

You can:
| Name | Type-1 angiotensin II receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | AGTR1 |
| Synonym | Type-1 angiotensin II receptor HAT1R Agtr-1a type-1A angiotensin II receptor AT1 [ Show all ] |
| Disease | Metabolic syndrome x Myocardial infarction Hypertension Restenosis Alzheimer disease [ Show all ] |
| Length | 359 |
| Amino acid sequence | MILNSSTEDGIKRIQDDCPKAGRHNYIFVMIPTLYSIIFVVGIFGNSLVVIVIYFYMKLKTVASVFLLNLALADLCFLLTLPLWAVYTAMEYRWPFGNYLCKIASASVSFNLYASVFLLTCLSIDRYLAIVHPMKSRLRRTMLVAKVTCIIIWLLAGLASLPAIIHRNVFFIENTNITVCAFHYESQNSTLPIGLGLTKNILGFLFPFLIILTSYTLIWKALKKAYEIQKNKPRNDDIFKIIMAIVLFFFFSWIPHQIFTFLDVLIQLGIIRDCRIADIVDTAMPITICIAYFNNCLNPLFYGFLGKKFKRYFLQLLKYIPPKAKSHSNLSTKMSTLSYRPSDNVSSSTKKPAPCFEVE |
| UniProt | P30556 |
| Protein Data Bank | 6do1, 4zud, 4yay |
| GPCR-HGmod model | P30556 |
| 3D structure model | This structure is from PDB ID 6do1. |
| BioLiP | BL0312790, BL0326733, BL0439004,BL0439005 |
| Therapeutic Target Database | T74456 |
| ChEMBL | CHEMBL227 |
| IUPHAR | 34 |
| DrugBank | BE0000062 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 102 | 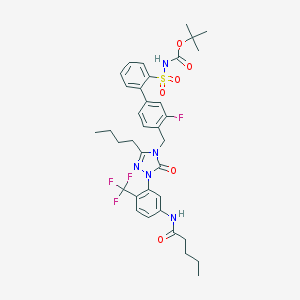 CHEMBL338434 CHEMBL338434 | C36H41F4N5O6S | 747.807 | 11 / 2 | 7.3 | No |
| 541 | 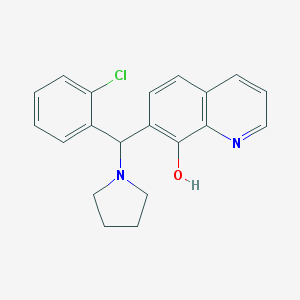 MLS000761483 MLS000761483 | C20H19ClN2O | 338.835 | 3 / 1 | 4.5 | Yes |
| 661 | 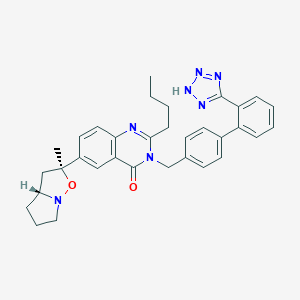 CHEMBL47177 CHEMBL47177 | C33H35N7O2 | 561.69 | 7 / 1 | 5.4 | No |
| 967 | 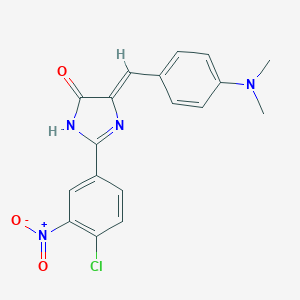 MLS000389682 MLS000389682 | C18H15ClN4O3 | 370.793 | 5 / 1 | 3.6 | Yes |
| 1344 | 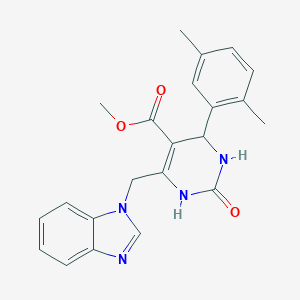 SR-01000300690 SR-01000300690 | C22H22N4O3 | 390.443 | 4 / 2 | 2.5 | Yes |
| 1405 | 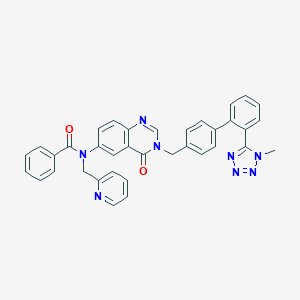 CHEMBL30086 CHEMBL30086 | C36H28N8O2 | 604.674 | 7 / 0 | 4.8 | No |
| 1548 | 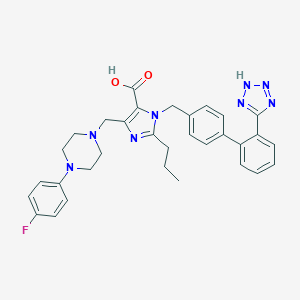 CHEMBL306216 CHEMBL306216 | C32H33FN8O2 | 580.668 | 9 / 2 | 2.8 | No |
| 1743 | 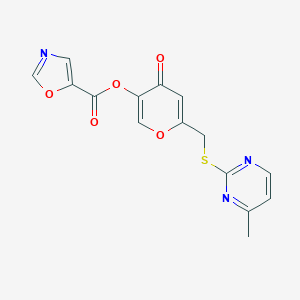 CHEMBL2158341 CHEMBL2158341 | C15H11N3O5S | 345.329 | 9 / 0 | 1.5 | Yes |
| 1985 | 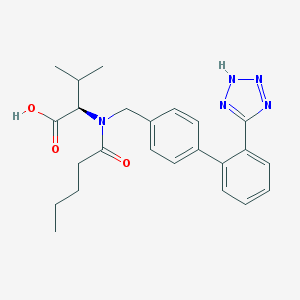 137862-87-4 137862-87-4 | C24H29N5O3 | 435.528 | 6 / 2 | 4.4 | Yes |
| 1986 | 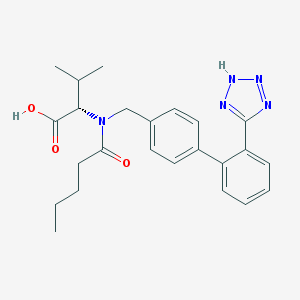 valsartan valsartan | C24H29N5O3 | 435.528 | 6 / 2 | 4.4 | Yes |
| 2117 | 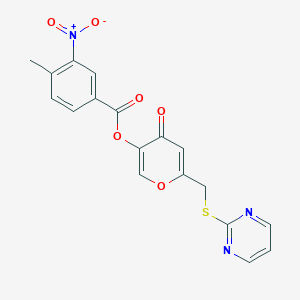 CHEMBL2158345 CHEMBL2158345 | C18H13N3O6S | 399.377 | 9 / 0 | 2.6 | Yes |
| 2229 | 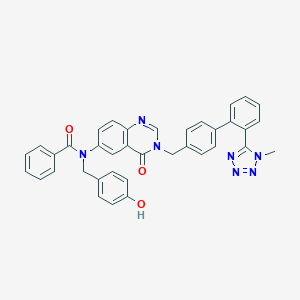 CHEMBL30015 CHEMBL30015 | C37H29N7O3 | 619.685 | 7 / 1 | 5.5 | No |
| 3204 | 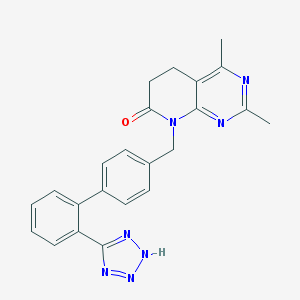 TASOSARTAN TASOSARTAN | C23H21N7O | 411.469 | 6 / 1 | 3.0 | Yes |
| 3538 | 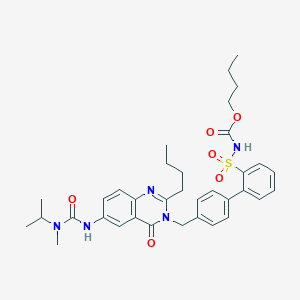 CHEMBL74476 CHEMBL74476 | C35H43N5O6S | 661.818 | 7 / 2 | 5.8 | No |
| 3847 | 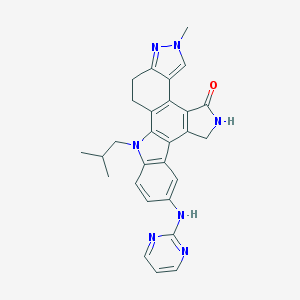 CEP-11981 CEP-11981 | C28H27N7O | 477.572 | 5 / 2 | 3.9 | Yes |
| 3933 | 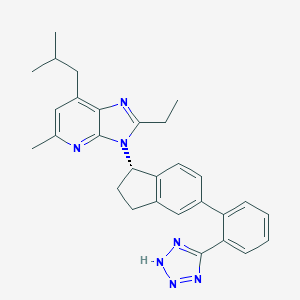 CHEMBL1801742 CHEMBL1801742 | C29H31N7 | 477.616 | 5 / 1 | 6.4 | No |
| 4693 | 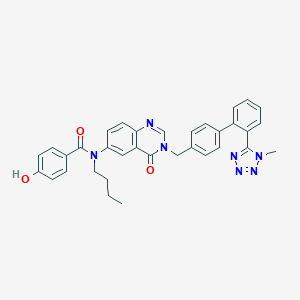 CHEMBL282168 CHEMBL282168 | C34H31N7O3 | 585.668 | 7 / 1 | 5.2 | No |
| 4914 | 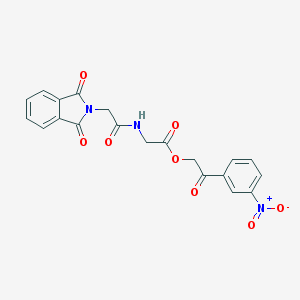 MLS000681499 MLS000681499 | C20H15N3O8 | 425.353 | 8 / 1 | 1.4 | Yes |
| 5329 | 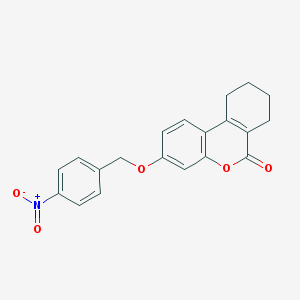 3-(4-Nitro-benzyloxy)-7,8,9,10-tetrahydro-benzo[c]chromen-6-one 3-(4-Nitro-benzyloxy)-7,8,9,10-tetrahydro-benzo[c]chromen-6-one | C20H17NO5 | 351.358 | 5 / 0 | 4.1 | Yes |
| 5431 | 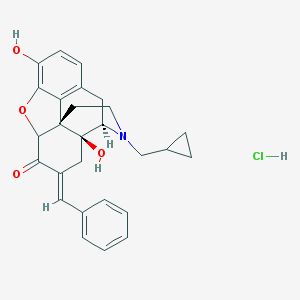 MLS002320750 MLS002320750 | C27H28ClNO4 | 465.974 | 5 / 3 | N/A | N/A |
| 6043 | 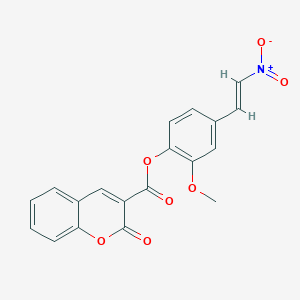 MLS001206870 MLS001206870 | C19H13NO7 | 367.313 | 7 / 0 | 3.8 | Yes |
| 6083 | 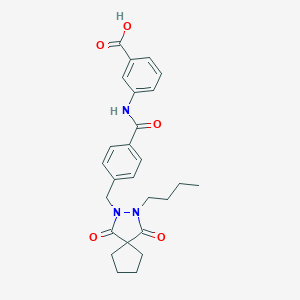 CHEMBL91069 CHEMBL91069 | C26H29N3O5 | 463.534 | 5 / 2 | 3.9 | Yes |
| 6592 | 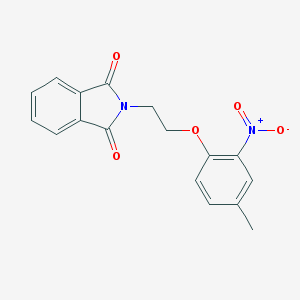 MLS000699844 MLS000699844 | C17H14N2O5 | 326.308 | 5 / 0 | 2.8 | Yes |
| 7054 | 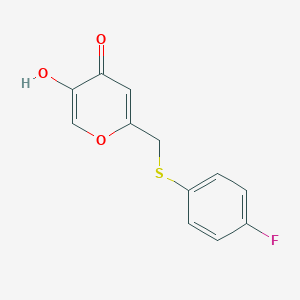 CHEMBL2158321 CHEMBL2158321 | C12H9FO3S | 252.259 | 5 / 1 | 2.1 | Yes |
| 7167 | 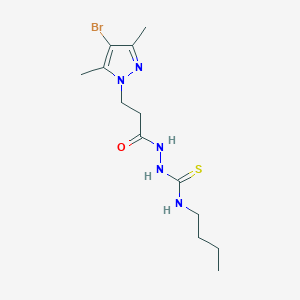 SMR000160170 SMR000160170 | C13H22BrN5OS | 376.317 | 3 / 3 | 2.1 | Yes |
| 8787 | 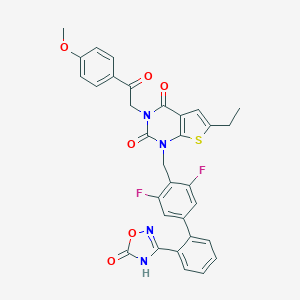 SCHEMBL2774146 SCHEMBL2774146 | C32H24F2N4O6S | 630.623 | 10 / 1 | 5.9 | No |
| 547983 | 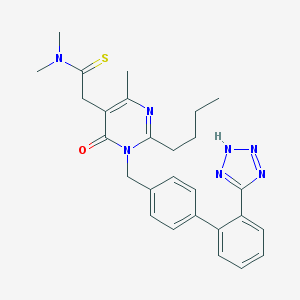 FIMASARTAN FIMASARTAN | C27H31N7OS | 501.653 | 6 / 1 | 3.5 | No |
| 9026 | 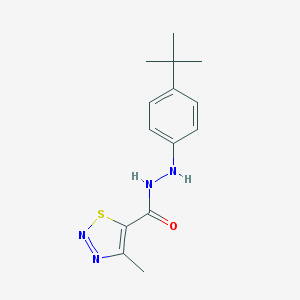 MLS000860444 MLS000860444 | C14H18N4OS | 290.385 | 5 / 2 | 4.0 | Yes |
| 9421 | 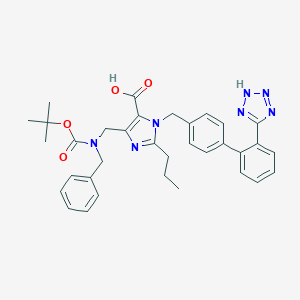 CHEMBL306922 CHEMBL306922 | C34H37N7O4 | 607.715 | 8 / 2 | 5.7 | No |
| 9567 | 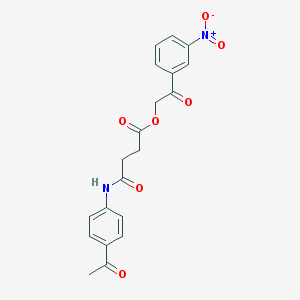 MLS000923770 MLS000923770 | C20H18N2O7 | 398.371 | 7 / 1 | 1.8 | Yes |
| 9599 | 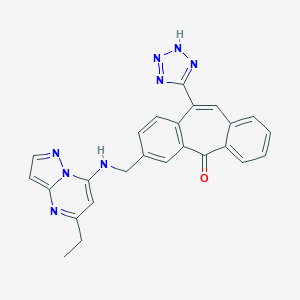 CHEMBL90513 CHEMBL90513 | C25H20N8O | 448.49 | 7 / 2 | 3.7 | Yes |
| 10035 | 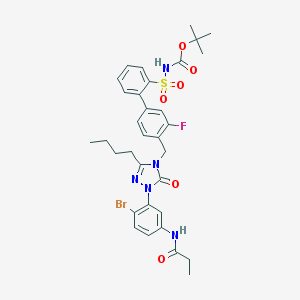 CHEMBL124771 CHEMBL124771 | C33H37BrFN5O6S | 730.65 | 8 / 2 | 6.2 | No |
| 10686 | 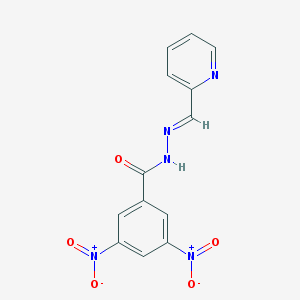 MLS000720075 MLS000720075 | C13H9N5O5 | 315.245 | 7 / 1 | 1.8 | Yes |
| 10941 | 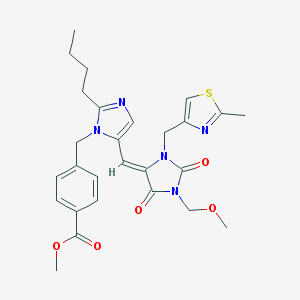 CHEMBL125835 CHEMBL125835 | C27H31N5O5S | 537.635 | 8 / 0 | 3.7 | No |
| 11015 | 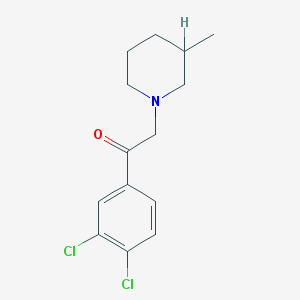 MLS001033014 MLS001033014 | C14H17Cl2NO | 286.196 | 2 / 0 | 4.2 | Yes |
| 521755 | 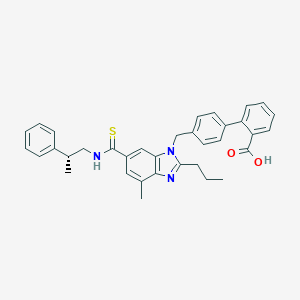 CHEMBL3734859 CHEMBL3734859 | C35H35N3O2S | 561.744 | 4 / 2 | 7.9 | No |
| 521756 | 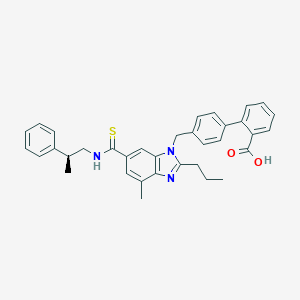 CHEMBL3735831 CHEMBL3735831 | C35H35N3O2S | 561.744 | 4 / 2 | 7.9 | No |
| 11142 | 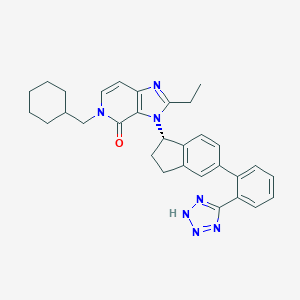 CHEMBL2322442 CHEMBL2322442 | C31H33N7O | 519.653 | 5 / 1 | 6.0 | No |
| 521760 | 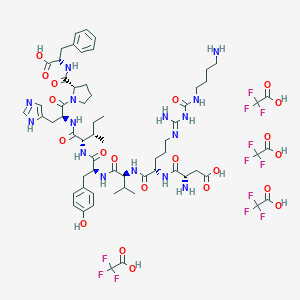 CHEMBL3786145 CHEMBL3786145 | C63H85F12N15O21 | 1616.44 | 37 / 19 | N/A | No |
| 521786 | 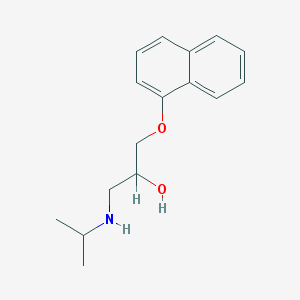 propranolol propranolol | C16H21NO2 | 259.349 | 3 / 2 | 3.0 | Yes |
| 12482 | 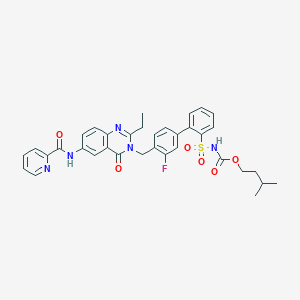 CHEMBL74144 CHEMBL74144 | C35H34FN5O6S | 671.744 | 9 / 2 | 5.5 | No |
| 12947 | 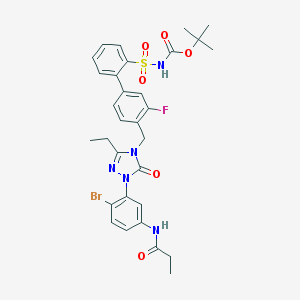 CHEMBL122157 CHEMBL122157 | C31H33BrFN5O6S | 702.596 | 8 / 2 | 5.3 | No |
| 14079 | 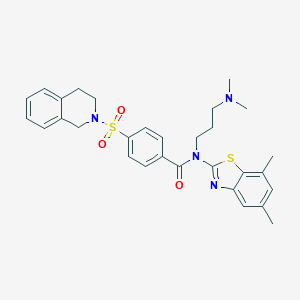 MLS001239249 MLS001239249 | C30H34N4O3S2 | 562.747 | 7 / 0 | 5.5 | No |
| 14361 | 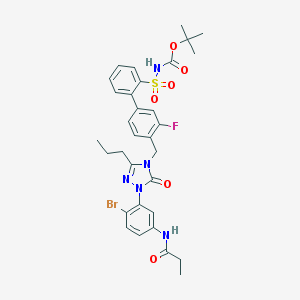 CHEMBL436396 CHEMBL436396 | C32H35BrFN5O6S | 716.623 | 8 / 2 | 5.6 | No |
| 14370 | 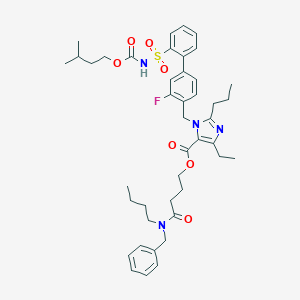 CHEMBL70868 CHEMBL70868 | C43H55FN4O7S | 790.992 | 9 / 1 | 8.9 | No |
| 14604 | 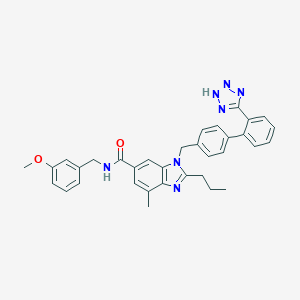 CHEMBL2058856 CHEMBL2058856 | C34H33N7O2 | 571.685 | 6 / 2 | 6.2 | No |
| 15019 | 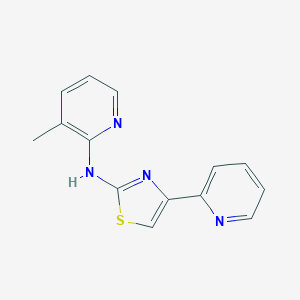 MLS000528306 MLS000528306 | C14H12N4S | 268.338 | 5 / 1 | 3.1 | Yes |
| 15046 | 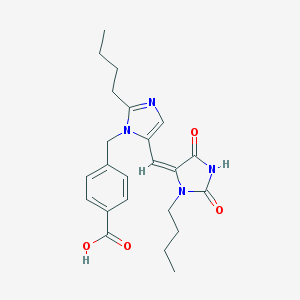 CHEMBL125766 CHEMBL125766 | C23H28N4O4 | 424.501 | 5 / 2 | 3.5 | Yes |
| 15047 | 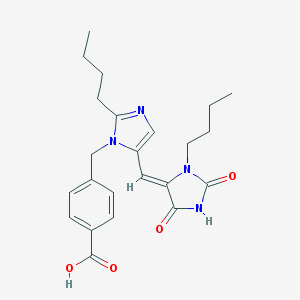 CHEMBL125937 CHEMBL125937 | C23H28N4O4 | 424.501 | 5 / 2 | 3.5 | Yes |
| 15096 | 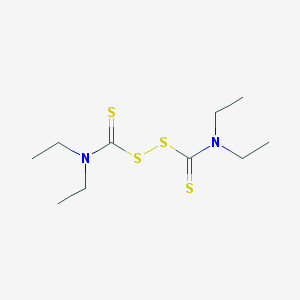 disulfiram disulfiram | C10H20N2S4 | 296.524 | 4 / 0 | 3.9 | Yes |
| 15504 | 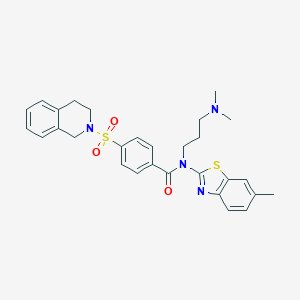 MLS001239478 MLS001239478 | C29H32N4O3S2 | 548.72 | 7 / 0 | 5.2 | No |
| 15539 | 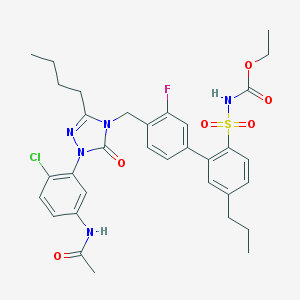 CHEMBL321313 CHEMBL321313 | C33H37ClFN5O6S | 686.196 | 8 / 2 | 6.4 | No |
| 15835 | 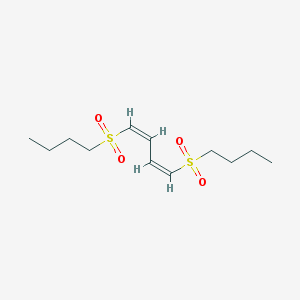 (1Z,3Z)-1,4-bis(butylsulfonyl)buta-1,3-diene (1Z,3Z)-1,4-bis(butylsulfonyl)buta-1,3-diene | C12H22O4S2 | 294.424 | 4 / 0 | 2.1 | Yes |
| 17180 | 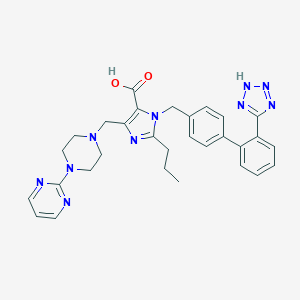 CHEMBL309165 CHEMBL309165 | C30H32N10O2 | 564.654 | 10 / 2 | 1.3 | No |
| 521988 | 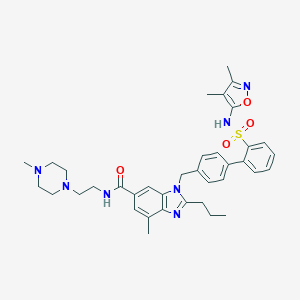 CHEMBL3735236 CHEMBL3735236 | C37H45N7O4S | 683.872 | 9 / 2 | 5.4 | No |
| 17619 | 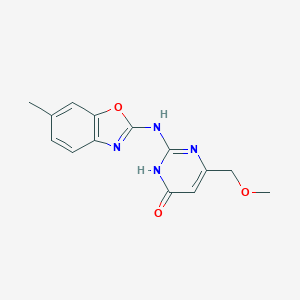 MLS000039260 MLS000039260 | C14H14N4O3 | 286.291 | 5 / 2 | 0.9 | Yes |
| 18767 | 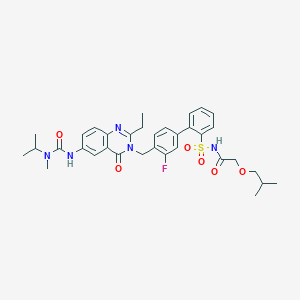 CHEMBL311386 CHEMBL311386 | C34H40FN5O6S | 665.781 | 8 / 2 | 4.5 | No |
| 19081 | 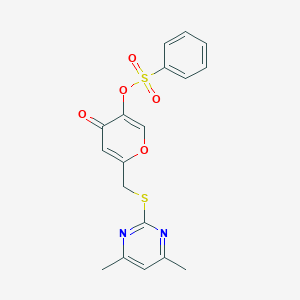 CHEMBL2158335 CHEMBL2158335 | C18H16N2O5S2 | 404.455 | 8 / 0 | 2.9 | Yes |
| 522040 | 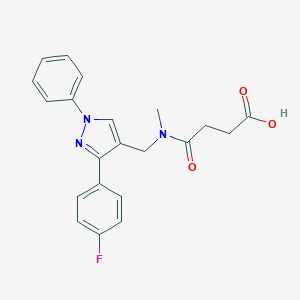 CHEMBL1716544 CHEMBL1716544 | C21H20FN3O3 | 381.407 | 5 / 1 | 2.4 | Yes |
| 19895 | 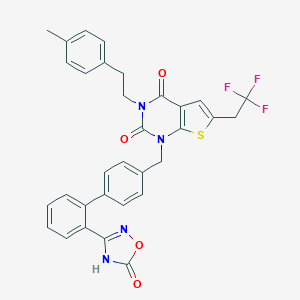 SCHEMBL2773794 SCHEMBL2773794 | C32H25F3N4O4S | 618.631 | 9 / 1 | 7.2 | No |
| 19957 | 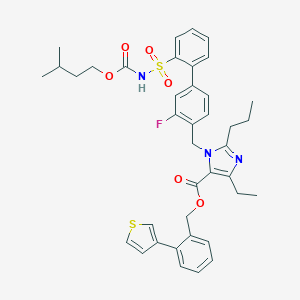 CHEMBL74236 CHEMBL74236 | C39H42FN3O6S2 | 731.898 | 9 / 1 | 9.2 | No |
| 19969 | 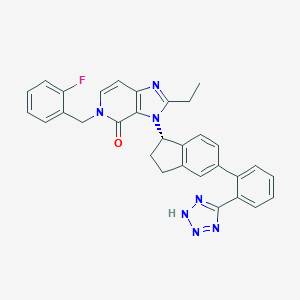 CHEMBL2322443 CHEMBL2322443 | C31H26FN7O | 531.595 | 6 / 1 | 5.3 | No |
| 20871 | 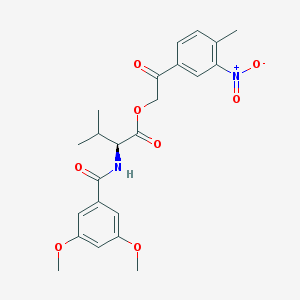 MLS002245453 MLS002245453 | C23H26N2O8 | 458.467 | 8 / 1 | 4.0 | Yes |
| 20946 | 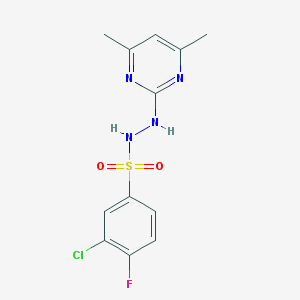 MLS001098424 MLS001098424 | C12H12ClFN4O2S | 330.762 | 7 / 2 | 2.9 | Yes |
| 20962 | 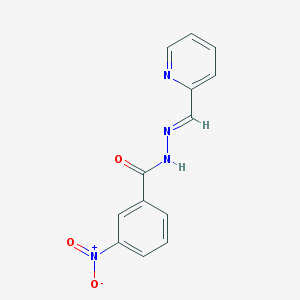 MLS000723451 MLS000723451 | C13H10N4O3 | 270.248 | 5 / 1 | 1.9 | Yes |
| 21088 | 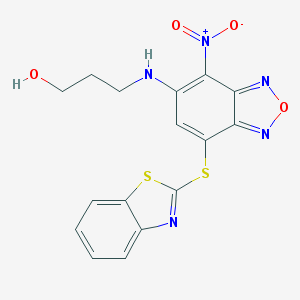 MLS000520994 MLS000520994 | C16H13N5O4S2 | 403.431 | 10 / 2 | 4.1 | Yes |
| 21577 | 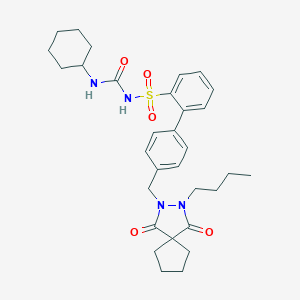 CHEMBL315873 CHEMBL315873 | C31H40N4O5S | 580.744 | 5 / 2 | 6.1 | No |
| 21674 | 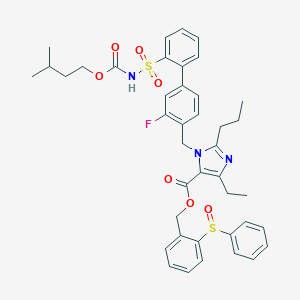 CHEMBL73283 CHEMBL73283 | C41H44FN3O7S2 | 773.935 | 10 / 1 | 8.7 | No |
| 21985 | 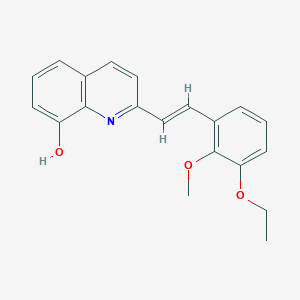 MLS001196044 MLS001196044 | C20H19NO3 | 321.376 | 4 / 1 | 4.4 | Yes |
| 522136 | 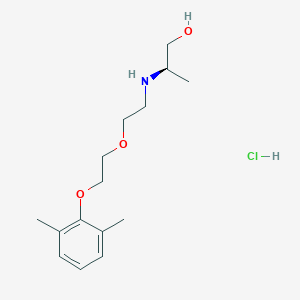 CHEMBL3775436 CHEMBL3775436 | C15H26ClNO3 | 303.827 | 4 / 3 | N/A | N/A |
| 522199 | 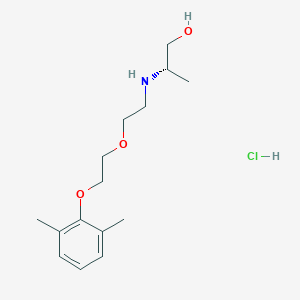 CHEMBL3775769 CHEMBL3775769 | C15H26ClNO3 | 303.827 | 4 / 3 | N/A | N/A |
| 22165 | 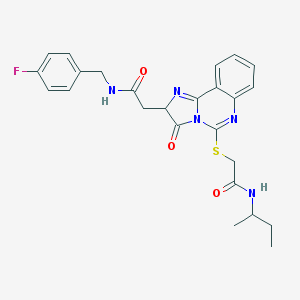 MLS000094900 MLS000094900 | C25H26FN5O3S | 495.573 | 7 / 2 | 3.0 | Yes |
| 22570 | 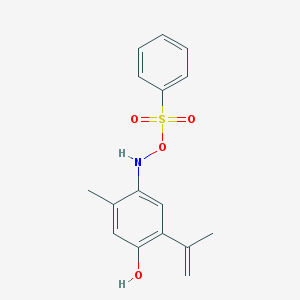 BDBM46978 BDBM46978 | C16H17NO4S | 319.375 | 5 / 2 | 4.5 | Yes |
| 23792 | 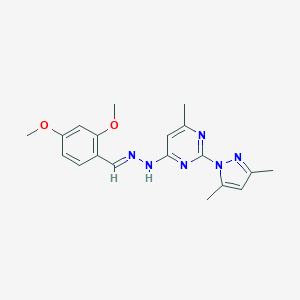 4-[(2E)-2-(2,4-dimethoxybenzylidene)hydrazinyl]-2-(3,5-dimethyl-1H-pyrazol-1-yl)-6-methylpyrimidine 4-[(2E)-2-(2,4-dimethoxybenzylidene)hydrazinyl]-2-(3,5-dimethyl-1H-pyrazol-1-yl)-6-methylpyrimidine | C19H22N6O2 | 366.425 | 7 / 1 | 3.6 | Yes |
| 24919 | 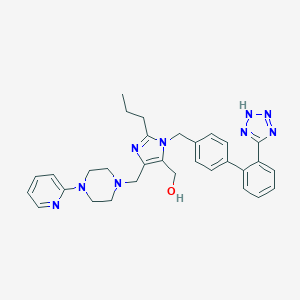 CHEMBL72409 CHEMBL72409 | C31H35N9O | 549.683 | 8 / 2 | 3.5 | No |
| 25201 | 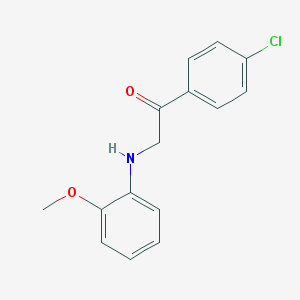 MLS001173828 MLS001173828 | C15H14ClNO2 | 275.732 | 3 / 1 | 3.8 | Yes |
| 522328 | 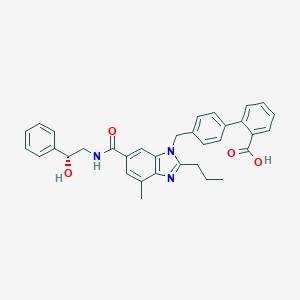 CHEMBL3734819 CHEMBL3734819 | C34H33N3O4 | 547.655 | 5 / 3 | 5.9 | No |
| 522329 | 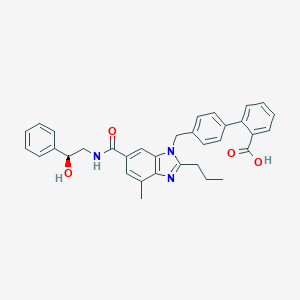 CHEMBL3735333 CHEMBL3735333 | C34H33N3O4 | 547.655 | 5 / 3 | 5.9 | No |
| 25592 | 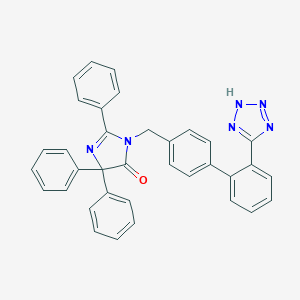 CHEMBL39542 CHEMBL39542 | C35H26N6O | 546.634 | 5 / 1 | 6.6 | No |
| 25597 | 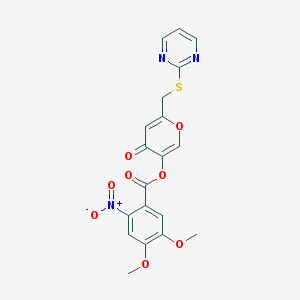 CHEMBL2158271 CHEMBL2158271 | C19H15N3O8S | 445.402 | 11 / 0 | 2.1 | No |
| 25849 | 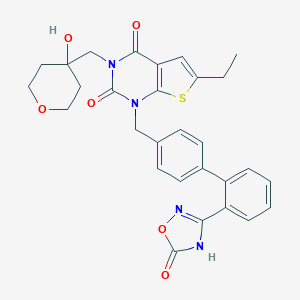 SCHEMBL2775820 SCHEMBL2775820 | C29H28N4O6S | 560.625 | 8 / 2 | 3.8 | No |
| 558075 | 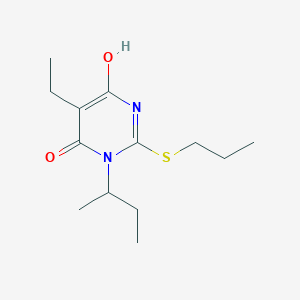 MLS000058898 MLS000058898 | C13H22N2O2S | 270.391 | 4 / 1 | 3.3 | Yes |
| 442715 | 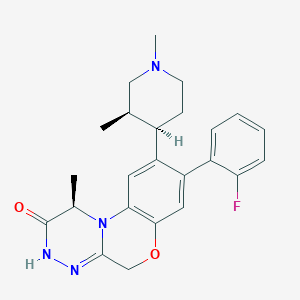 CHEMBL3355127 CHEMBL3355127 | C24H27FN4O2 | 422.504 | 5 / 1 | 3.6 | Yes |
| 27224 | 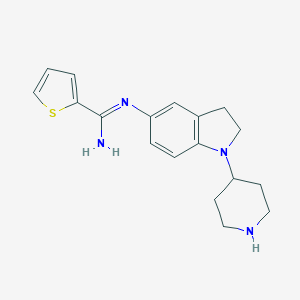 CHEMBL2203713 CHEMBL2203713 | C18H22N4S | 326.462 | 4 / 2 | 2.9 | Yes |
| 27278 | 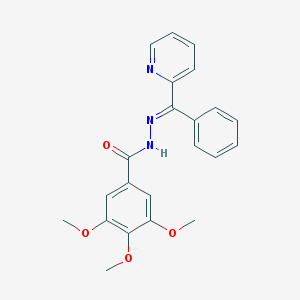 MLS000390992 MLS000390992 | C22H21N3O4 | 391.427 | 6 / 1 | 3.9 | Yes |
| 27536 | 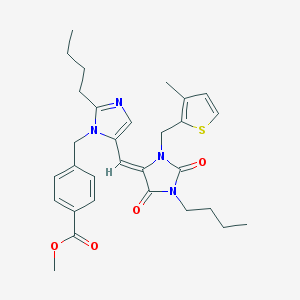 CHEMBL338123 CHEMBL338123 | C30H36N4O4S | 548.702 | 6 / 0 | 5.6 | No |
| 28059 | 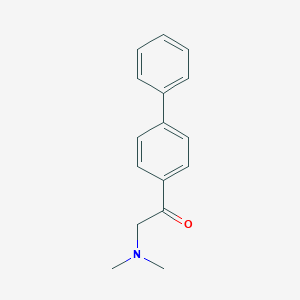 SMR000103373 SMR000103373 | C16H17NO | 239.318 | 2 / 0 | 3.8 | Yes |
| 28384 | 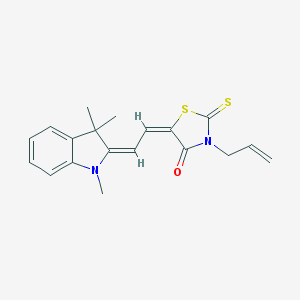 MLS000584397 MLS000584397 | C19H20N2OS2 | 356.502 | 4 / 0 | 5.0 | Yes |
| 28560 | 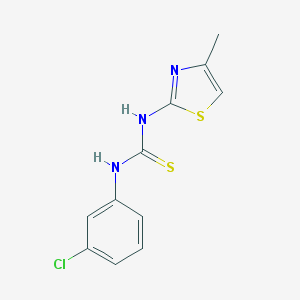 MLS000698760 MLS000698760 | C11H10ClN3S2 | 283.792 | 3 / 2 | 3.5 | Yes |
| 28787 | 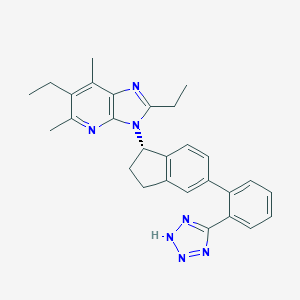 CHEMBL1801737 CHEMBL1801737 | C28H29N7 | 463.589 | 5 / 1 | 6.0 | No |
| 29561 | 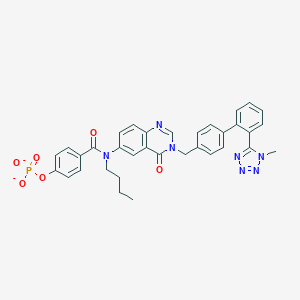 CHEMBL281890 CHEMBL281890 | C34H30N7O6P-2 | 663.631 | 10 / 0 | 3.8 | No |
| 29587 | 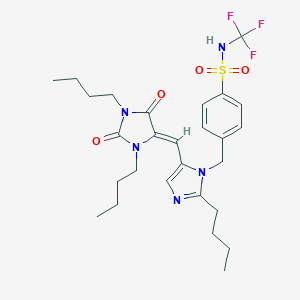 CHEMBL2259777 CHEMBL2259777 | C27H36F3N5O4S | 583.671 | 9 / 1 | 5.6 | No |
| 30560 | 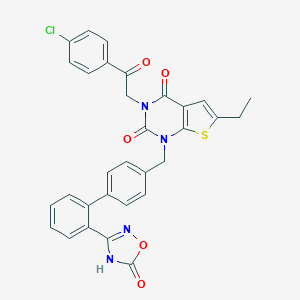 SCHEMBL2774156 SCHEMBL2774156 | C31H23ClN4O5S | 599.058 | 7 / 1 | 6.3 | No |
| 30568 | 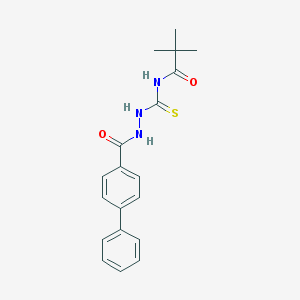 MLS000679816 MLS000679816 | C19H21N3O2S | 355.456 | 3 / 3 | 4.5 | Yes |
| 30583 | 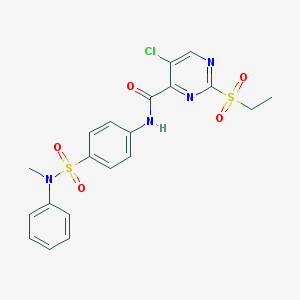 MLS001116397 MLS001116397 | C20H19ClN4O5S2 | 494.965 | 8 / 1 | 2.6 | Yes |
| 30610 | 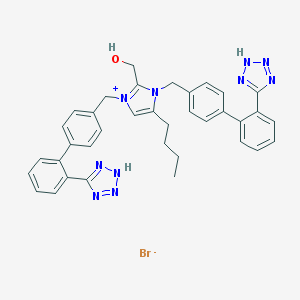 CHEMBL2337677 CHEMBL2337677 | C36H35BrN10O | 703.649 | 8 / 3 | N/A | No |
| 30689 | 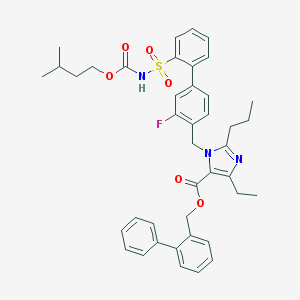 CHEMBL70793 CHEMBL70793 | C41H44FN3O6S | 725.876 | 8 / 1 | 9.5 | No |
| 30850 | 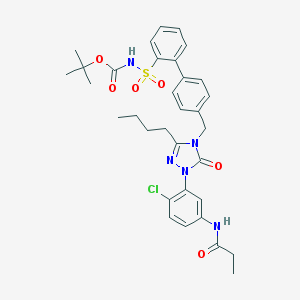 CHEMBL334300 CHEMBL334300 | C33H38ClN5O6S | 668.206 | 7 / 2 | 6.0 | No |
| 536787 | 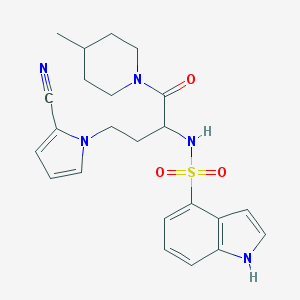 CHEMBL3889627 CHEMBL3889627 | C23H27N5O3S | 453.561 | 5 / 2 | 2.6 | Yes |
| 31203 | 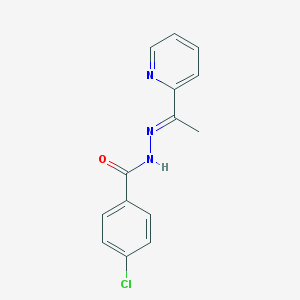 MLS001207850 MLS001207850 | C14H12ClN3O | 273.72 | 3 / 1 | 2.9 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218