

You can:
| Name | Somatostatin receptor type 2 |
|---|---|
| Species | Mus musculus (Mouse) |
| Gene | Sstr2 |
| Synonym | somatotropin release-inhibiting factor receptor SRIF-1 SS-2-R SS2-R SS2R [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 369 |
| Amino acid sequence | MEMSSEQLNGSQVWVSSPFDLNGSLGPSNGSNQTEPYYDMTSNAVLTFIYFVVCVVGLCGNTLVIYVILRYAKMKTITNIYILNLAIADELFMLGLPFLAMQVALVHWPFGKAICRVVMTVDGINQFTSIFCLTVMSIDRYLAVVHPIKSAKWRRPRTAKMINVAVWCVSLLVILPIMIYAGLRSNQWGRSSCTINWPGESGAWYTGFIIYAFILGFLVPLTIICLCYLFIIIKVKSSGIRVGSSKRKKSEKKVTRMVSIVVAVFIFCWLPFYIFNVSSVSVAISPTPALKGMFDFVVILTYANSCANPILYAFLSDNFKKSFQNVLCLVKVSGTEDGERSDSKQDKSRLNETTETQRTLLNGDLQTSI |
| UniProt | P30875 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3207 |
| IUPHAR | 356 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 555548 | 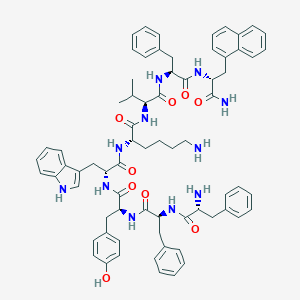 BDBM84629 BDBM84629 | C71H81N11O9 | 1232.5 | 11 / 12 | 8.1 | No |
| 555554 | 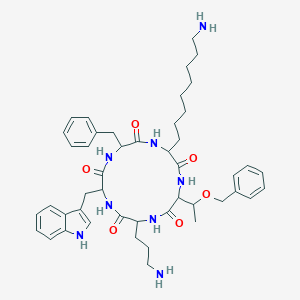 SA V SA V | C46H62N8O6 | 823.052 | 8 / 8 | 4.6 | No |
| 555593 | 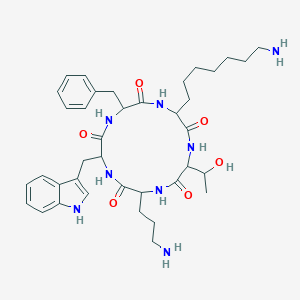 SA II SA II | C38H54N8O6 | 718.9 | 8 / 9 | 2.6 | No |
| 555627 | 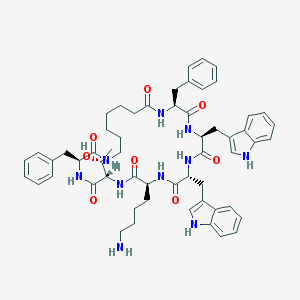 L-362855 L-362855 | C57H70N10O8 | 1023.25 | 9 / 11 | 5.7 | No |
| 555651 | 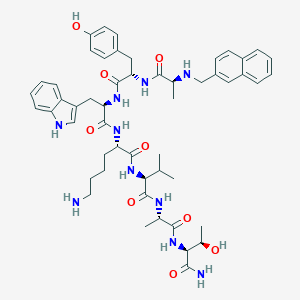 BDBM82458 BDBM82458 | C52H68N10O9 | 977.177 | 11 / 12 | 3.4 | No |
| 555663 | 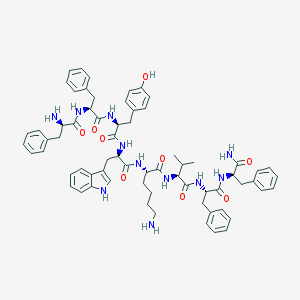 CHEMBL268112 CHEMBL268112 | C67H79N11O9 | 1182.44 | 11 / 12 | 5.6 | No |
| 555674 | 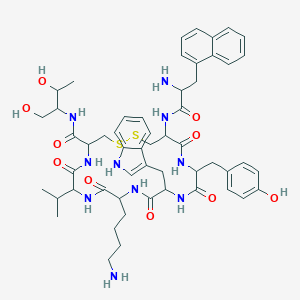 BDBM84620 BDBM84620 | C54H70N10O10S2 | 1083.33 | 14 / 13 | 2.9 | No |
| 443978 | 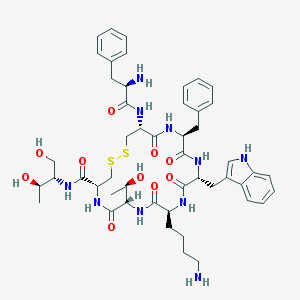 CHEMBL3350037 CHEMBL3350037 | C49H66N10O10S2 | 1019.25 | 14 / 13 | 1.0 | No |
| 555692 | 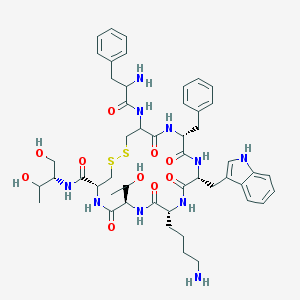 Octreotide Octreotide | C49H66N10O10S2 | 1019.25 | 14 / 13 | 1.0 | No |
| 58033 | 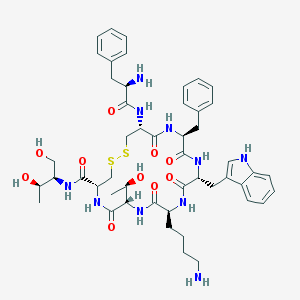 Octreotide Octreotide | C49H66N10O10S2 | 1019.25 | 14 / 13 | 1.0 | No |
| 553532 | 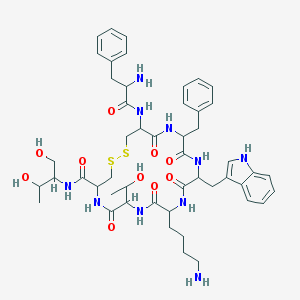 Octreotide Octreotide | C49H66N10O10S2 | 1019.25 | 14 / 13 | 1.0 | No |
| 555695 | 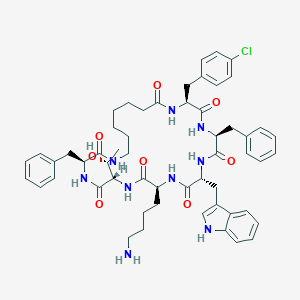 BDBM82466 BDBM82466 | C55H68ClN9O8 | 1018.65 | 9 / 10 | 6.2 | No |
| 555710 | 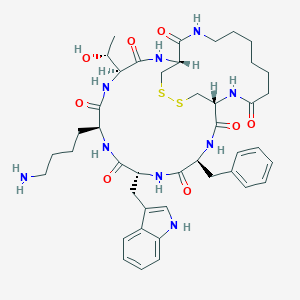 CHEMBL2369699 CHEMBL2369699 | C43H59N9O8S2 | 894.12 | 11 / 10 | 2.1 | No |
| 555728 | 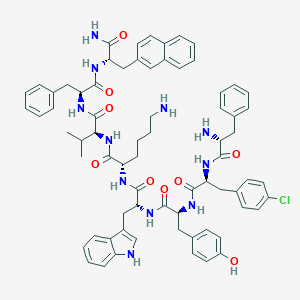 BDBM84640 BDBM84640 | C71H80ClN11O9 | 1266.94 | 11 / 12 | 8.8 | No |
| 68359 | 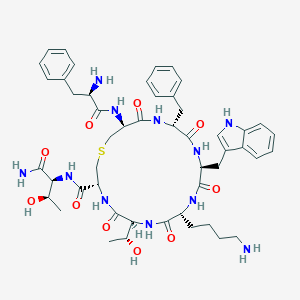 CHEMBL436783 CHEMBL436783 | C49H65N11O10S | 1000.19 | 13 / 13 | 0.5 | No |
| 555774 | 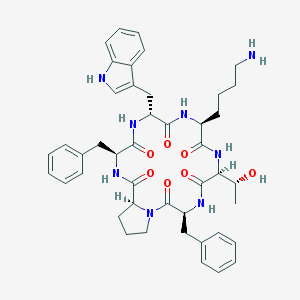 CHEMBL421362 CHEMBL421362 | C44H54N8O7 | 806.965 | 8 / 8 | 3.4 | No |
| 93067 | 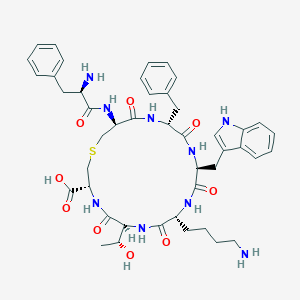 CHEMBL69379 CHEMBL69379 | C45H57N9O9S | 900.065 | 12 / 11 | -0.2 | No |
| 555827 | 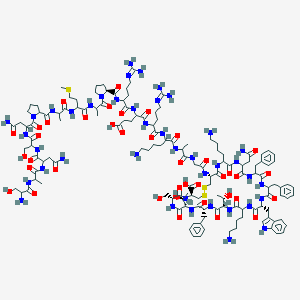 BDBM82460 BDBM82460 | C137H207N41O39S3 | 3148.59 | 48 / 44 | -16.0 | No |
| 553729 | 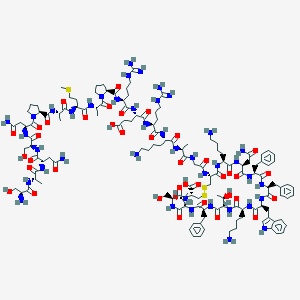 Somatostatin 28 Somatostatin 28 | C137H207N41O39S3 | 3148.59 | 48 / 46 | -15.2 | No |
| 555882 | 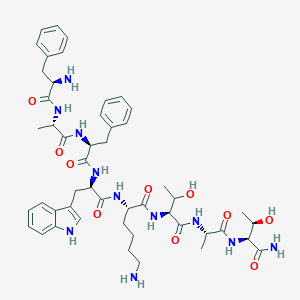 BDBM84625 BDBM84625 | C49H67N11O10 | 970.142 | 12 / 13 | -0.5 | No |
| 555884 | 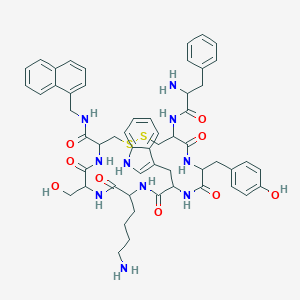 BDBM84623 BDBM84623 | C55H64N10O9S2 | 1073.3 | 13 / 12 | 3.8 | No |
| 114083 | 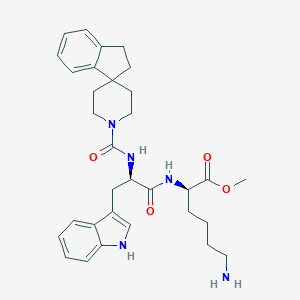 CHEMBL147319 CHEMBL147319 | C32H41N5O4 | 559.711 | 5 / 4 | 3.9 | No |
| 555927 | 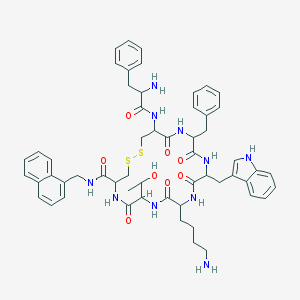 BDBM84628 BDBM84628 | C56H66N10O8S2 | 1071.33 | 12 / 11 | 4.6 | No |
| 555935 | 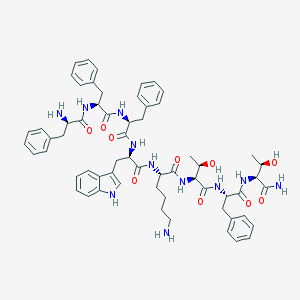 BIM 23052 BIM 23052 | C61H75N11O10 | 1122.34 | 12 / 13 | 2.7 | No |
| 555965 | 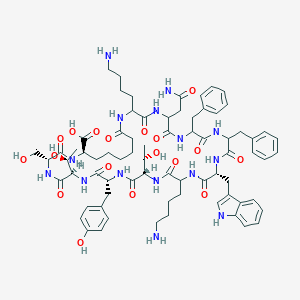 BDBM82467 BDBM82467 | C73H99N15O18 | 1474.68 | 20 / 20 | -0.8 | No |
| 555990 | 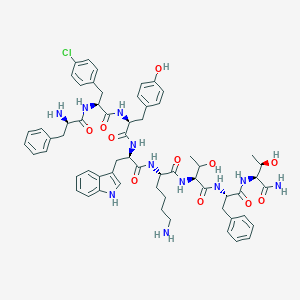 BDBM84624 BDBM84624 | C61H74ClN11O11 | 1172.78 | 13 / 14 | 3.7 | No |
| 128918 | 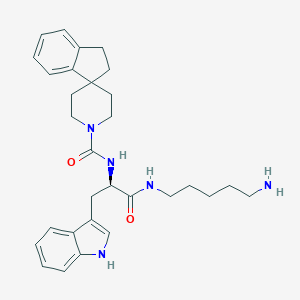 CHEMBL146128 CHEMBL146128 | C30H39N5O2 | 501.675 | 3 / 4 | 4.0 | No |
| 556042 | 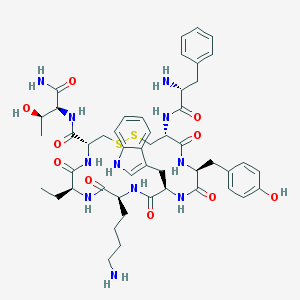 BIM-23023 BIM-23023 | C49H65N11O10S2 | 1032.25 | 14 / 13 | 0.8 | No |
| 556061 | 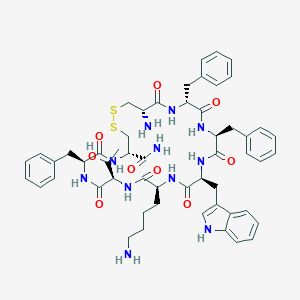 BIM 23268 BIM 23268 | C54H67N11O9S2 | 1078.32 | 13 / 12 | 2.8 | No |
| 556090 | 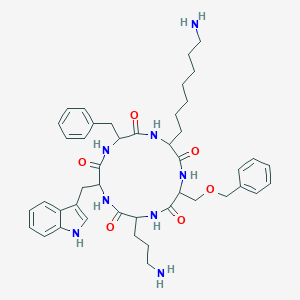 BDBM84644 BDBM84644 | C44H58N8O6 | 794.998 | 8 / 8 | 3.6 | No |
| 166626 | 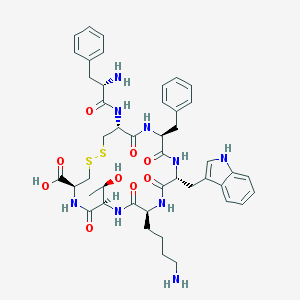 CHEMBL421493 CHEMBL421493 | C45H57N9O9S2 | 932.125 | 13 / 11 | -0.2 | No |
| 167139 | 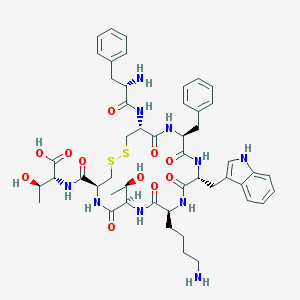 CHEMBL415860 CHEMBL415860 | C49H64N10O11S2 | 1033.23 | 15 / 13 | -1.1 | No |
| 556161 | 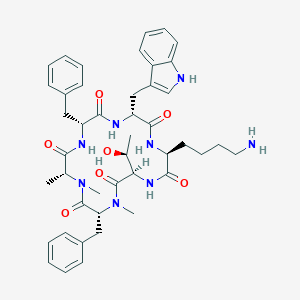 BDBM82454 BDBM82454 | C44H56N8O7 | 808.981 | 8 / 7 | 3.4 | No |
| 556163 | 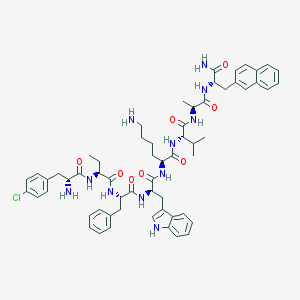 BDBM84617 BDBM84617 | C60H74ClN11O8 | 1112.77 | 10 / 11 | 6.5 | No |
| 556215 | 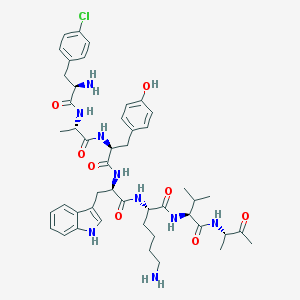 BDBM84616 BDBM84616 | C47H62ClN9O8 | 916.518 | 10 / 10 | 2.5 | No |
| 556216 | 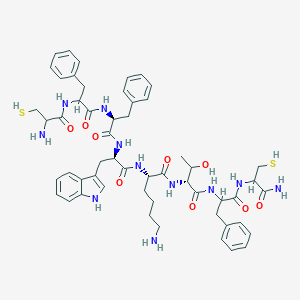 BDBM85230 BDBM85230 | C54H69N11O9S2 | 1080.33 | 13 / 14 | 2.3 | No |
| 186163 | 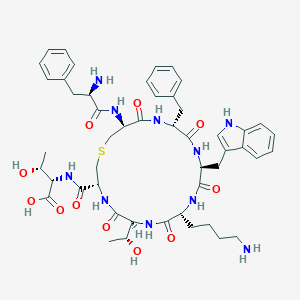 CHEMBL266469 CHEMBL266469 | C49H64N10O11S | 1001.17 | 14 / 13 | -1.1 | No |
| 556276 | 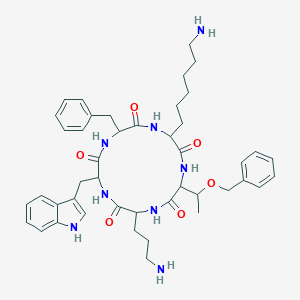 SA IV SA IV | C44H58N8O6 | 794.998 | 8 / 8 | 3.5 | No |
| 223705 | 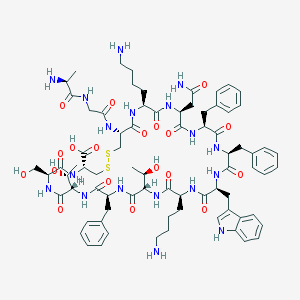 SOMATOSTATIN SOMATOSTATIN | C76H104N18O19S2 | 1637.9 | 24 / 22 | -3.1 | No |
| 556356 | 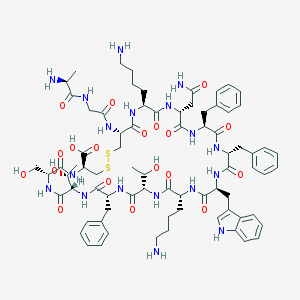 SOMATOSTATIN SOMATOSTATIN | C76H104N18O19S2 | 1637.9 | 24 / 22 | -3.1 | No |
| 554413 | 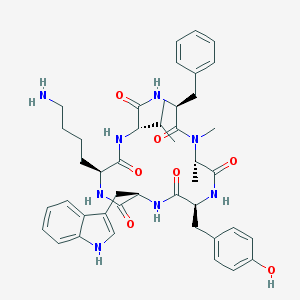 seglitide seglitide | C44H56N8O7 | 808.981 | 8 / 8 | 3.9 | No |
| 556374 | 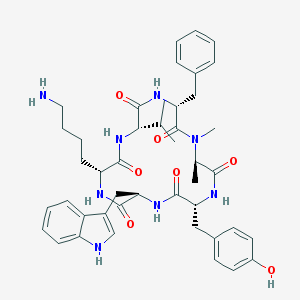 CHEMBL413535 CHEMBL413535 | C44H56N8O7 | 808.981 | 8 / 8 | 3.9 | No |
| 556376 | 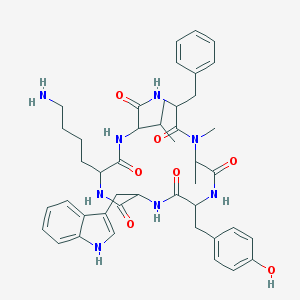 BDBM81766 BDBM81766 | C44H56N8O7 | 808.981 | 8 / 8 | 3.9 | No |
| 556482 | 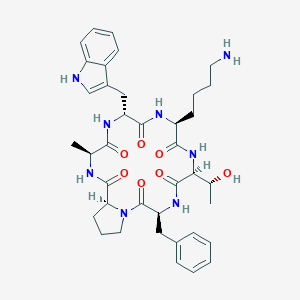 CHEMBL3349593 CHEMBL3349593 | C38H50N8O7 | 730.867 | 8 / 8 | 1.8 | No |
| 556490 | 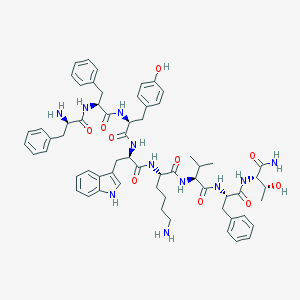 BIM-23058 BIM-23058 | C62H77N11O10 | 1136.36 | 12 / 13 | 4.7 | No |
| 556522 | 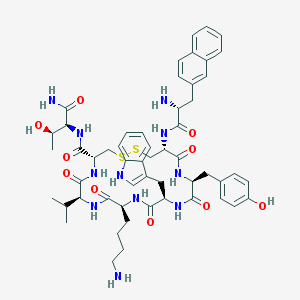 Lanreotide Lanreotide | C54H69N11O10S2 | 1096.33 | 14 / 13 | 2.5 | No |
| 554605 | 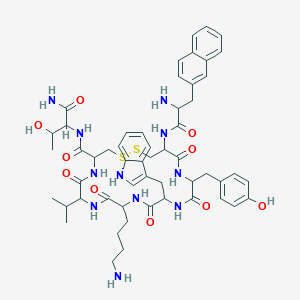 Lanreotide acetate Lanreotide acetate | C54H69N11O10S2 | 1096.33 | 14 / 13 | 2.5 | No |
| 556546 | 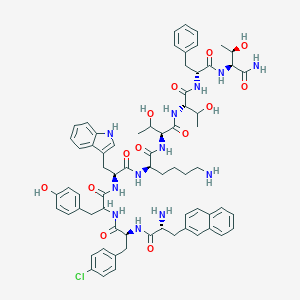 BDBM84619 BDBM84619 | C69H83ClN12O13 | 1323.94 | 15 / 16 | 4.1 | No |
| 556617 | 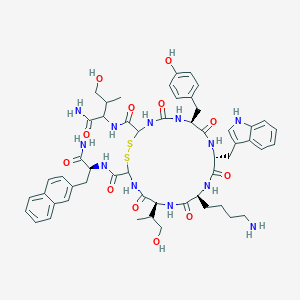 BDBM82462 BDBM82462 | C54H68N12O12S2 | 1141.33 | 15 / 15 | 1.5 | No |
| 299000 | 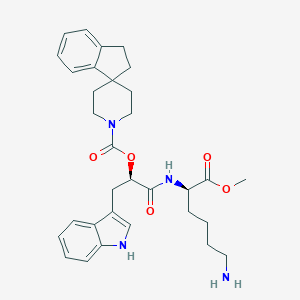 CHEMBL146022 CHEMBL146022 | C32H40N4O5 | 560.695 | 6 / 3 | 4.5 | No |
| 308753 | 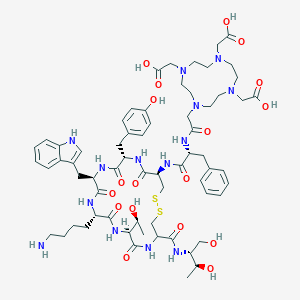 CHEMBL1791313 CHEMBL1791313 | C65H92N14O18S2 | 1421.65 | 25 / 17 | -7.2 | No |
| 556742 | 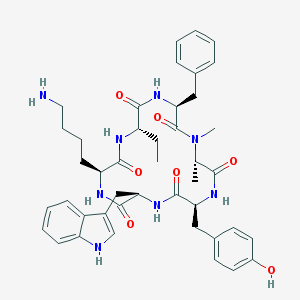 BIM-23027 BIM-23027 | C43H54N8O7 | 794.954 | 8 / 8 | 3.5 | No |
| 556793 | 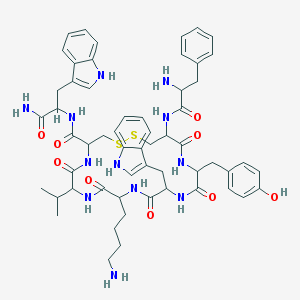 Vapreotide Vapreotide | C57H70N12O9S2 | 1131.38 | 13 / 13 | 3.6 | No |
| 556802 | 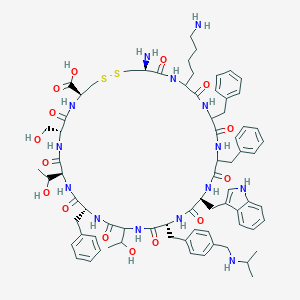 CH-275 CH-275 | C74H96N14O15S2 | 1485.79 | 20 / 18 | 1.6 | No |
| 556838 | 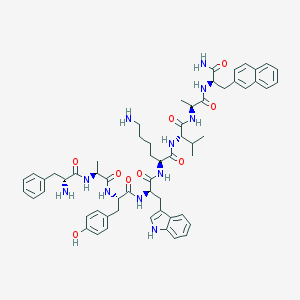 BDBM82461 BDBM82461 | C59H73N11O9 | 1080.3 | 11 / 12 | 5.0 | No |
| 556893 | 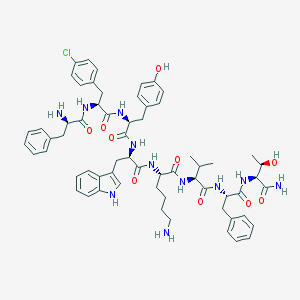 SCHEMBL1579410 SCHEMBL1579410 | C62H76ClN11O10 | 1170.81 | 12 / 13 | 5.3 | No |
| 556896 | 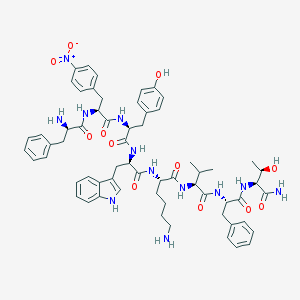 CHEMBL3350185 CHEMBL3350185 | C62H76N12O12 | 1181.36 | 14 / 13 | 4.5 | No |
| 346567 | 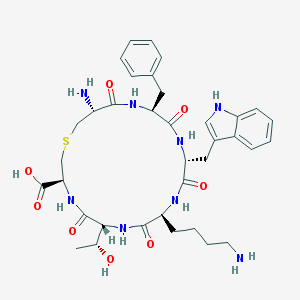 CHEMBL306702 CHEMBL306702 | C36H48N8O8S | 752.888 | 11 / 10 | -1.6 | No |
| 556903 | 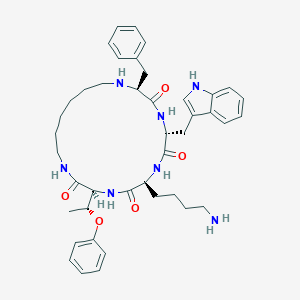 cycloantagonist SA cycloantagonist SA | C42H55N7O5 | 737.946 | 7 / 7 | 4.8 | No |
| 556955 | 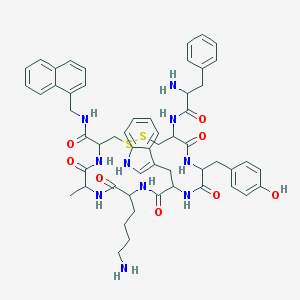 BDBM84626 BDBM84626 | C55H64N10O8S2 | 1057.3 | 12 / 11 | 4.3 | No |
| 556979 | 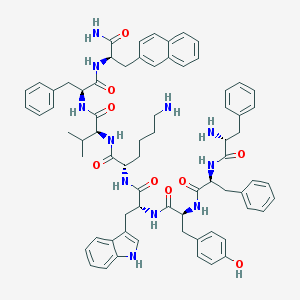 BIM 23056 BIM 23056 | C71H81N11O9 | 1232.5 | 11 / 12 | 8.1 | No |
| 556980 | 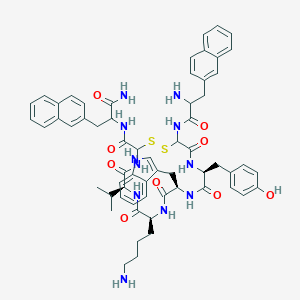 BDBM84621 BDBM84621 | C61H69N11O9S2 | 1164.41 | 13 / 12 | 6.1 | No |
| 555100 | 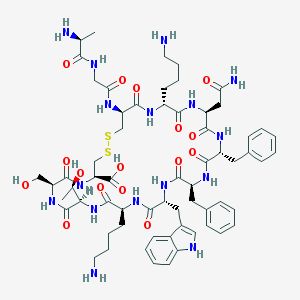 SS14 SS14 | C63H88N16O16S2 | 1389.61 | 21 / 19 | -4.1 | No |
| 381832 | 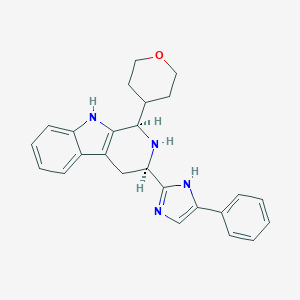 CHEMBL2069502 CHEMBL2069502 | C25H26N4O | 398.51 | 3 / 3 | 3.5 | Yes |
| 557084 | 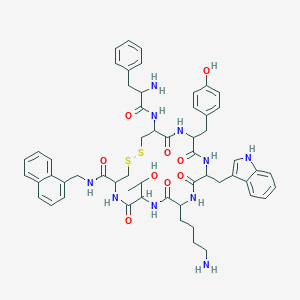 BDBM84630 BDBM84630 | C56H66N10O9S2 | 1087.32 | 13 / 12 | 4.2 | No |
| 557118 | 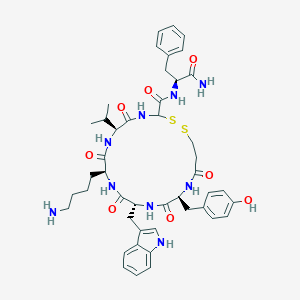 BDBM82456 BDBM82456 | C45H57N9O8S2 | 916.126 | 11 / 10 | 3.0 | No |
| 394310 | 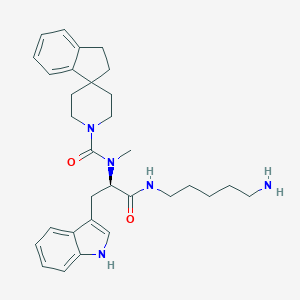 CHEMBL144146 CHEMBL144146 | C31H41N5O2 | 515.702 | 3 / 3 | 4.1 | No |
| 557137 | 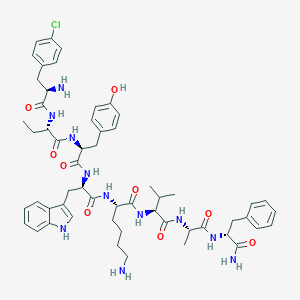 BDBM84627 BDBM84627 | C56H72ClN11O9 | 1078.71 | 11 / 12 | 4.2 | No |
| 557154 | 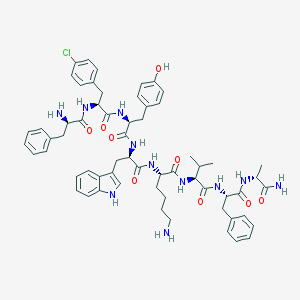 CHEMBL264562 CHEMBL264562 | C61H74ClN11O9 | 1140.78 | 11 / 12 | 5.1 | No |
| 557211 | 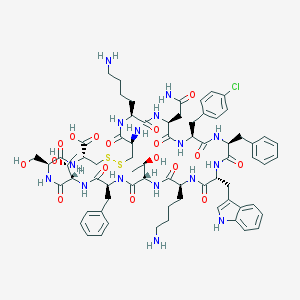 Bim-23003 Bim-23003 | C71H95ClN16O17S2 | 1544.21 | 22 / 20 | -1.6 | No |
| 557265 | 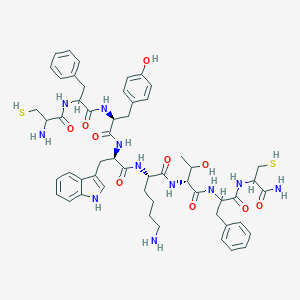 BDBM85229 BDBM85229 | C54H69N11O10S2 | 1096.33 | 14 / 15 | 2.0 | No |
| 557280 | 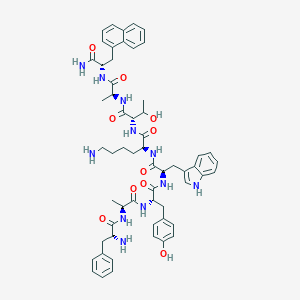 BDBM84638 BDBM84638 | C58H71N11O10 | 1082.27 | 12 / 13 | 3.4 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218