

You can:
| Name | Atypical chemokine receptor 3 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | ACKR3 |
| Synonym | RDC-1 GPR159 G-protein coupled receptor RDC1 homolog G-protein coupled receptor 159 Cxcr7 [ Show all ] |
| Disease | Cancer Asthma |
| Length | 362 |
| Amino acid sequence | MDLHLFDYSEPGNFSDISWPCNSSDCIVVDTVMCPNMPNKSVLLYTLSFIYIFIFVIGMIANSVVVWVNIQAKTTGYDTHCYILNLAIADLWVVLTIPVWVVSLVQHNQWPMGELTCKVTHLIFSINLFGSIFFLTCMSVDRYLSITYFTNTPSSRKKMVRRVVCILVWLLAFCVSLPDTYYLKTVTSASNNETYCRSFYPEHSIKEWLIGMELVSVVLGFAVPFSIIAVFYFLLARAISASSDQEKHSSRKIIFSYVVVFLVCWLPYHVAVLLDIFSILHYIPFTCRLEHALFTALHVTQCLSLVHCCVNPVLYSFINRNYRYELMKAFIFKYSAKTGLTKLIDASRVSETEYSALEQSTK |
| UniProt | P25106 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P25106 |
| 3D structure model | This predicted structure model is from GPCR-EXP P25106. |
| BioLiP | N/A |
| Therapeutic Target Database | T10491 |
| ChEMBL | CHEMBL2010631 |
| IUPHAR | 80 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 533914 | 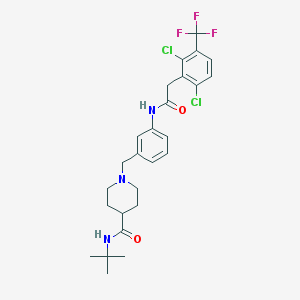 CHEMBL3921107 CHEMBL3921107 | C26H30Cl2F3N3O2 | 544.44 | 6 / 2 | 5.4 | No |
| 533917 | 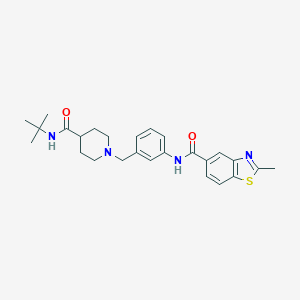 CHEMBL3978976 CHEMBL3978976 | C26H32N4O2S | 464.628 | 5 / 2 | 4.2 | Yes |
| 535946 | 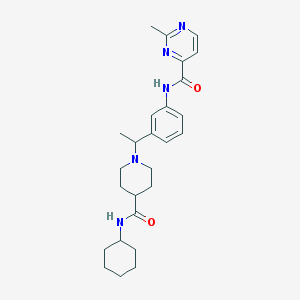 CHEMBL3915132 CHEMBL3915132 | C26H35N5O2 | 449.599 | 5 / 2 | 3.6 | Yes |
| 533922 | 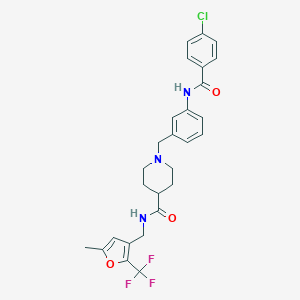 CHEMBL3915468 CHEMBL3915468 | C27H27ClF3N3O3 | 533.976 | 7 / 2 | 4.9 | No |
| 536039 | 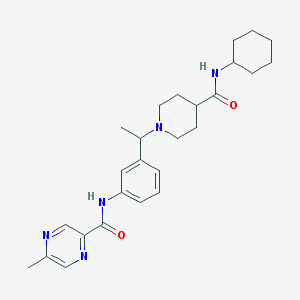 CHEMBL3951108 CHEMBL3951108 | C26H35N5O2 | 449.599 | 5 / 2 | 3.2 | Yes |
| 533931 | 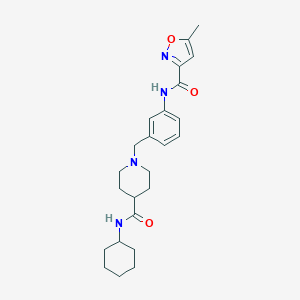 CHEMBL3899484 CHEMBL3899484 | C24H32N4O3 | 424.545 | 5 / 2 | 3.4 | Yes |
| 533935 | 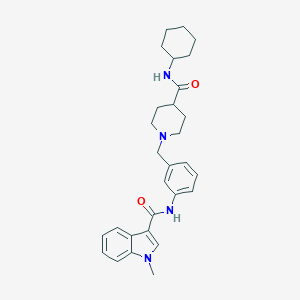 CHEMBL3982167 CHEMBL3982167 | C29H36N4O2 | 472.633 | 3 / 2 | 4.3 | Yes |
| 463830 | 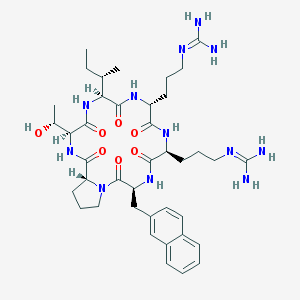 CHEMBL3581286 CHEMBL3581286 | C40H60N12O7 | 820.997 | 9 / 10 | 0.5 | No |
| 533938 | 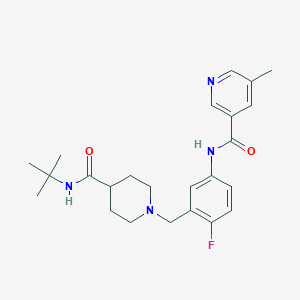 CHEMBL3982702 CHEMBL3982702 | C24H31FN4O2 | 426.536 | 5 / 2 | 2.7 | Yes |
| 533939 | 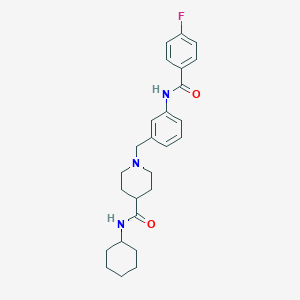 CHEMBL3923052 CHEMBL3923052 | C26H32FN3O2 | 437.559 | 4 / 2 | 4.3 | Yes |
| 533941 | 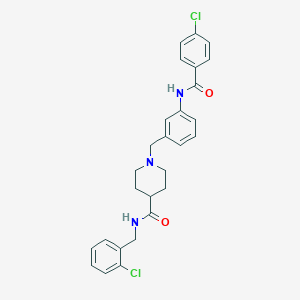 CHEMBL3905974 CHEMBL3905974 | C27H27Cl2N3O2 | 496.432 | 3 / 2 | 5.1 | No |
| 533946 | 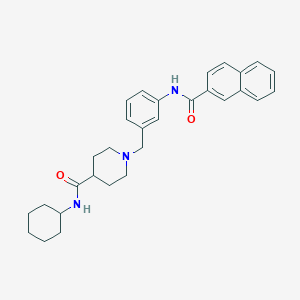 CHEMBL3942036 CHEMBL3942036 | C30H35N3O2 | 469.629 | 3 / 2 | 5.4 | No |
| 533947 | 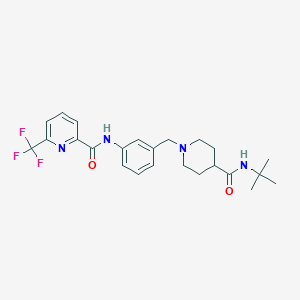 CHEMBL3889615 CHEMBL3889615 | C24H29F3N4O2 | 462.517 | 7 / 2 | 3.5 | Yes |
| 536229 | 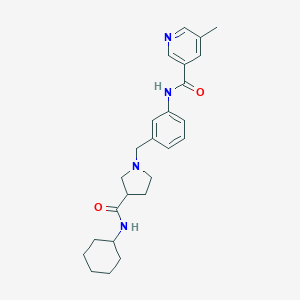 CHEMBL3974763 CHEMBL3974763 | C25H32N4O2 | 420.557 | 4 / 2 | 3.1 | Yes |
| 533950 | 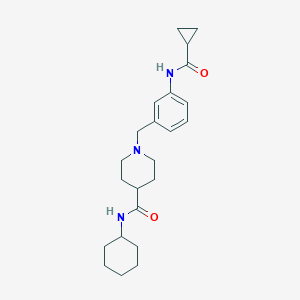 CHEMBL3905756 CHEMBL3905756 | C23H33N3O2 | 383.536 | 3 / 2 | 3.0 | Yes |
| 533955 | 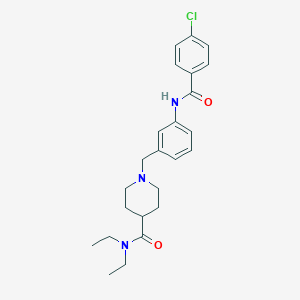 CHEMBL3901696 CHEMBL3901696 | C24H30ClN3O2 | 427.973 | 3 / 1 | 3.9 | Yes |
| 533959 | 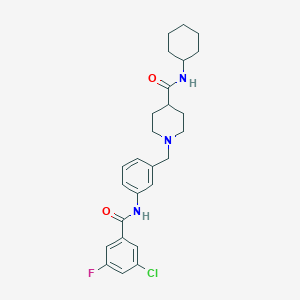 CHEMBL3970605 CHEMBL3970605 | C26H31ClFN3O2 | 472.001 | 4 / 2 | 4.9 | Yes |
| 533960 | 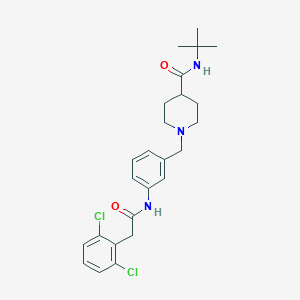 CHEMBL3948534 CHEMBL3948534 | C25H31Cl2N3O2 | 476.442 | 3 / 2 | 4.5 | Yes |
| 533963 | 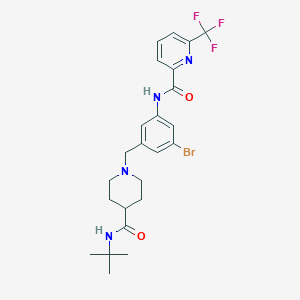 CHEMBL3957872 CHEMBL3957872 | C24H28BrF3N4O2 | 541.413 | 7 / 2 | 4.2 | No |
| 533964 | 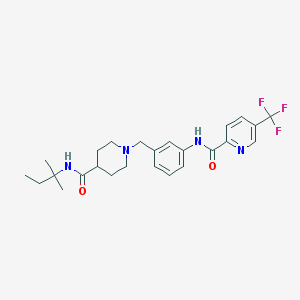 CHEMBL3979681 CHEMBL3979681 | C25H31F3N4O2 | 476.544 | 7 / 2 | 4.0 | Yes |
| 533966 | 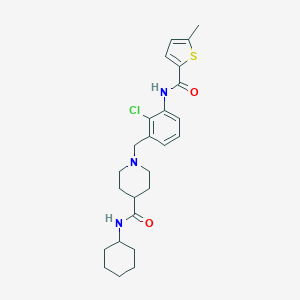 CHEMBL3983033 CHEMBL3983033 | C25H32ClN3O2S | 474.06 | 4 / 2 | 5.2 | No |
| 533971 | 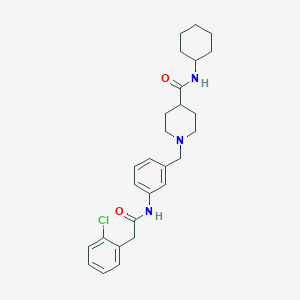 CHEMBL3985336 CHEMBL3985336 | C27H34ClN3O2 | 468.038 | 3 / 2 | 4.7 | Yes |
| 536363 | 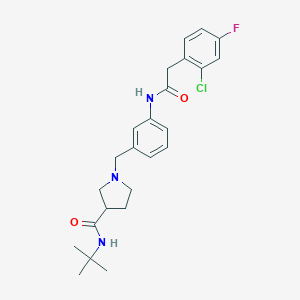 CHEMBL3912367 CHEMBL3912367 | C24H29ClFN3O2 | 445.963 | 4 / 2 | 3.7 | Yes |
| 464764 | 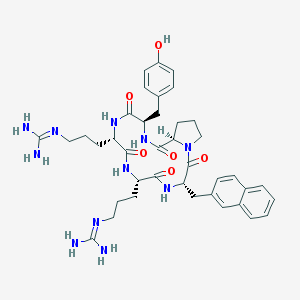 CHEMBL3581265 CHEMBL3581265 | C39H51N11O6 | 769.908 | 8 / 9 | 0.7 | No |
| 464765 | 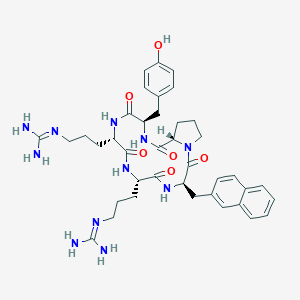 CHEMBL3581273 CHEMBL3581273 | C39H51N11O6 | 769.908 | 8 / 9 | 0.7 | No |
| 464767 | 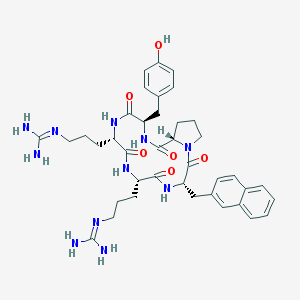 CHEMBL3581264 CHEMBL3581264 | C39H51N11O6 | 769.908 | 8 / 9 | 0.7 | No |
| 464770 | 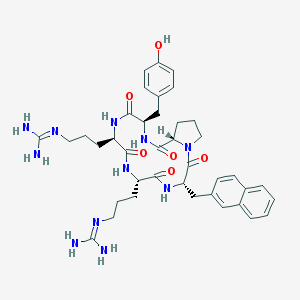 CHEMBL3581271 CHEMBL3581271 | C39H51N11O6 | 769.908 | 8 / 9 | 0.7 | No |
| 464771 | 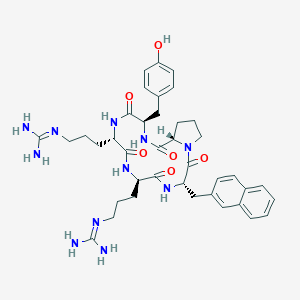 CHEMBL3581272 CHEMBL3581272 | C39H51N11O6 | 769.908 | 8 / 9 | 0.7 | No |
| 464774 | 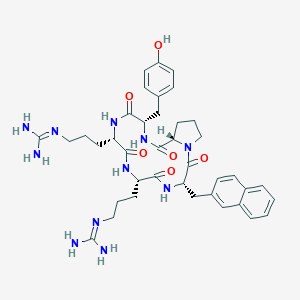 CHEMBL3581270 CHEMBL3581270 | C39H51N11O6 | 769.908 | 8 / 9 | 0.7 | No |
| 533974 | 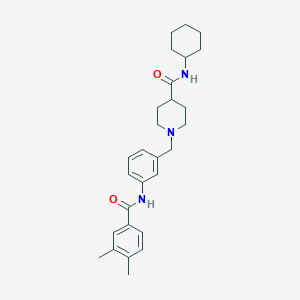 CHEMBL3911168 CHEMBL3911168 | C28H37N3O2 | 447.623 | 3 / 2 | 4.9 | Yes |
| 533979 | 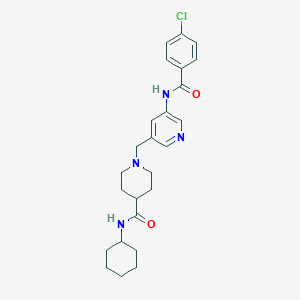 CHEMBL3917580 CHEMBL3917580 | C25H31ClN4O2 | 454.999 | 4 / 2 | 3.7 | Yes |
| 19550 | 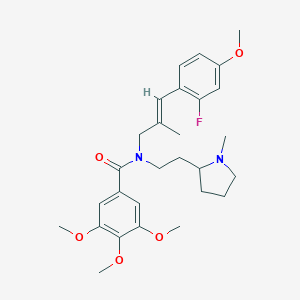 CHEMBL2013232 CHEMBL2013232 | C28H37FN2O5 | 500.611 | 7 / 0 | 5.1 | No |
| 19821 | 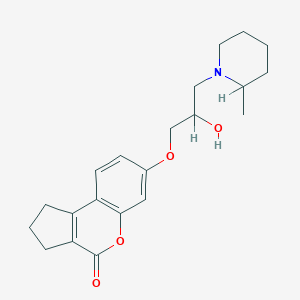 CHEMBL2381324 CHEMBL2381324 | C21H27NO4 | 357.45 | 5 / 1 | 3.0 | Yes |
| 536480 | 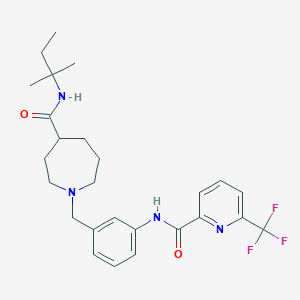 CHEMBL3946330 CHEMBL3946330 | C26H33F3N4O2 | 490.571 | 7 / 2 | 4.4 | Yes |
| 536521 | 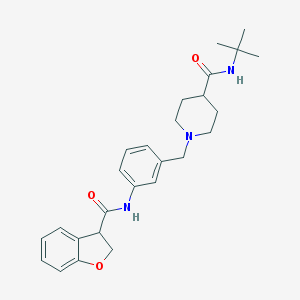 CHEMBL3961653 CHEMBL3961653 | C26H33N3O3 | 435.568 | 4 / 2 | 3.1 | Yes |
| 533994 | 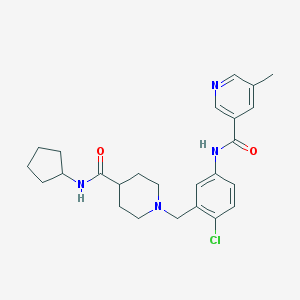 CHEMBL3963631 CHEMBL3963631 | C25H31ClN4O2 | 454.999 | 4 / 2 | 3.6 | Yes |
| 534001 | 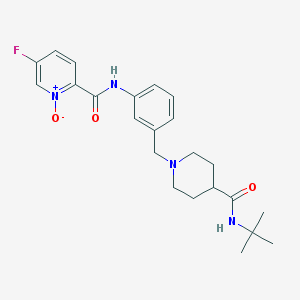 CHEMBL3983752 CHEMBL3983752 | C23H29FN4O3 | 428.508 | 5 / 2 | 1.7 | Yes |
| 534003 | 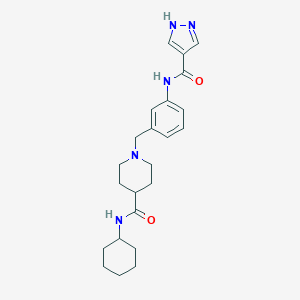 CHEMBL3930062 CHEMBL3930062 | C23H31N5O2 | 409.534 | 4 / 3 | 2.5 | Yes |
| 534007 | 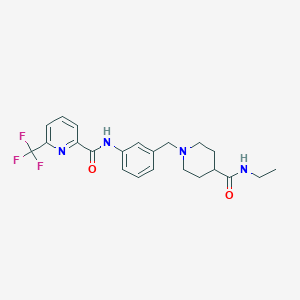 CHEMBL3981571 CHEMBL3981571 | C22H25F3N4O2 | 434.463 | 7 / 2 | 2.9 | Yes |
| 26135 | 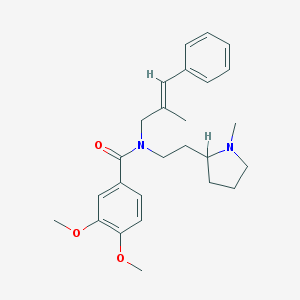 CHEMBL2013219 CHEMBL2013219 | C26H34N2O3 | 422.569 | 4 / 0 | 5.0 | Yes |
| 534011 | 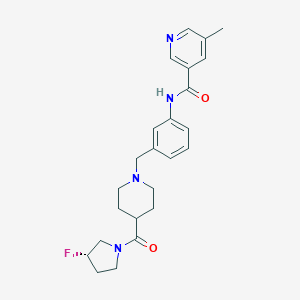 CHEMBL3975274 CHEMBL3975274 | C24H29FN4O2 | 424.52 | 5 / 1 | 2.4 | Yes |
| 534012 | 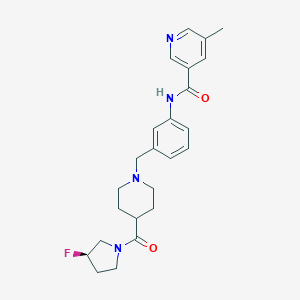 CHEMBL3973660 CHEMBL3973660 | C24H29FN4O2 | 424.52 | 5 / 1 | 2.4 | Yes |
| 534013 | 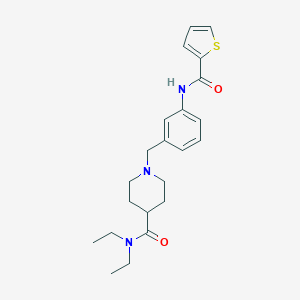 CHEMBL3891911 CHEMBL3891911 | C22H29N3O2S | 399.553 | 4 / 1 | 3.3 | Yes |
| 534014 | 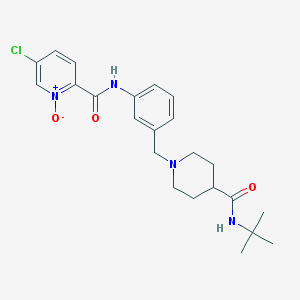 CHEMBL3944347 CHEMBL3944347 | C23H29ClN4O3 | 444.96 | 4 / 2 | 2.2 | Yes |
| 534016 | 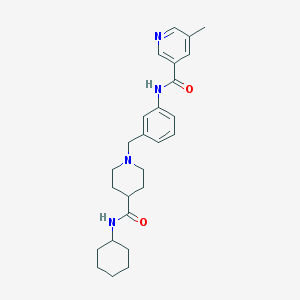 CHEMBL3937850 CHEMBL3937850 | C26H34N4O2 | 434.584 | 4 / 2 | 3.5 | Yes |
| 534019 | 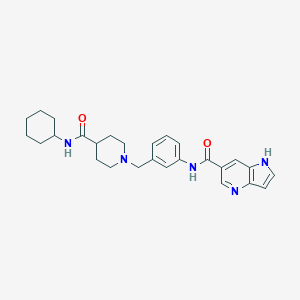 CHEMBL3918035 CHEMBL3918035 | C27H33N5O2 | 459.594 | 4 / 3 | 3.3 | Yes |
| 534021 | 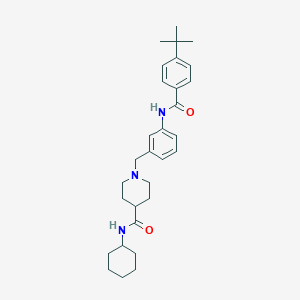 CHEMBL3966602 CHEMBL3966602 | C30H41N3O2 | 475.677 | 3 / 2 | 5.8 | No |
| 534023 | 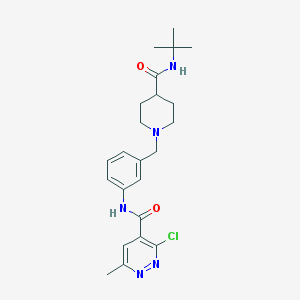 CHEMBL3971888 CHEMBL3971888 | C23H30ClN5O2 | 443.976 | 5 / 2 | 2.6 | Yes |
| 534025 | 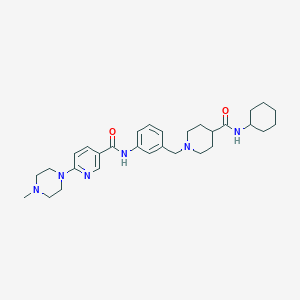 CHEMBL3935099 CHEMBL3935099 | C30H42N6O2 | 518.706 | 6 / 2 | 3.4 | No |
| 534026 | 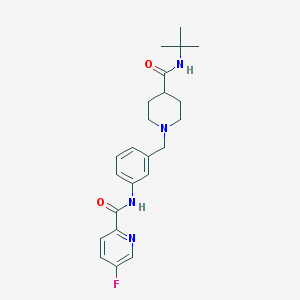 CHEMBL3922943 CHEMBL3922943 | C23H29FN4O2 | 412.509 | 5 / 2 | 2.7 | Yes |
| 534028 | 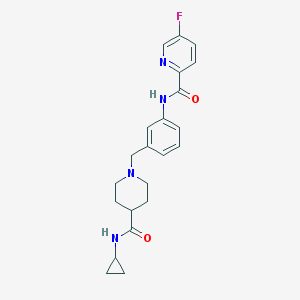 CHEMBL3971504 CHEMBL3971504 | C22H25FN4O2 | 396.466 | 5 / 2 | 2.3 | Yes |
| 466876 | 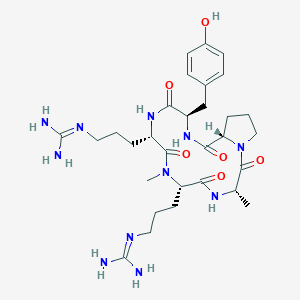 CHEMBL3581285 CHEMBL3581285 | C30H47N11O6 | 657.777 | 8 / 8 | -1.9 | No |
| 534040 | 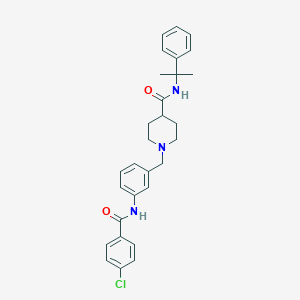 CHEMBL3947322 CHEMBL3947322 | C29H32ClN3O2 | 490.044 | 3 / 2 | 5.1 | No |
| 534041 | 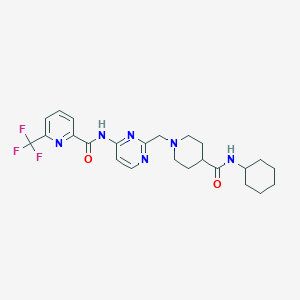 CHEMBL3924510 CHEMBL3924510 | C24H29F3N6O2 | 490.531 | 9 / 2 | 3.0 | Yes |
| 534042 | 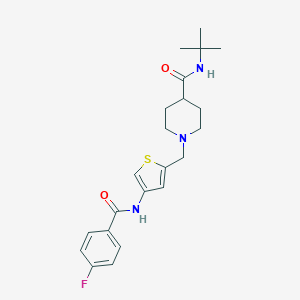 CHEMBL3938037 CHEMBL3938037 | C22H28FN3O2S | 417.543 | 5 / 2 | 3.2 | Yes |
| 534043 | 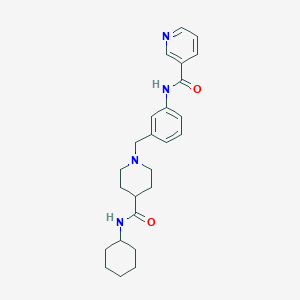 CHEMBL3917109 CHEMBL3917109 | C25H32N4O2 | 420.557 | 4 / 2 | 3.1 | Yes |
| 534048 | 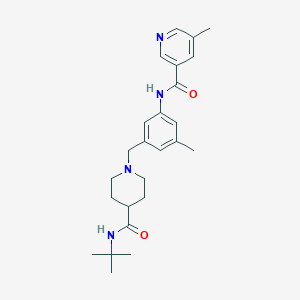 CHEMBL3899145 CHEMBL3899145 | C25H34N4O2 | 422.573 | 4 / 2 | 3.0 | Yes |
| 536889 | 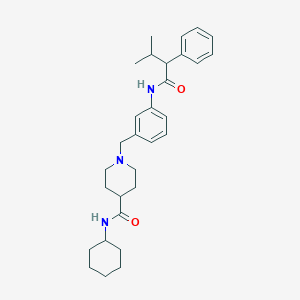 CHEMBL3963024 CHEMBL3963024 | C30H41N3O2 | 475.677 | 3 / 2 | 5.5 | No |
| 534055 | 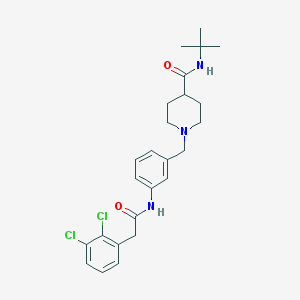 CHEMBL3924230 CHEMBL3924230 | C25H31Cl2N3O2 | 476.442 | 3 / 2 | 4.5 | Yes |
| 534056 | 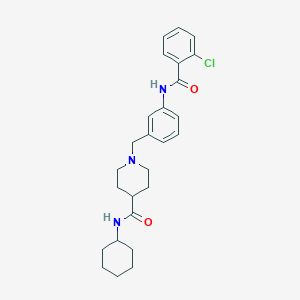 CHEMBL3943089 CHEMBL3943089 | C26H32ClN3O2 | 454.011 | 3 / 2 | 4.8 | Yes |
| 534058 | 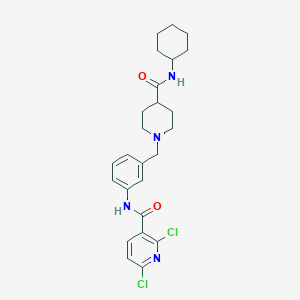 CHEMBL3913386 CHEMBL3913386 | C25H30Cl2N4O2 | 489.441 | 4 / 2 | 5.0 | Yes |
| 534059 | 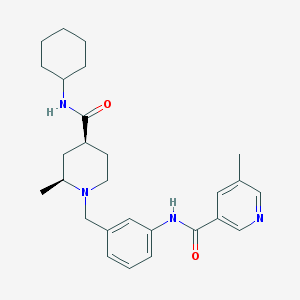 CHEMBL3936492 CHEMBL3936492 | C27H36N4O2 | 448.611 | 4 / 2 | 3.9 | Yes |
| 534069 | 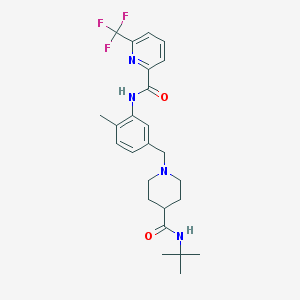 CHEMBL3978234 CHEMBL3978234 | C25H31F3N4O2 | 476.544 | 7 / 2 | 3.9 | Yes |
| 536942 | 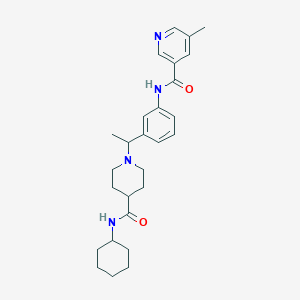 CHEMBL3972572 CHEMBL3972572 | C27H36N4O2 | 448.611 | 4 / 2 | 3.9 | Yes |
| 534072 | 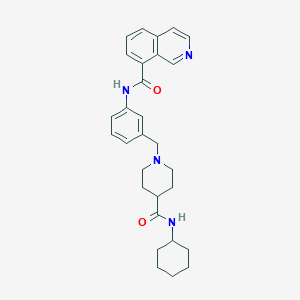 CHEMBL3929887 CHEMBL3929887 | C29H34N4O2 | 470.617 | 4 / 2 | 4.4 | Yes |
| 536964 | 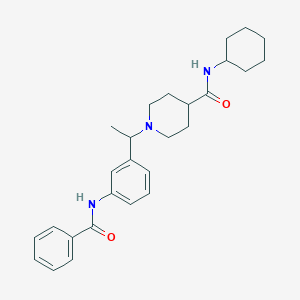 CHEMBL3904295 CHEMBL3904295 | C27H35N3O2 | 433.596 | 3 / 2 | 4.6 | Yes |
| 534083 | 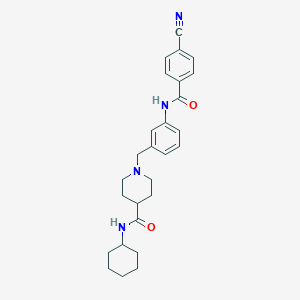 CHEMBL3898127 CHEMBL3898127 | C27H32N4O2 | 444.579 | 4 / 2 | 3.9 | Yes |
| 40727 | 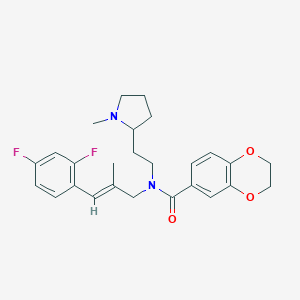 CHEMBL2013233 CHEMBL2013233 | C26H30F2N2O3 | 456.534 | 6 / 0 | 5.0 | Yes |
| 534092 | 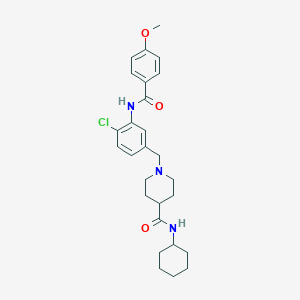 CHEMBL3914298 CHEMBL3914298 | C27H34ClN3O3 | 484.037 | 4 / 2 | 4.8 | Yes |
| 537067 | 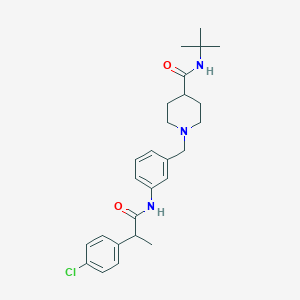 CHEMBL3975469 CHEMBL3975469 | C26H34ClN3O2 | 456.027 | 3 / 2 | 4.5 | Yes |
| 534093 | 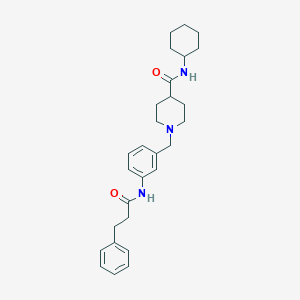 CHEMBL3972023 CHEMBL3972023 | C28H37N3O2 | 447.623 | 3 / 2 | 4.4 | Yes |
| 534094 | 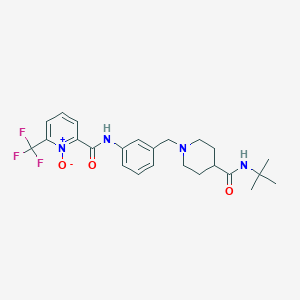 CHEMBL3933763 CHEMBL3933763 | C24H29F3N4O3 | 478.516 | 7 / 2 | 2.5 | Yes |
| 534102 | 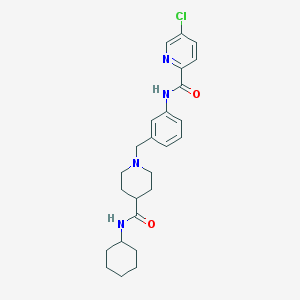 CHEMBL3890801 CHEMBL3890801 | C25H31ClN4O2 | 454.999 | 4 / 2 | 4.1 | Yes |
| 534107 | 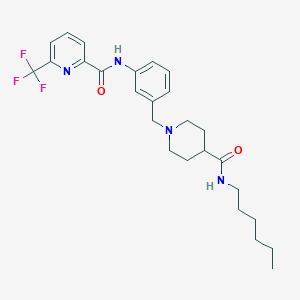 CHEMBL3919580 CHEMBL3919580 | C26H33F3N4O2 | 490.571 | 7 / 2 | 4.9 | Yes |
| 534108 | 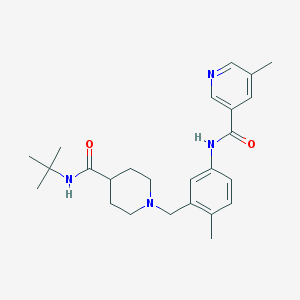 CHEMBL3972507 CHEMBL3972507 | C25H34N4O2 | 422.573 | 4 / 2 | 3.0 | Yes |
| 534110 | 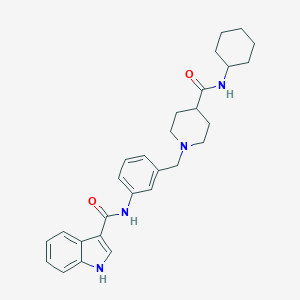 CHEMBL3983113 CHEMBL3983113 | C28H34N4O2 | 458.606 | 3 / 3 | 4.3 | Yes |
| 47289 | 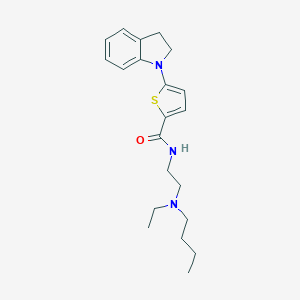 CHEMBL2381337 CHEMBL2381337 | C21H29N3OS | 371.543 | 4 / 1 | 4.9 | Yes |
| 534120 | 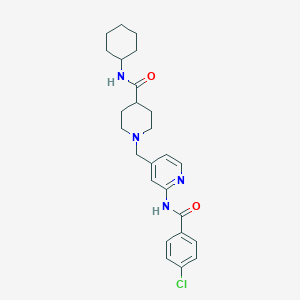 CHEMBL3980568 CHEMBL3980568 | C25H31ClN4O2 | 454.999 | 4 / 2 | 4.1 | Yes |
| 47860 | 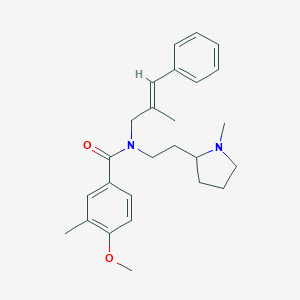 CHEMBL2013220 CHEMBL2013220 | C26H34N2O2 | 406.57 | 3 / 0 | 5.4 | No |
| 534128 | 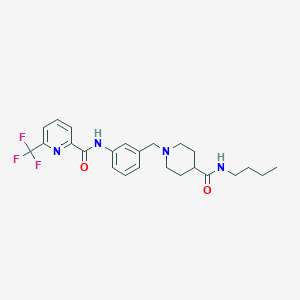 CHEMBL3974670 CHEMBL3974670 | C24H29F3N4O2 | 462.517 | 7 / 2 | 3.8 | Yes |
| 537223 | 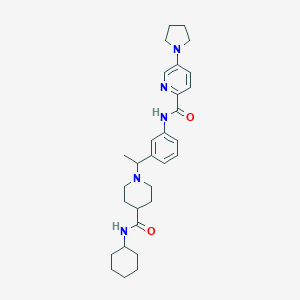 CHEMBL3923474 CHEMBL3923474 | C30H41N5O2 | 503.691 | 5 / 2 | 4.4 | No |
| 534129 | 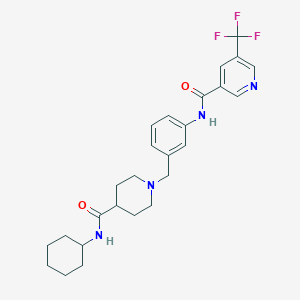 CHEMBL3946666 CHEMBL3946666 | C26H31F3N4O2 | 488.555 | 7 / 2 | 4.0 | Yes |
| 534141 | 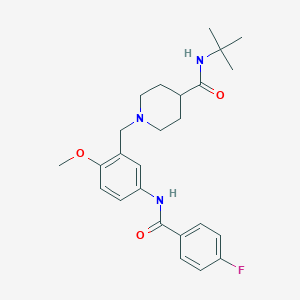 CHEMBL3982357 CHEMBL3982357 | C25H32FN3O3 | 441.547 | 5 / 2 | 3.4 | Yes |
| 534143 | 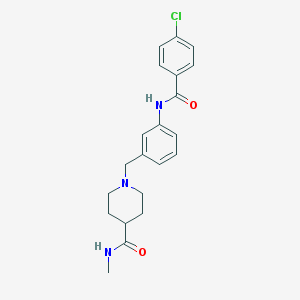 CHEMBL3911655 CHEMBL3911655 | C21H24ClN3O2 | 385.892 | 3 / 2 | 3.0 | Yes |
| 534146 | 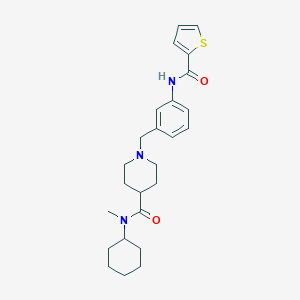 CHEMBL3947371 CHEMBL3947371 | C25H33N3O2S | 439.618 | 4 / 1 | 4.4 | Yes |
| 534147 | 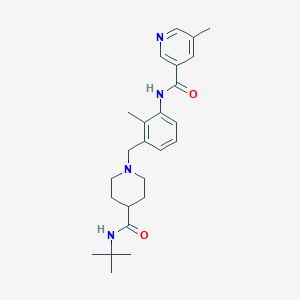 CHEMBL3986916 CHEMBL3986916 | C25H34N4O2 | 422.573 | 4 / 2 | 3.0 | Yes |
| 534148 | 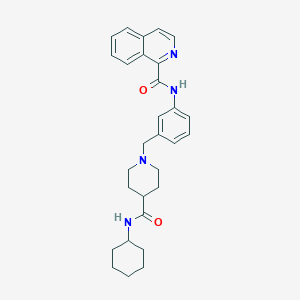 CHEMBL3902012 CHEMBL3902012 | C29H34N4O2 | 470.617 | 4 / 2 | 4.7 | Yes |
| 534151 | 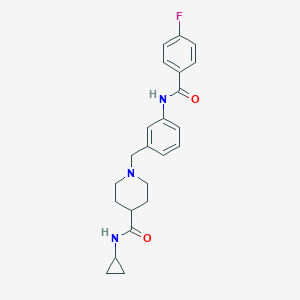 CHEMBL3905718 CHEMBL3905718 | C23H26FN3O2 | 395.478 | 4 / 2 | 3.0 | Yes |
| 534155 | 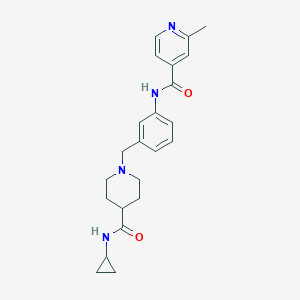 CHEMBL3899041 CHEMBL3899041 | C23H28N4O2 | 392.503 | 4 / 2 | 2.2 | Yes |
| 534156 | 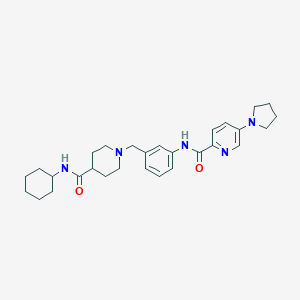 CHEMBL3935694 CHEMBL3935694 | C29H39N5O2 | 489.664 | 5 / 2 | 4.0 | Yes |
| 534161 | 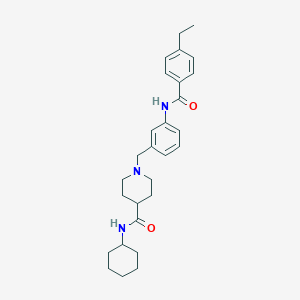 CHEMBL3925733 CHEMBL3925733 | C28H37N3O2 | 447.623 | 3 / 2 | 5.0 | Yes |
| 534163 | 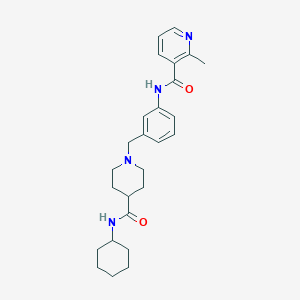 CHEMBL3942373 CHEMBL3942373 | C26H34N4O2 | 434.584 | 4 / 2 | 3.5 | Yes |
| 537411 | 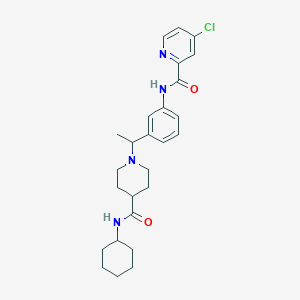 CHEMBL3984153 CHEMBL3984153 | C26H33ClN4O2 | 469.026 | 4 / 2 | 4.5 | Yes |
| 534174 | 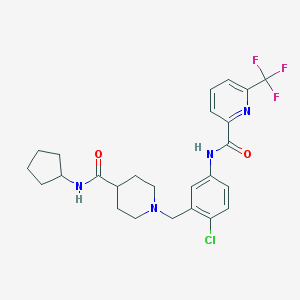 CHEMBL3966751 CHEMBL3966751 | C25H28ClF3N4O2 | 508.97 | 7 / 2 | 4.4 | No |
| 534175 | 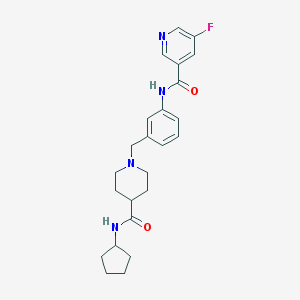 CHEMBL3957834 CHEMBL3957834 | C24H29FN4O2 | 424.52 | 5 / 2 | 2.7 | Yes |
| 534178 | 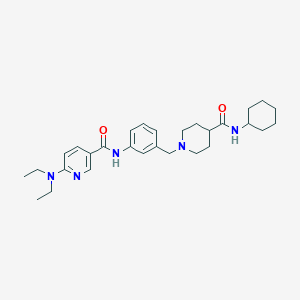 CHEMBL3968214 CHEMBL3968214 | C29H41N5O2 | 491.68 | 5 / 2 | 4.3 | Yes |
| 534182 | 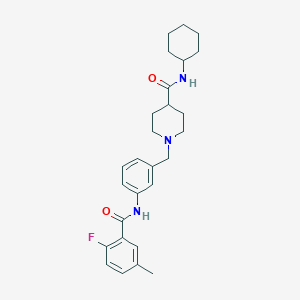 CHEMBL3956018 CHEMBL3956018 | C27H34FN3O2 | 451.586 | 4 / 2 | 4.6 | Yes |
| 534185 | 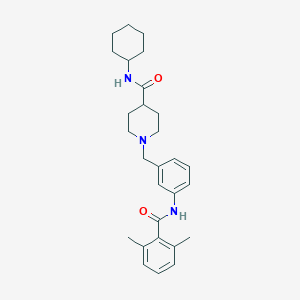 CHEMBL3922915 CHEMBL3922915 | C28H37N3O2 | 447.623 | 3 / 2 | 4.9 | Yes |
| 534186 | 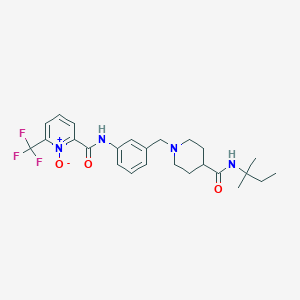 CHEMBL3982018 CHEMBL3982018 | C25H31F3N4O3 | 492.543 | 7 / 2 | 3.1 | Yes |
| 63141 | 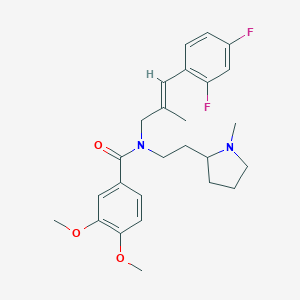 CHEMBL2013223 CHEMBL2013223 | C26H32F2N2O3 | 458.55 | 6 / 0 | 5.2 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218