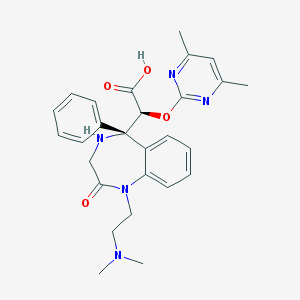 CHEMBL450628 CHEMBL450628 | C27H31N5O4 | 489.576 | 8 / 2 | 0.4 | Yes |
| 12823 | 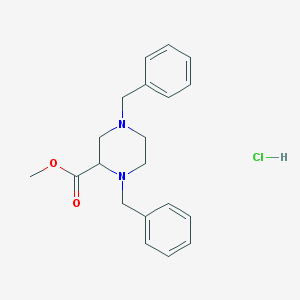 CHEMBL541312 CHEMBL541312 | C20H25ClN2O2 | 360.882 | 4 / 1 | N/A | N/A |
| 12877 | 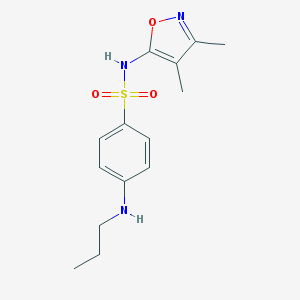 CHEMBL286682 CHEMBL286682 | C14H19N3O3S | 309.384 | 6 / 2 | 2.6 | Yes |
| 13421 | 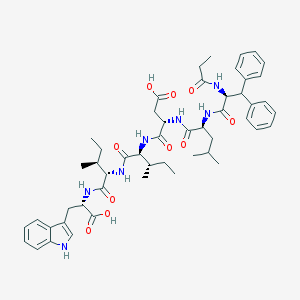 CHEMBL323331 CHEMBL323331 | C51H67N7O10 | 938.136 | 10 / 9 | 6.6 | No |
| 13539 | 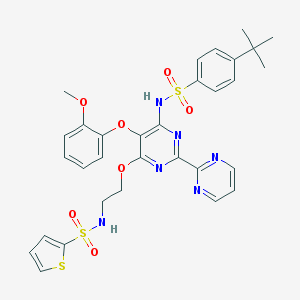 CHEMBL172030 CHEMBL172030 | C31H32N6O7S3 | 696.812 | 14 / 2 | 5.2 | No |
| 13718 | 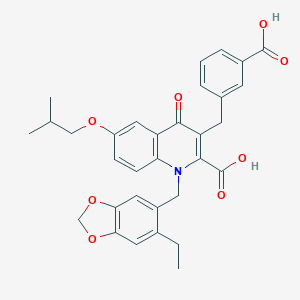 CHEMBL1271640 CHEMBL1271640 | C32H31NO8 | 557.599 | 9 / 2 | 6.3 | No |
| 13892 | 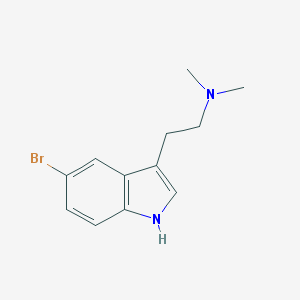 5-Bromo-N,N-dimethyltryptamine 5-Bromo-N,N-dimethyltryptamine | C12H15BrN2 | 267.17 | 1 / 1 | 2.2 | Yes |
| 14948 | 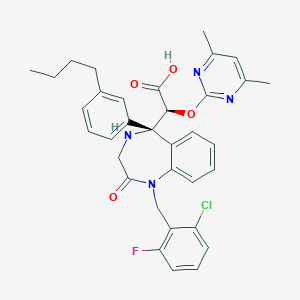 CHEMBL330135 CHEMBL330135 | C34H34ClFN4O4 | 617.118 | 8 / 2 | 4.5 | No |
| 15336 | 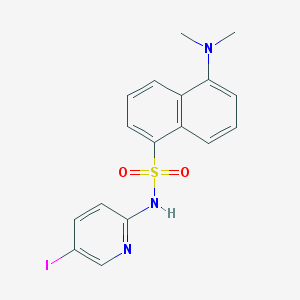 CHEMBL266684 CHEMBL266684 | C17H16IN3O2S | 453.298 | 5 / 1 | 3.7 | Yes |
| 15620 | 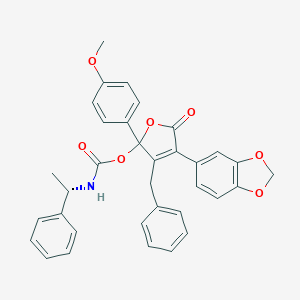 CHEMBL103501 CHEMBL103501 | C34H29NO7 | 563.606 | 7 / 1 | 6.3 | No |
| 15657 | 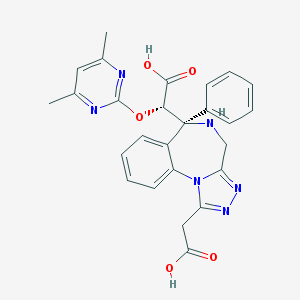 SCHEMBL5351769 SCHEMBL5351769 | C26H24N6O5 | 500.515 | 10 / 3 | -0.5 | No |
| 15770 | 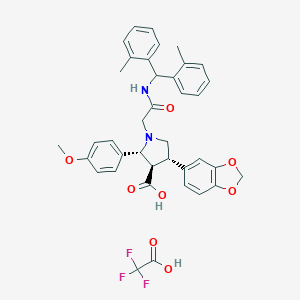 CHEMBL332109 CHEMBL332109 | C38H37F3N2O8 | 706.715 | 12 / 3 | N/A | No |
| 15862 | 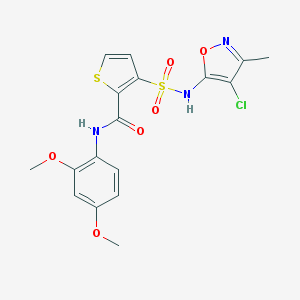 CHEMBL43788 CHEMBL43788 | C17H16ClN3O6S2 | 457.9 | 9 / 2 | 3.0 | Yes |
| 16367 | 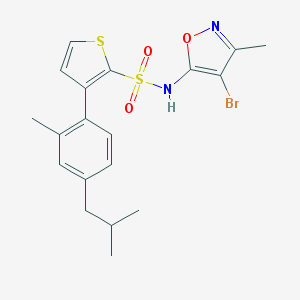 CHEMBL432815 CHEMBL432815 | C19H21BrN2O3S2 | 469.412 | 6 / 1 | 5.9 | No |
| 16568 | 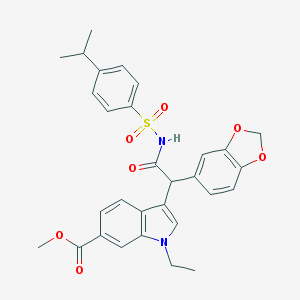 CHEMBL23418 CHEMBL23418 | C30H30N2O7S | 562.637 | 7 / 1 | 5.1 | No |
| 16660 | 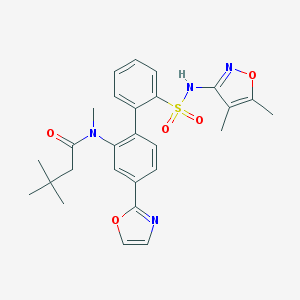 CHEMBL274489 CHEMBL274489 | C27H30N4O5S | 522.62 | 8 / 1 | 4.7 | No |
| 16894 | 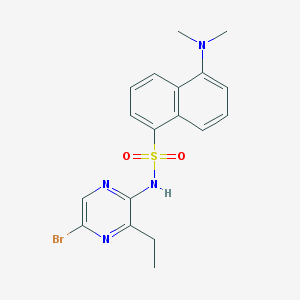 CHEMBL6543 CHEMBL6543 | C18H19BrN4O2S | 435.34 | 6 / 1 | 3.9 | Yes |
| 16912 | 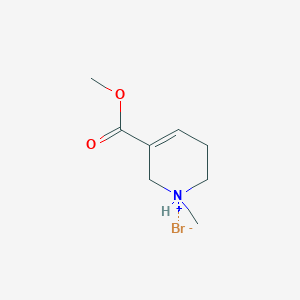 CID 57486785 CID 57486785 | C8H14BrNO2 | 236.109 | 3 / 1 | N/A | N/A |
| 17105 | 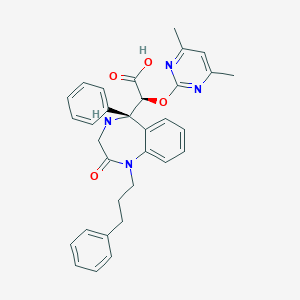 CHEMBL96114 CHEMBL96114 | C32H32N4O4 | 536.632 | 7 / 2 | 2.7 | No |
| 17133 | 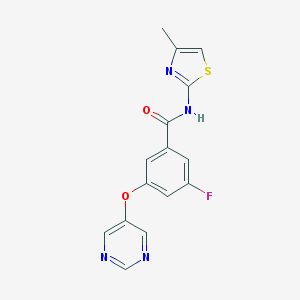 CHEMBL2440659 CHEMBL2440659 | C15H11FN4O2S | 330.337 | 7 / 1 | 2.4 | Yes |
| 521989 | 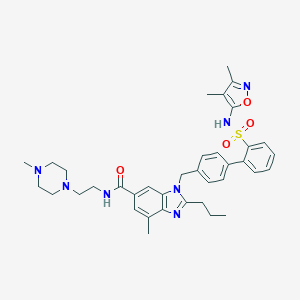 CHEMBL3735236 CHEMBL3735236 | C37H45N7O4S | 683.872 | 9 / 2 | 5.4 | No |
| 17658 | 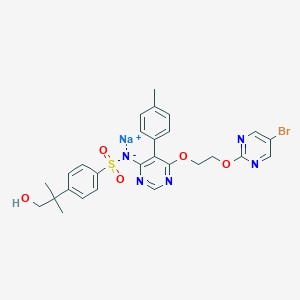 CHEMBL1627022 CHEMBL1627022 | C27H27BrN5NaO5S | 636.497 | 10 / 1 | N/A | No |
| 18021 | 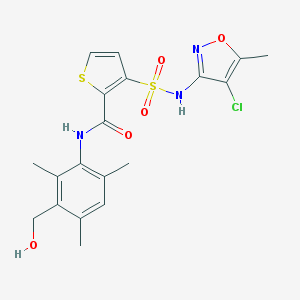 CHEMBL424101 CHEMBL424101 | C19H20ClN3O5S2 | 469.955 | 8 / 3 | 3.3 | Yes |
| 18582 | 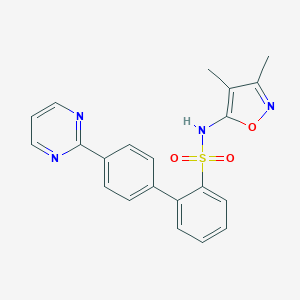 CHEMBL359273 CHEMBL359273 | C21H18N4O3S | 406.46 | 7 / 1 | 3.6 | Yes |
| 18736 | 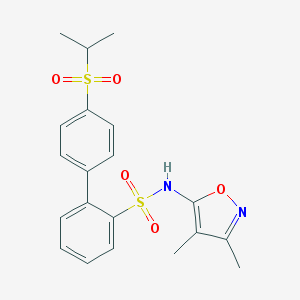 CHEMBL149050 CHEMBL149050 | C20H22N2O5S2 | 434.525 | 7 / 1 | 3.4 | Yes |
| 19173 | 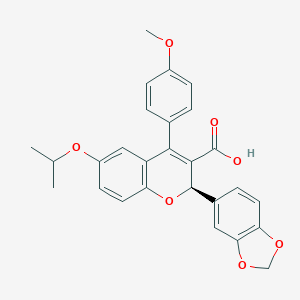 CHEMBL61211 CHEMBL61211 | C27H24O7 | 460.482 | 7 / 1 | 4.9 | Yes |
| 19651 | 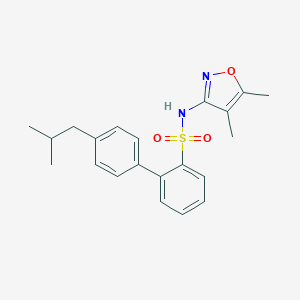 CHEMBL434082 CHEMBL434082 | C21H24N2O3S | 384.494 | 5 / 1 | 5.2 | No |
| 20155 | 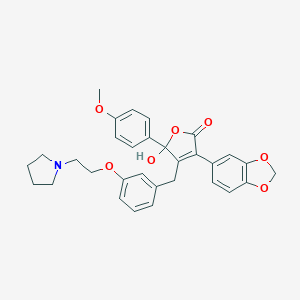 CHEMBL65939 CHEMBL65939 | C31H31NO7 | 529.589 | 8 / 1 | 4.5 | No |
| 20388 | 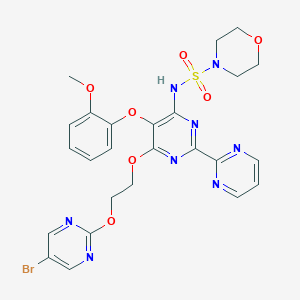 CHEMBL2163722 CHEMBL2163722 | C25H25BrN8O7S | 661.488 | 15 / 1 | 1.8 | No |
| 20438 | 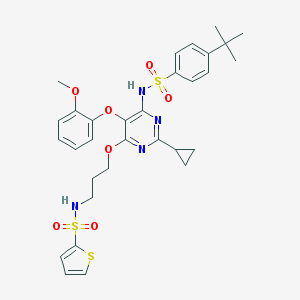 CHEMBL172297 CHEMBL172297 | C31H36N4O7S3 | 672.83 | 12 / 2 | 6.1 | No |
| 20895 | 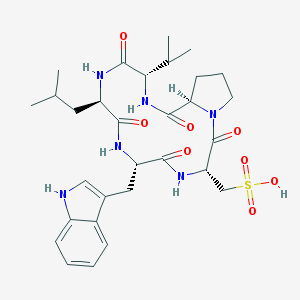 CHEMBL342615 CHEMBL342615 | C30H42N6O8S | 646.76 | 8 / 6 | 1.6 | No |
| 20904 | 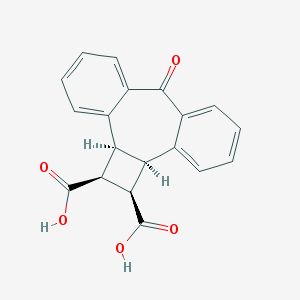 CHEMBL286808 CHEMBL286808 | C19H14O5 | 322.316 | 5 / 2 | 2.1 | Yes |
| 21071 | 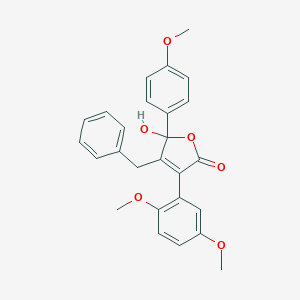 CHEMBL267096 CHEMBL267096 | C26H24O6 | 432.472 | 6 / 1 | 4.2 | Yes |
| 21168 | 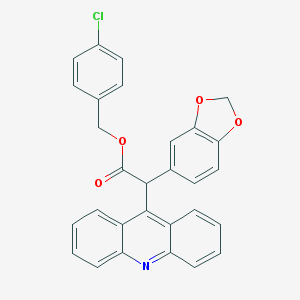 CHEMBL101292 CHEMBL101292 | C29H20ClNO4 | 481.932 | 5 / 0 | 6.9 | No |
| 21299 | 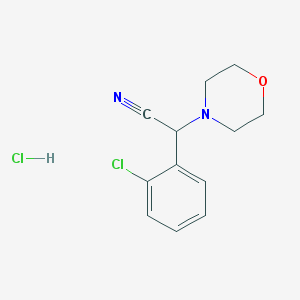 CHEMBL545270 CHEMBL545270 | C12H14Cl2N2O | 273.157 | 3 / 1 | N/A | N/A |
| 21535 | 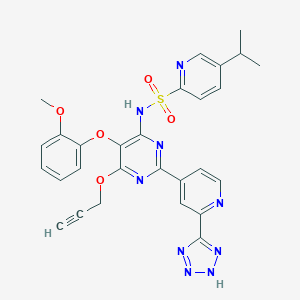 CHEMBL175057 CHEMBL175057 | C28H25N9O5S | 599.626 | 13 / 2 | 3.2 | No |
| 21718 | 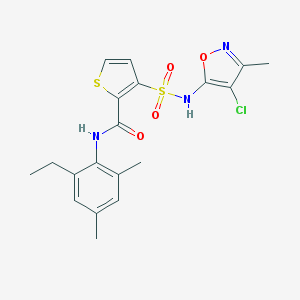 CHEMBL283582 CHEMBL283582 | C19H20ClN3O4S2 | 453.956 | 7 / 2 | 4.6 | Yes |
| 21765 | 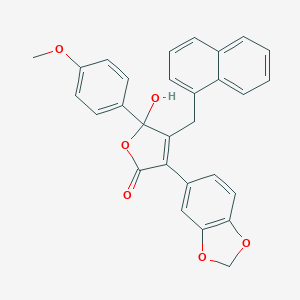 CHEMBL269409 CHEMBL269409 | C29H22O6 | 466.489 | 6 / 1 | 5.3 | No |
| 522150 | 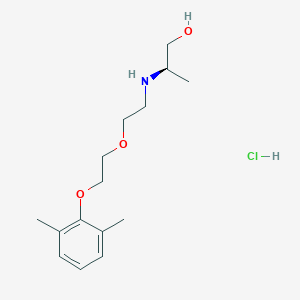 CHEMBL3775436 CHEMBL3775436 | C15H26ClNO3 | 303.827 | 4 / 3 | N/A | N/A |
| 522183 | 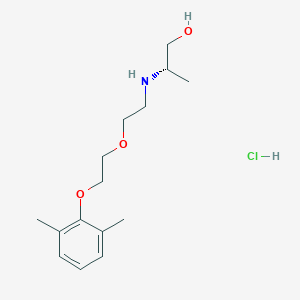 CHEMBL3775769 CHEMBL3775769 | C15H26ClNO3 | 303.827 | 4 / 3 | N/A | N/A |
| 22294 | 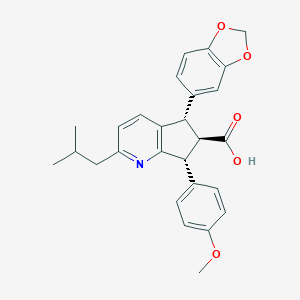 CHEMBL102314 CHEMBL102314 | C27H27NO5 | 445.515 | 6 / 1 | 5.0 | Yes |
| 22329 | 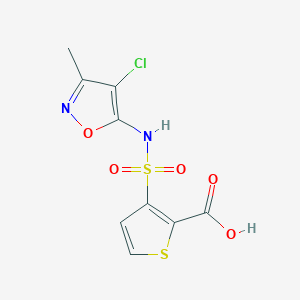 184040-74-2 184040-74-2 | C9H7ClN2O5S2 | 322.734 | 8 / 2 | 1.8 | Yes |
| 22723 | 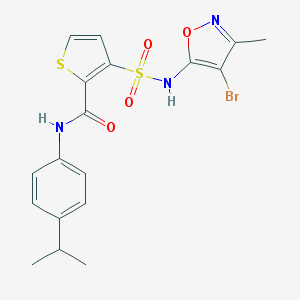 CHEMBL47417 CHEMBL47417 | C18H18BrN3O4S2 | 484.383 | 7 / 2 | 4.3 | Yes |
| 23059 | 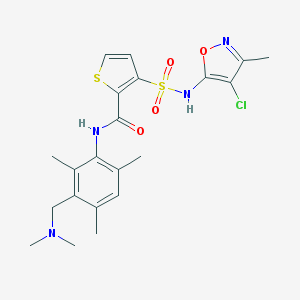 CHEMBL343578 CHEMBL343578 | C21H25ClN4O4S2 | 497.025 | 8 / 2 | 4.0 | Yes |
| 23458 | 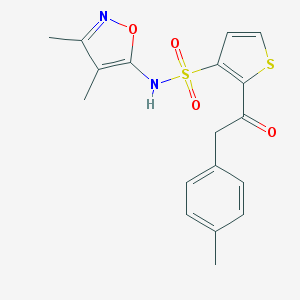 CHEMBL45875 CHEMBL45875 | C18H18N2O4S2 | 390.472 | 7 / 1 | 3.6 | Yes |
| 24316 | 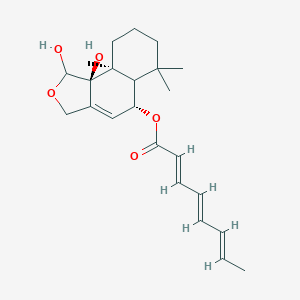 CHEMBL170335 CHEMBL170335 | C23H32O5 | 388.504 | 5 / 2 | 3.4 | Yes |
| 24458 | 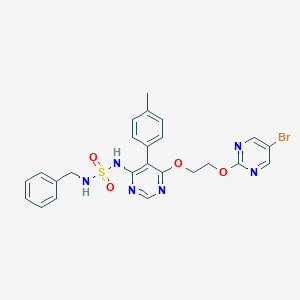 CHEMBL2165332 CHEMBL2165332 | C24H23BrN6O4S | 571.45 | 10 / 2 | 4.0 | No |
| 24824 | 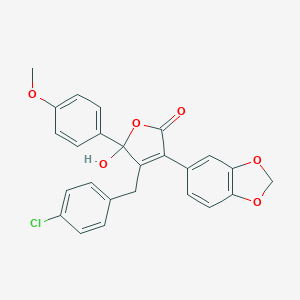 CHEMBL10733 CHEMBL10733 | C25H19ClO6 | 450.871 | 6 / 1 | 4.7 | Yes |
| 25431 | 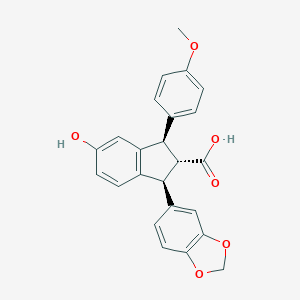 CHEMBL41087 CHEMBL41087 | C24H20O6 | 404.418 | 6 / 2 | 4.1 | Yes |
| 25453 | 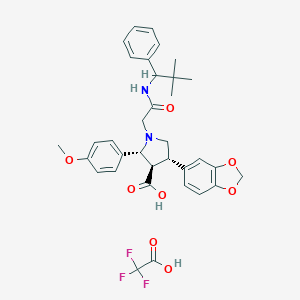 CHEMBL123316 CHEMBL123316 | C34H37F3N2O8 | 658.671 | 12 / 3 | N/A | No |
| 25617 | 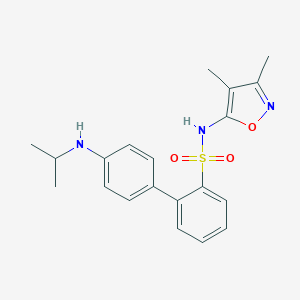 CHEMBL343867 CHEMBL343867 | C20H23N3O3S | 385.482 | 6 / 2 | 4.1 | Yes |
| 25765 | 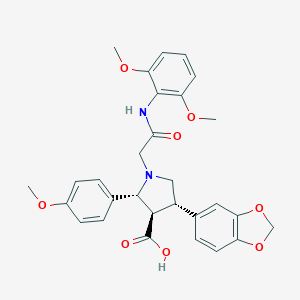 CHEMBL122832 CHEMBL122832 | C29H30N2O8 | 534.565 | 9 / 2 | 1.2 | No |
| 25911 | 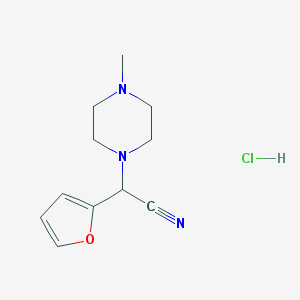 CHEMBL554094 CHEMBL554094 | C11H16ClN3O | 241.719 | 4 / 1 | N/A | N/A |
| 25973 | 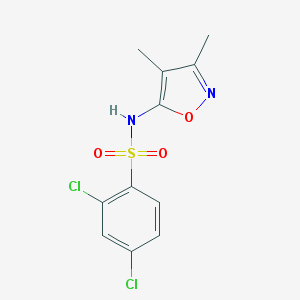 CHEMBL79054 CHEMBL79054 | C11H10Cl2N2O3S | 321.172 | 5 / 1 | 2.9 | Yes |
| 26174 | 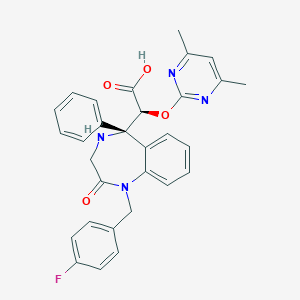 CHEMBL314755 CHEMBL314755 | C30H27FN4O4 | 526.568 | 8 / 2 | 2.0 | No |
| 26889 | 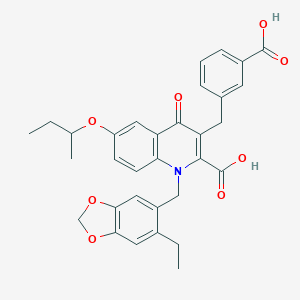 CHEMBL1271691 CHEMBL1271691 | C32H31NO8 | 557.599 | 9 / 2 | 6.3 | No |
| 26958 | 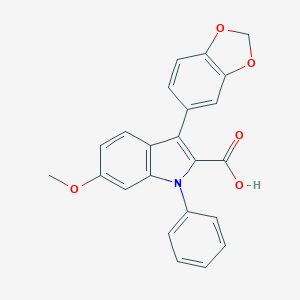 CHEMBL276615 CHEMBL276615 | C23H17NO5 | 387.391 | 5 / 1 | 4.9 | Yes |
| 27221 | 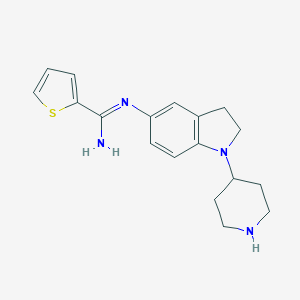 CHEMBL2203713 CHEMBL2203713 | C18H22N4S | 326.462 | 4 / 2 | 2.9 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218