

You can:
| Name | Calcitonin receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | CALCR |
| Synonym | CTRI1- CTR2 CTR CT-R CT receptor [ Show all ] |
| Disease | Pain; Postmenopausal osteoporosis Paget's disease Hypertension Osteoporosis |
| Length | 508 |
| Amino acid sequence | MQFSGEKISGQRDLQKSKMRFTFTSRCLALFLLLNHPTPILPAFSNQTYPTIEPKPFLYVVGRKKMMDAQYKCYDRMQQLPAYQGEGPYCNRTWDGWLCWDDTPAGVLSYQFCPDYFPDFDPSEKVTKYCDEKGVWFKHPENNRTWSNYTMCNAFTPEKLKNAYVLYYLAIVGHSLSIFTLVISLGIFVFFRKLTTIFPLNWKYRKALSLGCQRVTLHKNMFLTYILNSMIIIIHLVEVVPNGELVRRDPVSCKILHFFHQYMMACNYFWMLCEGIYLHTLIVVAVFTEKQRLRWYYLLGWGFPLVPTTIHAITRAVYFNDNCWLSVETHLLYIIHGPVMAALVVNFFFLLNIVRVLVTKMRETHEAESHMYLKAVKATMILVPLLGIQFVVFPWRPSNKMLGKIYDYVMHSLIHFQGFFVATIYCFCNNEVQTTVKRQWAQFKIQWNQRWGRRPSNRSARAAAAAAEAGDIPIYICHQEPRNEPANNQGEESAEIIPLNIIEQESSA |
| UniProt | P30988 |
| Protein Data Bank | 6niy, 5ii0 |
| GPCR-HGmod model | P30988 |
| 3D structure model | This structure is from PDB ID 6niy. |
| BioLiP | BL0346076,BL0346077,BL0346078, BL0438754 |
| Therapeutic Target Database | T32247 |
| ChEMBL | CHEMBL1832 |
| IUPHAR | 43 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 18026 |  CHEMBL2369907 CHEMBL2369907 | C145H242N44O48S2 | 3433.91 | 55 / 53 | -18.5 | No |
| 108614 |  CHEMBL294095 CHEMBL294095 | C14H13FN2O4S | 324.326 | 5 / 1 | 0.7 | Yes |
| 110726 |  BMS-694153 BMS-694153 | C35H45FN8O3 | 644.796 | 6 / 3 | 3.6 | No |
| 113888 |  BDBM50422415 BDBM50422415 | C149H222N38O44S3 | 3345.82 | 50 / 44 | -10.7 | No |
| 130548 | 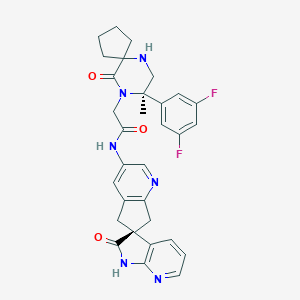 MK-8825 MK-8825 | C31H30F2N6O3 | 572.617 | 8 / 3 | 1.9 | No |
| 187600 | 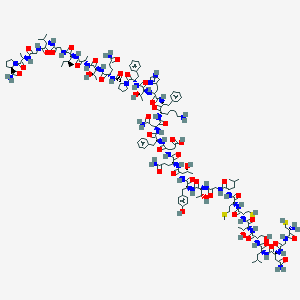 CHEMBL2369892 CHEMBL2369892 | C150H227N39O44S3 | 3376.88 | 51 / 46 | -11.3 | No |
| 187601 | 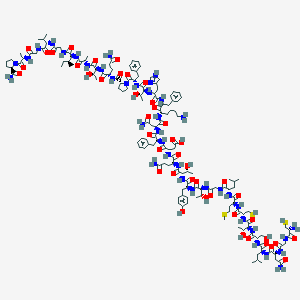 CGNLSTCMLGTYTQDFNKFHTFPQTAIGVGAP-amide CGNLSTCMLGTYTQDFNKFHTFPQTAIGVGAP-amide | C150H227N39O44S3 | 3376.88 | 51 / 46 | -11.3 | No |
| 190393 | 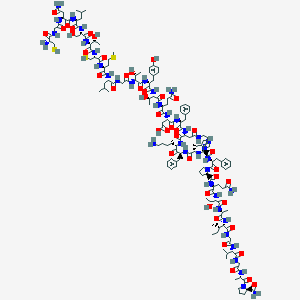 CHEMBL2369899 CHEMBL2369899 | C152H229N39O43S3 | 3386.91 | 50 / 45 | -9.7 | No |
| 190395 | 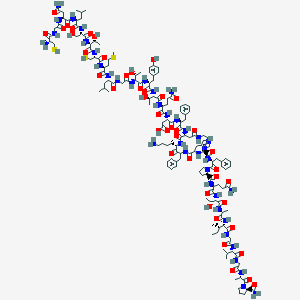 D03DPD D03DPD | C152H229N39O43S3 | 3386.91 | 50 / 45 | -9.7 | No |
| 203788 | 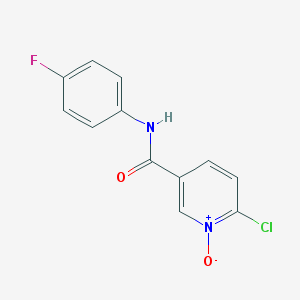 CHEMBL60066 CHEMBL60066 | C12H8ClFN2O2 | 266.656 | 3 / 1 | 1.8 | Yes |
| 225992 | 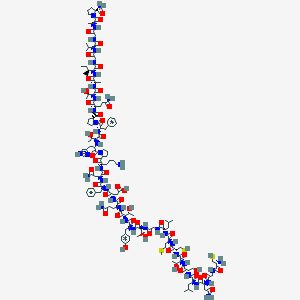 CGNLSTCMLGTYTQDFNKPHTFPQTAIGVGAP-amide CGNLSTCMLGTYTQDFNKPHTFPQTAIGVGAP-amide | C146H225N39O44S3 | 3326.81 | 51 / 45 | -12.5 | No |
| 225995 | 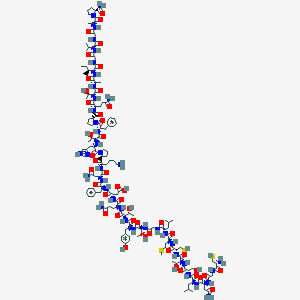 CHEMBL2369905 CHEMBL2369905 | C146H225N39O44S3 | 3326.81 | 51 / 45 | -12.5 | No |
| 230441 |  D0A8VZ D0A8VZ | C152H229N39O43S2 | 3354.85 | 49 / 45 | -9.2 | No |
| 230444 |  CHEMBL2369893 CHEMBL2369893 | C152H229N39O43S2 | 3354.85 | 49 / 45 | -9.2 | No |
| 232345 |  CGNLSTCBLGTYTQDFNKFHZYPQTAIGVGAP-amide CGNLSTCBLGTYTQDFNKFHZYPQTAIGVGAP-amide | C152H231N39O44S2 | 3372.87 | 50 / 46 | -9.3 | No |
| 232347 |  CHEMBL2369897 CHEMBL2369897 | C152H231N39O44S2 | 3372.87 | 50 / 46 | -9.3 | No |
| 243836 | 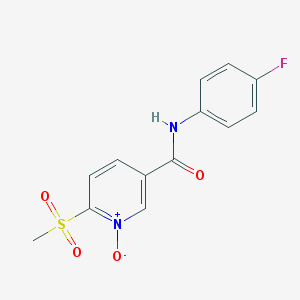 CHEMBL61835 CHEMBL61835 | C13H11FN2O4S | 310.299 | 5 / 1 | 0.4 | Yes |
| 262414 | 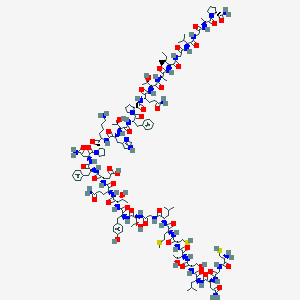 CHEMBL2369904 CHEMBL2369904 | C146H225N39O44S3 | 3326.81 | 51 / 45 | -12.5 | No |
| 262417 | 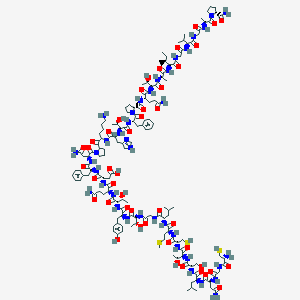 D0Y0NM D0Y0NM | C146H225N39O44S3 | 3326.81 | 51 / 45 | -12.5 | No |
| 265577 | 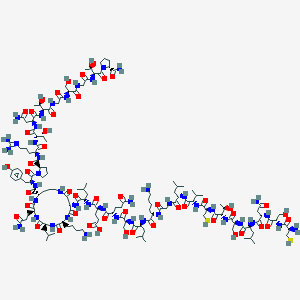 CHEMBL2369895 CHEMBL2369895 | C144H241N43O48S2 | 3406.89 | 54 / 52 | -18.7 | No |
| 265580 |  CSNLSTCVLGKLSQELc[DKLQK]YPRTNTGSGTP-amide CSNLSTCVLGKLSQELc[DKLQK]YPRTNTGSGTP-amide | C144H241N43O48S2 | 3406.89 | 54 / 52 | -18.7 | No |
| 314692 |  CHEMBL62108 CHEMBL62108 | C17H17FN2O4S | 364.391 | 5 / 1 | 1.6 | Yes |
| 335513 |  D00JEO D00JEO | C146H243N45O47S2 | 3444.94 | 55 / 52 | -17.7 | No |
| 335514 |  CHEMBL2369912 CHEMBL2369912 | C146H243N45O47S2 | 3444.94 | 55 / 52 | -17.7 | No |
| 352696 | 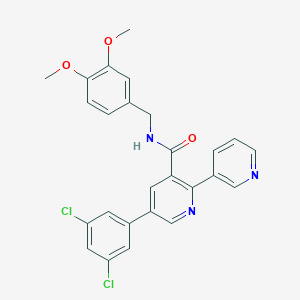 CHEMBL3099899 CHEMBL3099899 | C26H21Cl2N3O3 | 494.372 | 5 / 1 | 5.0 | Yes |
| 401265 | 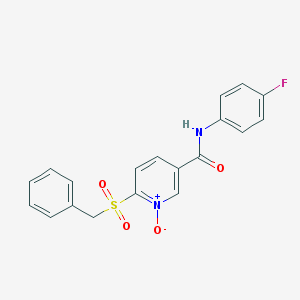 CHEMBL58619 CHEMBL58619 | C19H15FN2O4S | 386.397 | 5 / 1 | 1.9 | Yes |
| 414771 | 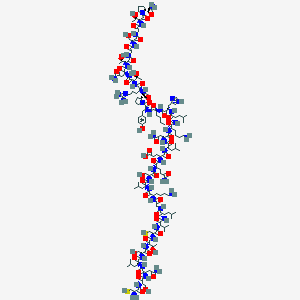 CHEMBL2369915 CHEMBL2369915 | C146H244N44O47S2 | 3431.94 | 54 / 52 | -16.8 | No |
| 416693 | 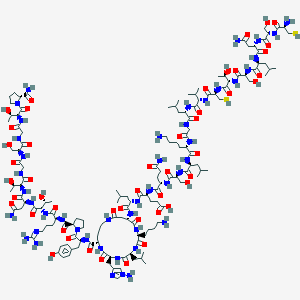 CHEMBL2369886 CHEMBL2369886 | C145H241N45O47S2 | 3430.91 | 55 / 52 | -18.0 | No |
| 416696 |  CSNLSTCVLGKLSQELc[DKLHO]YPRTNTGSGTP-amide CSNLSTCVLGKLSQELc[DKLHO]YPRTNTGSGTP-amide | C145H241N45O47S2 | 3430.91 | 55 / 52 | -18.0 | No |
| 422262 |  CHEMBL2369898 CHEMBL2369898 | C142H216N38O44S3 | 3255.69 | 50 / 44 | -12.7 | No |
| 555427 |  AC 187 AC 187 | C127H205N37O40 | 2890.26 | 44 / 44 | -13.2 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218