

You can:
| Name | Histamine H1 receptor |
|---|---|
| Species | Cavia porcellus (Guinea pig) |
| Gene | HRH1 |
| Synonym | H1R HH1R |
| Disease | N/A for non-human GPCRs |
| Length | 488 |
| Amino acid sequence | MSFLPGMTPVTLSNFSWALEDRMLEGNSTTTPTRQLMPLVVVLSSVSLVTVALNLLVLYAVRSERKLHTVGNLYIVSLSVADLIVGAVVMPMSILYLHRSAWILGRPLCLFWLSMDYVASTASIFSVFILCIDRYRSVQQPLRYLRYRTKTRASATILGAWLLSFLWVIPILGWHHFMAPTSEPREKKCETDFYDVTWFKVMTAIINFYLPTLLMLWFYIRIYKAVRRHCQHRQLINSSLPSFSEMKLKLENAKVDTRRMGKESPWEDPKRCSKDASGVHTPMPSSQHLVDMPCAAVLSEDEGGEVGTRQMPMLAVGDGRCCEALNHMHSQLELSGQSRATHSISARPEEWTVVDGQSFPITDSDTSTEAAPMGGQPRSGSNSGLDYIKFTWRRLRSHSRQYTSGLHLNRERKAAKQLGCIMAAFILCWIPYFVFFMVIAFCKSCSNEPVHMFTIWLGYLNSTLNPLIYPLCNENFRKTFKRILRIPP |
| UniProt | P31389 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3943 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 190 | 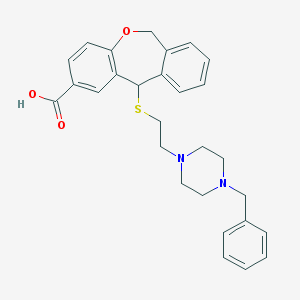 CHEMBL134543 CHEMBL134543 | C28H30N2O3S | 474.619 | 6 / 1 | 2.2 | Yes |
| 759 | 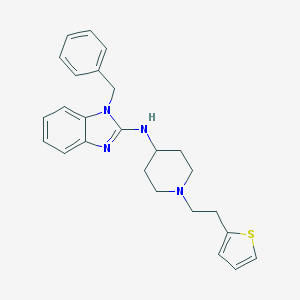 CHEMBL300353 CHEMBL300353 | C25H28N4S | 416.587 | 4 / 1 | 5.6 | No |
| 1294 | 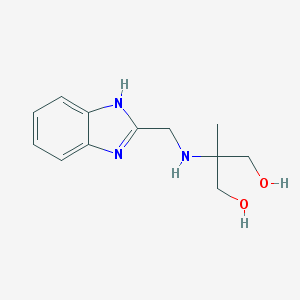 CHEMBL2262480 CHEMBL2262480 | C12H17N3O2 | 235.287 | 4 / 4 | -0.2 | Yes |
| 1387 | 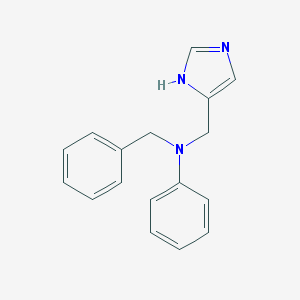 CHEMBL348369 CHEMBL348369 | C17H17N3 | 263.344 | 2 / 1 | 3.2 | Yes |
| 1888 | 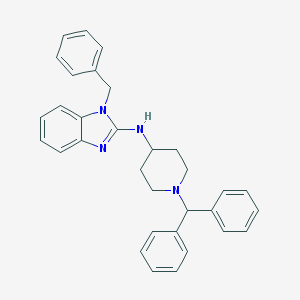 CHEMBL301238 CHEMBL301238 | C32H32N4 | 472.636 | 3 / 1 | 7.1 | No |
| 2956 | 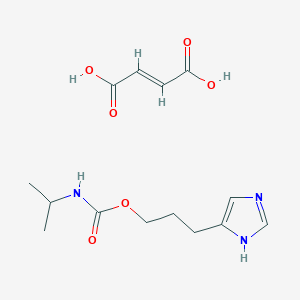 CID 44370594 CID 44370594 | C14H21N3O6 | 327.337 | 7 / 4 | N/A | N/A |
| 3343 | 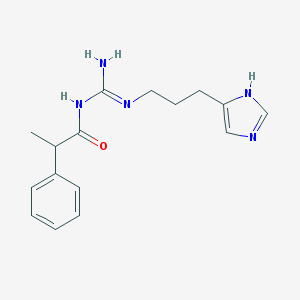 UR-AK68 UR-AK68 | C16H21N5O | 299.378 | 3 / 3 | 2.1 | Yes |
| 3646 | 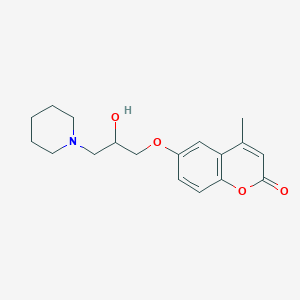 CHEMBL518264 CHEMBL518264 | C18H23NO4 | 317.385 | 5 / 1 | 2.1 | Yes |
| 3698 | 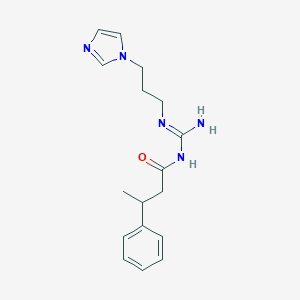 CHEMBL3220892 CHEMBL3220892 | C17H23N5O | 313.405 | 3 / 2 | 1.7 | Yes |
| 4473 | 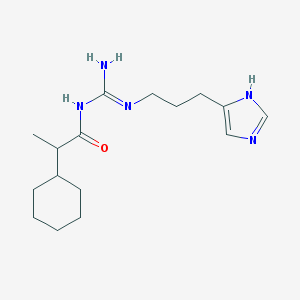 UR-AK59 UR-AK59 | C16H27N5O | 305.426 | 3 / 3 | 3.1 | Yes |
| 5297 | 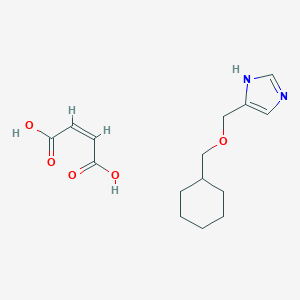 CID 44323707 CID 44323707 | C15H22N2O5 | 310.35 | 6 / 3 | N/A | N/A |
| 5324 | 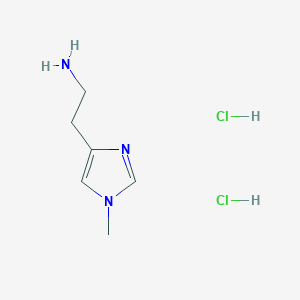 1-Methylhistamine dihydrochloride 1-Methylhistamine dihydrochloride | C6H13Cl2N3 | 198.091 | 2 / 3 | N/A | N/A |
| 5750 | 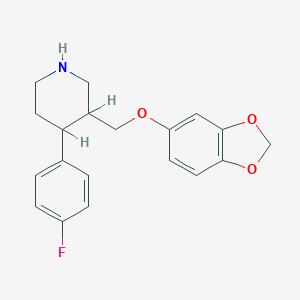 AHOUBRCZNHFOSL-UHFFFAOYSA-N AHOUBRCZNHFOSL-UHFFFAOYSA-N | C19H20FNO3 | 329.371 | 5 / 1 | 3.5 | Yes |
| 5764 | 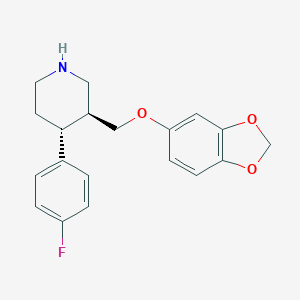 paroxetine paroxetine | C19H20FNO3 | 329.371 | 5 / 1 | 3.5 | Yes |
| 5941 | 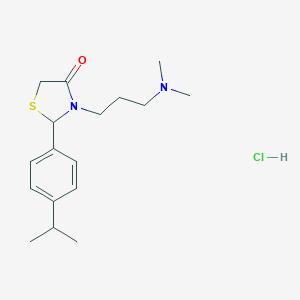 CHEMBL1202934 CHEMBL1202934 | C17H27ClN2OS | 342.926 | 3 / 1 | N/A | N/A |
| 6500 | 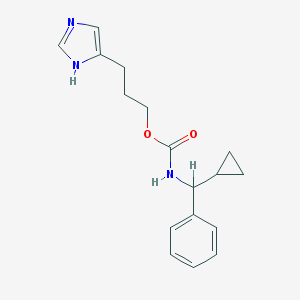 CHEMBL128585 CHEMBL128585 | C17H21N3O2 | 299.374 | 3 / 2 | 2.7 | Yes |
| 6973 | 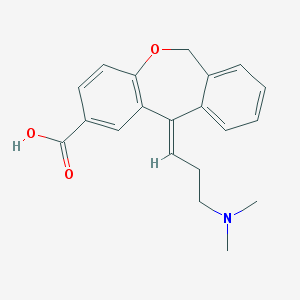 CHEMBL65822 CHEMBL65822 | C20H21NO3 | 323.392 | 4 / 1 | 1.5 | Yes |
| 6974 | 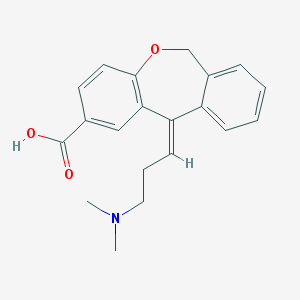 CHEMBL64067 CHEMBL64067 | C20H21NO3 | 323.392 | 4 / 1 | 1.5 | Yes |
| 8214 | 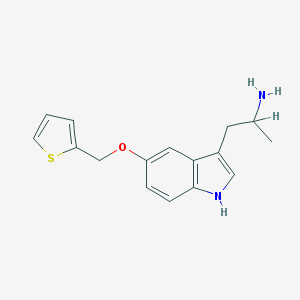 BW-723C86 BW-723C86 | C16H18N2OS | 286.393 | 3 / 2 | 3.0 | Yes |
| 8369 | 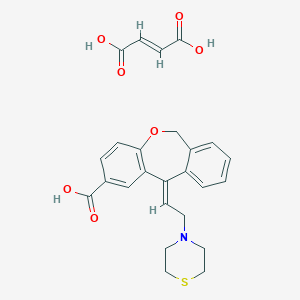 CHEMBL63746 CHEMBL63746 | C25H25NO7S | 483.535 | 9 / 3 | N/A | N/A |
| 8416 | 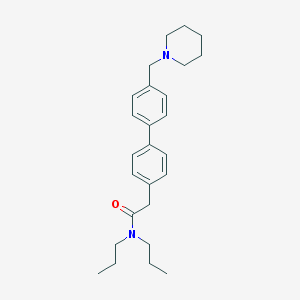 CHEMBL458507 CHEMBL458507 | C26H36N2O | 392.587 | 2 / 0 | 5.4 | No |
| 8724 | 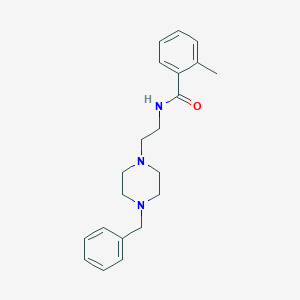 CHEMBL328170 CHEMBL328170 | C21H27N3O | 337.467 | 3 / 1 | 2.9 | Yes |
| 9376 | 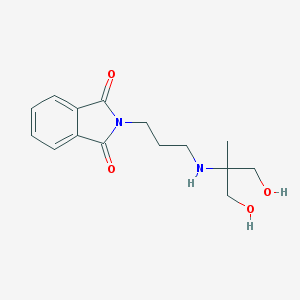 CHEMBL2262485 CHEMBL2262485 | C15H20N2O4 | 292.335 | 5 / 3 | 0.3 | Yes |
| 9844 | 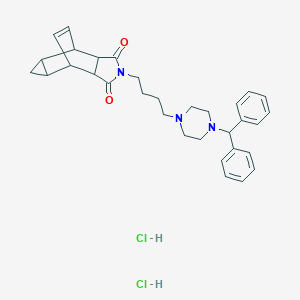 CHEMBL541594 CHEMBL541594 | C32H39Cl2N3O2 | 568.583 | 4 / 2 | N/A | No |
| 9902 | 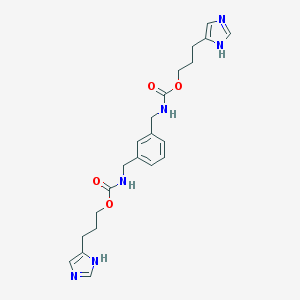 CHEMBL415989 CHEMBL415989 | C22H28N6O4 | 440.504 | 6 / 4 | 1.9 | Yes |
| 11133 | 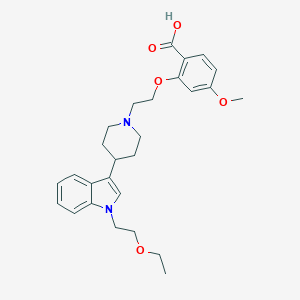 CHEMBL181278 CHEMBL181278 | C27H34N2O5 | 466.578 | 6 / 1 | 1.6 | Yes |
| 11654 | 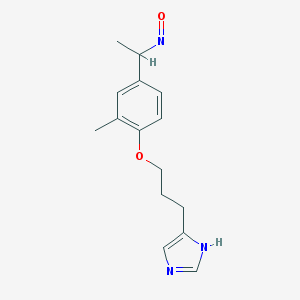 BDBM50091375 BDBM50091375 | C15H19N3O2 | 273.336 | 4 / 1 | 2.5 | Yes |
| 11754 | 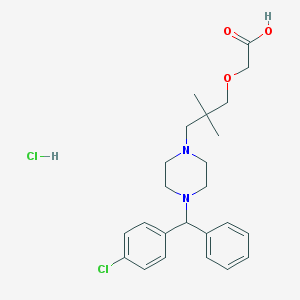 CHEMBL542427 CHEMBL542427 | C24H32Cl2N2O3 | 467.431 | 5 / 2 | N/A | N/A |
| 11985 | 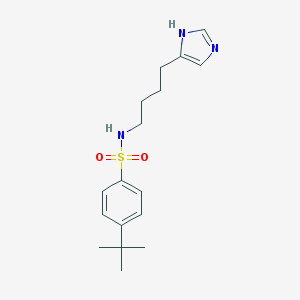 CHEMBL68764 CHEMBL68764 | C17H25N3O2S | 335.466 | 4 / 2 | 3.4 | Yes |
| 12270 | 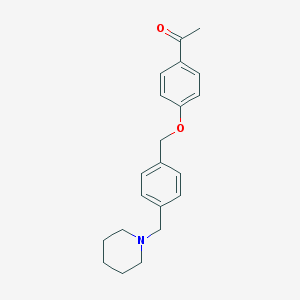 CHEMBL39603 CHEMBL39603 | C21H25NO2 | 323.436 | 3 / 0 | 3.8 | Yes |
| 12407 | 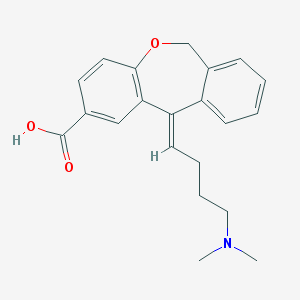 CHEMBL304042 CHEMBL304042 | C21H23NO3 | 337.419 | 4 / 1 | 1.9 | Yes |
| 12409 | 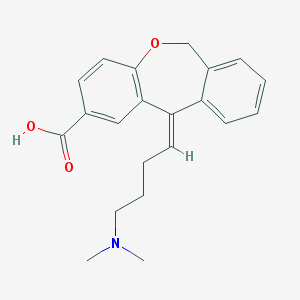 CHEMBL294345 CHEMBL294345 | C21H23NO3 | 337.419 | 4 / 1 | 1.9 | Yes |
| 13441 | 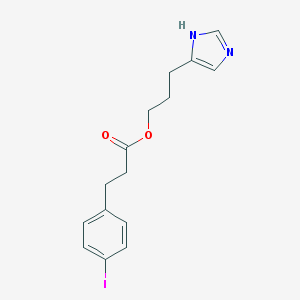 CHEMBL423473 CHEMBL423473 | C15H17IN2O2 | 384.217 | 3 / 1 | 3.1 | Yes |
| 14253 | 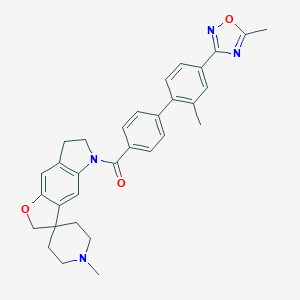 180083-23-2 180083-23-2 | C32H32N4O3 | 520.633 | 6 / 0 | 5.6 | No |
| 14681 | 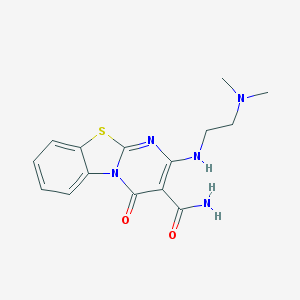 CHEMBL2283392 CHEMBL2283392 | C15H17N5O2S | 331.394 | 6 / 2 | 2.0 | Yes |
| 14749 | 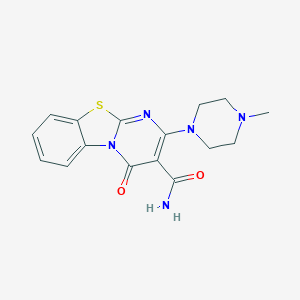 CHEMBL2283395 CHEMBL2283395 | C16H17N5O2S | 343.405 | 6 / 1 | 1.4 | Yes |
| 15411 | 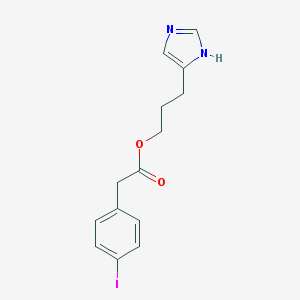 CHEMBL277227 CHEMBL277227 | C14H15IN2O2 | 370.19 | 3 / 1 | 2.8 | Yes |
| 15764 | 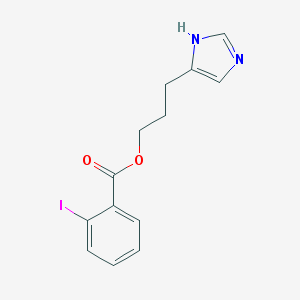 CHEMBL20191 CHEMBL20191 | C13H13IN2O2 | 356.163 | 3 / 1 | 3.3 | Yes |
| 16201 | 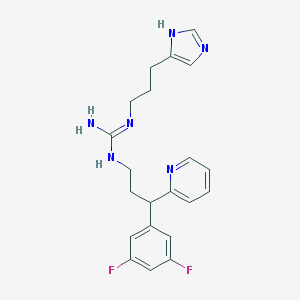 BU-E-76 BU-E-76 | C21H24F2N6 | 398.462 | 5 / 3 | 2.5 | Yes |
| 16783 | 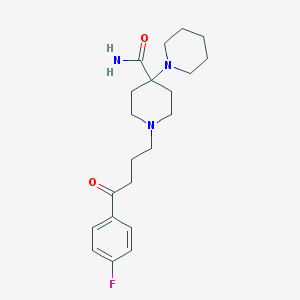 Pipamperone Pipamperone | C21H30FN3O2 | 375.488 | 5 / 1 | 2.0 | Yes |
| 17387 | 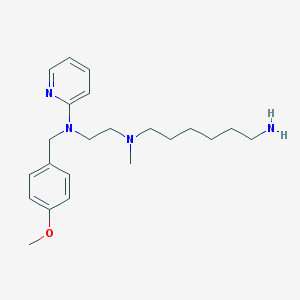 CHEMBL21278 CHEMBL21278 | C22H34N4O | 370.541 | 5 / 1 | 3.8 | Yes |
| 18778 | 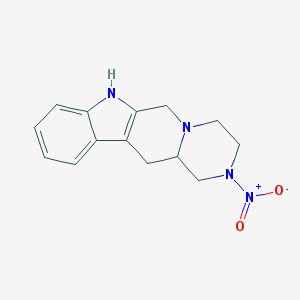 CHEMBL97938 CHEMBL97938 | C14H16N4O2 | 272.308 | 4 / 1 | 2.2 | Yes |
| 18815 | 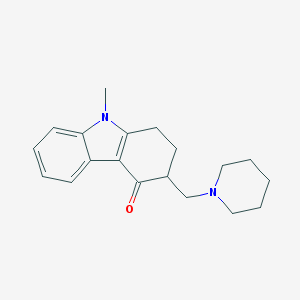 BRN 0418992 BRN 0418992 | C19H24N2O | 296.414 | 2 / 0 | 3.1 | Yes |
| 18875 | 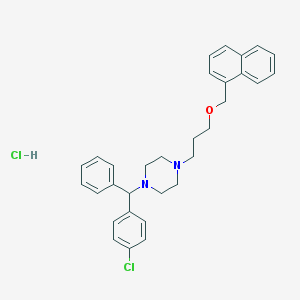 CHEMBL552864 CHEMBL552864 | C31H34Cl2N2O | 521.526 | 3 / 1 | N/A | No |
| 18898 | 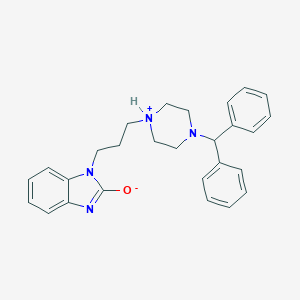 AC1LQZR1 AC1LQZR1 | C27H30N4O | 426.564 | 3 / 1 | 5.4 | No |
| 18998 | 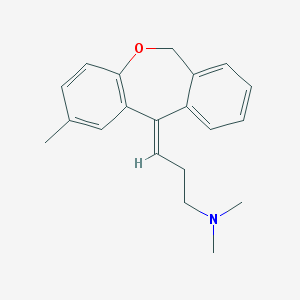 CHEMBL366965 CHEMBL366965 | C20H23NO | 293.41 | 2 / 0 | 4.7 | Yes |
| 19725 | 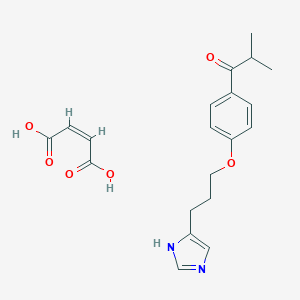 CHEMBL131021 CHEMBL131021 | C20H24N2O6 | 388.42 | 7 / 3 | N/A | N/A |
| 20232 | 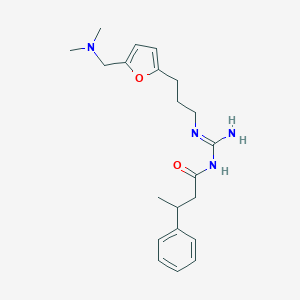 CHEMBL3220888 CHEMBL3220888 | C21H30N4O2 | 370.497 | 4 / 2 | 2.9 | Yes |
| 20249 | 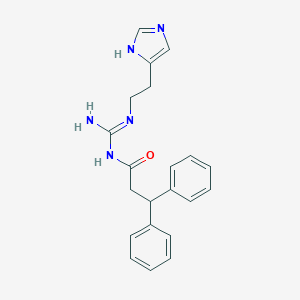 CHEMBL499301 CHEMBL499301 | C21H23N5O | 361.449 | 3 / 3 | 2.9 | Yes |
| 21680 | 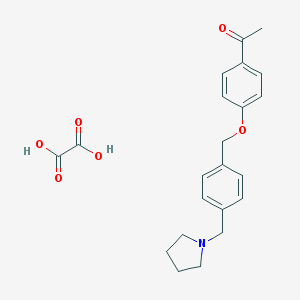 CHEMBL290613 CHEMBL290613 | C22H25NO6 | 399.443 | 7 / 2 | N/A | N/A |
| 22338 | 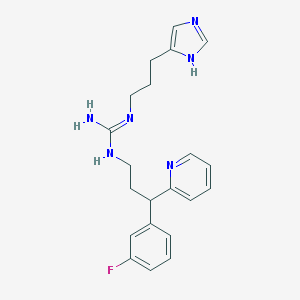 CHEMBL61675 CHEMBL61675 | C21H25FN6 | 380.471 | 4 / 3 | 2.4 | Yes |
| 23527 | 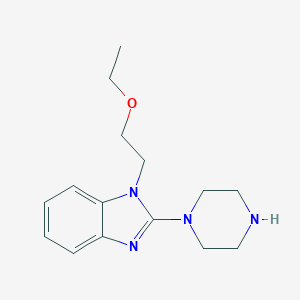 CHEMBL278053 CHEMBL278053 | C15H22N4O | 274.368 | 4 / 1 | 1.4 | Yes |
| 24183 | 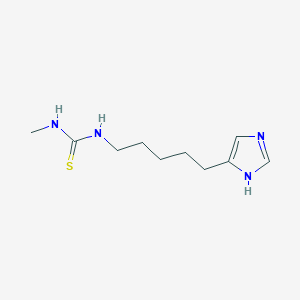 CHEMBL44220 CHEMBL44220 | C10H18N4S | 226.342 | 2 / 3 | 0.8 | Yes |
| 24388 | 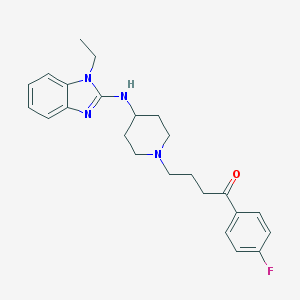 CHEMBL56659 CHEMBL56659 | C24H29FN4O | 408.521 | 5 / 1 | 4.4 | Yes |
| 24432 | 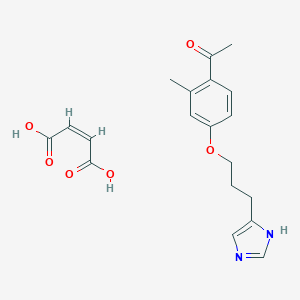 CHEMBL128513 CHEMBL128513 | C19H22N2O6 | 374.393 | 7 / 3 | N/A | N/A |
| 24487 | 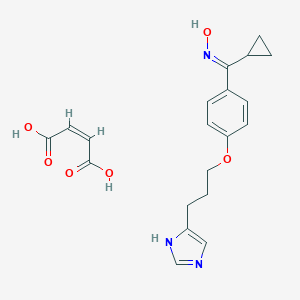 CID 44341209 CID 44341209 | C20H23N3O6 | 401.419 | 8 / 4 | N/A | N/A |
| 25378 | 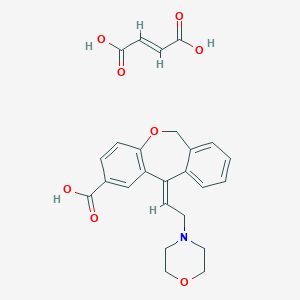 CHEMBL64263 CHEMBL64263 | C25H25NO8 | 467.474 | 9 / 3 | N/A | N/A |
| 26061 | 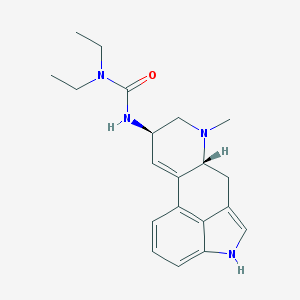 S-(-)-Lisuride S-(-)-Lisuride | C20H26N4O | 338.455 | 2 / 2 | 2.7 | Yes |
| 26071 | 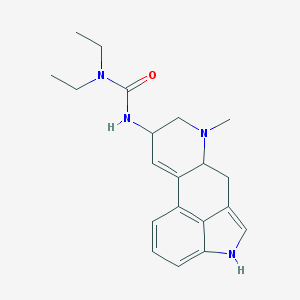 AC1L1H1T AC1L1H1T | C20H26N4O | 338.455 | 2 / 2 | 2.7 | Yes |
| 26487 | 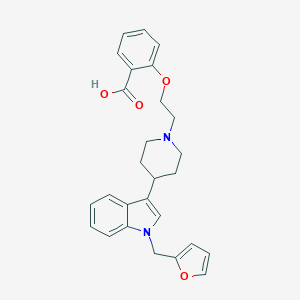 CHEMBL374218 CHEMBL374218 | C27H28N2O4 | 444.531 | 5 / 1 | 2.2 | Yes |
| 26675 | 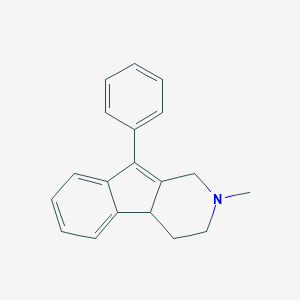 CHEMBL22215 CHEMBL22215 | C19H19N | 261.368 | 1 / 0 | 3.2 | Yes |
| 26967 | 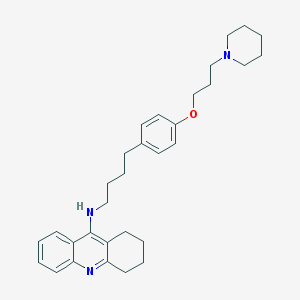 CHEMBL14868 CHEMBL14868 | C31H41N3O | 471.689 | 4 / 1 | 7.3 | No |
| 27148 | 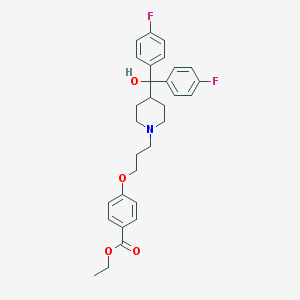 CHEMBL555593 CHEMBL555593 | C30H33F2NO4 | 509.594 | 7 / 1 | 5.8 | No |
| 27629 | 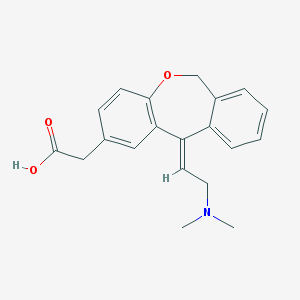 CHEMBL302005 CHEMBL302005 | C20H21NO3 | 323.392 | 4 / 1 | 1.0 | Yes |
| 27635 | 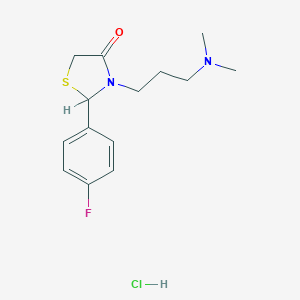 CHEMBL1202939 CHEMBL1202939 | C14H20ClFN2OS | 318.835 | 4 / 1 | N/A | N/A |
| 27738 | 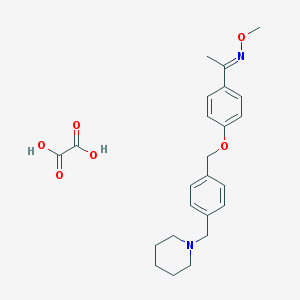 CHEMBL35869 CHEMBL35869 | C24H30N2O6 | 442.512 | 8 / 2 | N/A | N/A |
| 27792 | 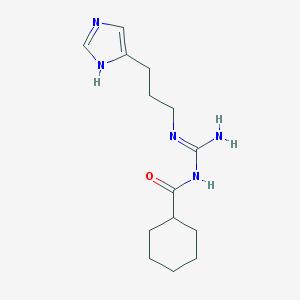 UR-AK46 UR-AK46 | C14H23N5O | 277.372 | 3 / 3 | 2.0 | Yes |
| 442794 | 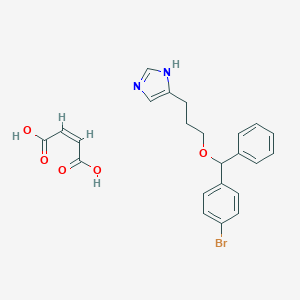 CHEMBL59851 CHEMBL59851 | C23H23BrN2O5 | 487.35 | 6 / 3 | N/A | N/A |
| 29320 | 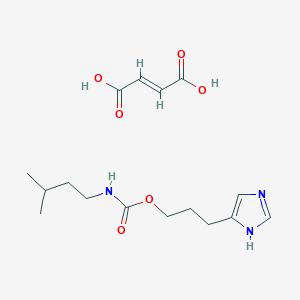 CID 44370355 CID 44370355 | C16H25N3O6 | 355.391 | 7 / 4 | N/A | N/A |
| 29683 | 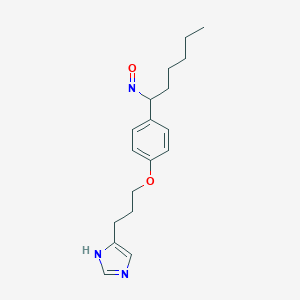 BDBM50091370 BDBM50091370 | C18H25N3O2 | 315.417 | 4 / 1 | 4.1 | Yes |
| 29702 | 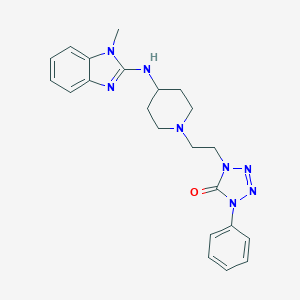 CHEMBL300383 CHEMBL300383 | C22H26N8O | 418.505 | 6 / 1 | 3.9 | Yes |
| 29843 | 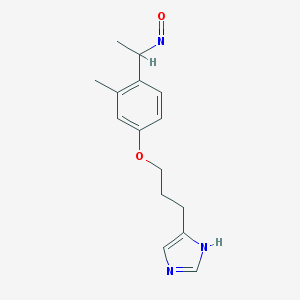 BDBM50091371 BDBM50091371 | C15H19N3O2 | 273.336 | 4 / 1 | 2.5 | Yes |
| 30030 | 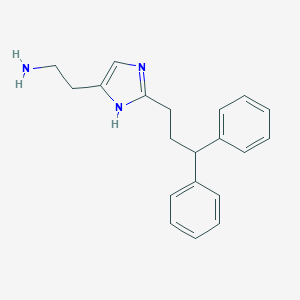 Histaprodifen Histaprodifen | C20H23N3 | 305.425 | 2 / 2 | 3.5 | Yes |
| 30723 | 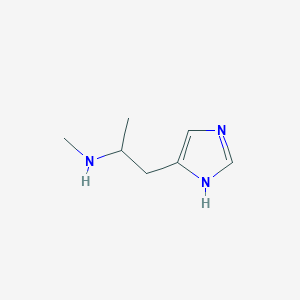 76721-87-4 76721-87-4 | C7H13N3 | 139.202 | 2 / 2 | 0.4 | Yes |
| 30801 | 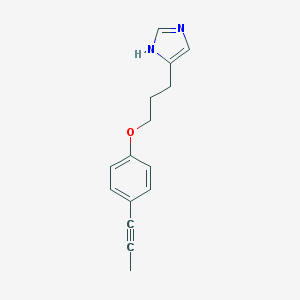 CHEMBL132798 CHEMBL132798 | C15H16N2O | 240.306 | 2 / 1 | 3.1 | Yes |
| 31228 | 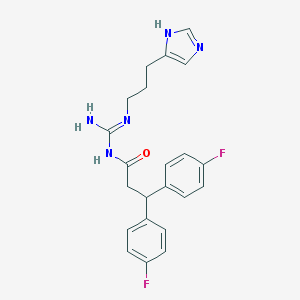 UR-PG55B UR-PG55B | C22H23F2N5O | 411.457 | 5 / 3 | 3.4 | Yes |
| 31550 | 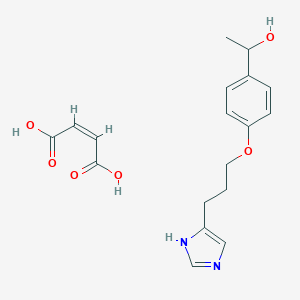 CID 44341348 CID 44341348 | C18H22N2O6 | 362.382 | 7 / 4 | N/A | N/A |
| 32407 | 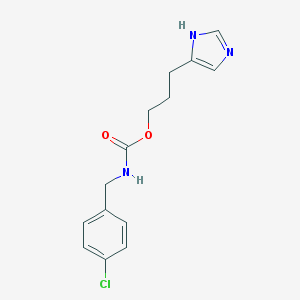 CHEMBL277149 CHEMBL277149 | C14H16ClN3O2 | 293.751 | 3 / 2 | 2.6 | Yes |
| 32497 | 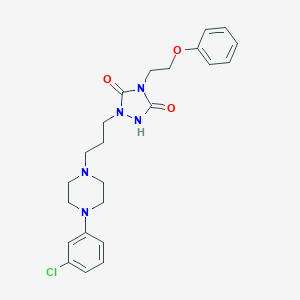 Triazoledione Triazoledione | C23H28ClN5O3 | 457.959 | 5 / 1 | 3.6 | Yes |
| 32672 | 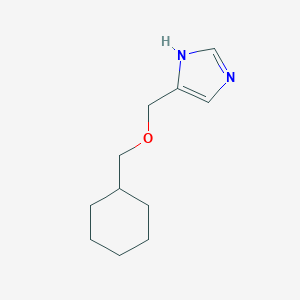 SCHEMBL8106368 SCHEMBL8106368 | C11H18N2O | 194.278 | 2 / 1 | 2.1 | Yes |
| 32674 | 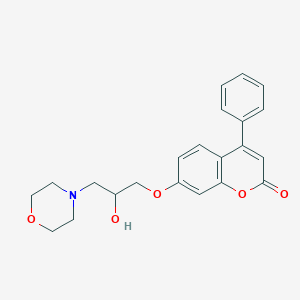 CHEMBL477410 CHEMBL477410 | C22H23NO5 | 381.428 | 6 / 1 | 2.1 | Yes |
| 32905 | 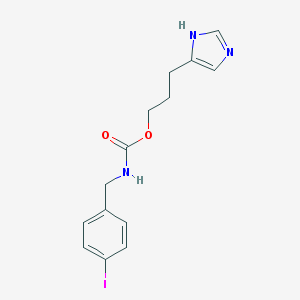 CHEMBL16496 CHEMBL16496 | C14H16IN3O2 | 385.205 | 3 / 2 | 2.6 | Yes |
| 33576 | 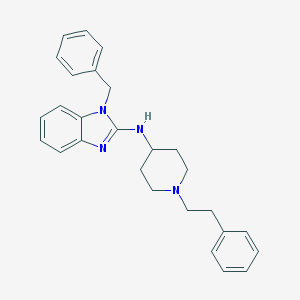 CHEMBL418296 CHEMBL418296 | C27H30N4 | 410.565 | 3 / 1 | 5.9 | No |
| 33847 | 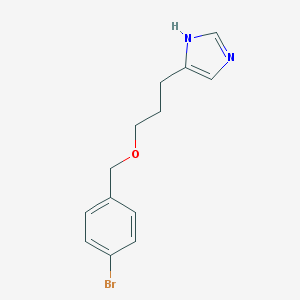 CHEMBL19172 CHEMBL19172 | C13H15BrN2O | 295.18 | 2 / 1 | 2.8 | Yes |
| 442984 | 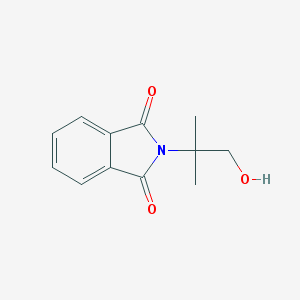 4490-74-8 4490-74-8 | C12H13NO3 | 219.24 | 3 / 1 | 1.2 | Yes |
| 33992 | 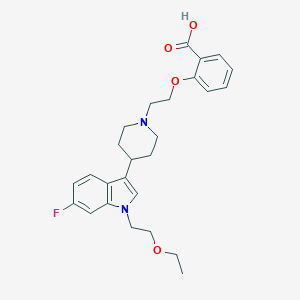 CHEMBL221413 CHEMBL221413 | C26H31FN2O4 | 454.542 | 6 / 1 | 1.7 | Yes |
| 34289 | 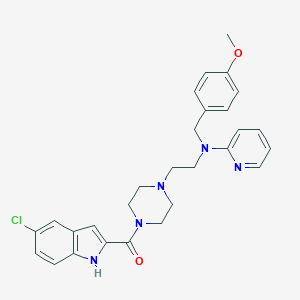 CHEMBL1910379 CHEMBL1910379 | C28H30ClN5O2 | 504.031 | 5 / 1 | 4.8 | No |
| 35818 | 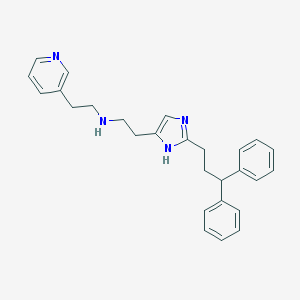 CHEMBL356449 CHEMBL356449 | C27H30N4 | 410.565 | 3 / 2 | 4.9 | Yes |
| 36318 | 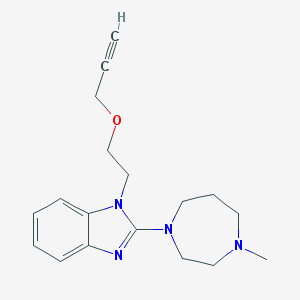 CHEMBL279875 CHEMBL279875 | C18H24N4O | 312.417 | 4 / 0 | 2.0 | Yes |
| 36654 | 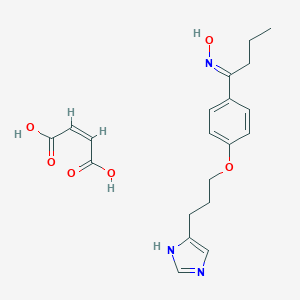 CID 44341037 CID 44341037 | C20H25N3O6 | 403.435 | 8 / 4 | N/A | N/A |
| 36815 | 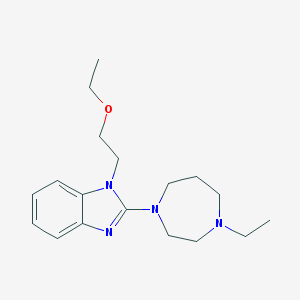 CHEMBL17474 CHEMBL17474 | C18H28N4O | 316.449 | 4 / 0 | 2.6 | Yes |
| 37801 | 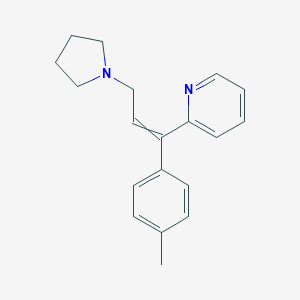 10191-42-1 10191-42-1 | C19H22N2 | 278.399 | 2 / 0 | 3.9 | Yes |
| 37802 | 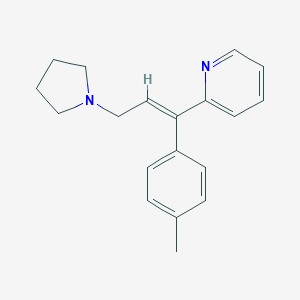 triprolidine triprolidine | C19H22N2 | 278.399 | 2 / 0 | 3.9 | Yes |
| 38049 | 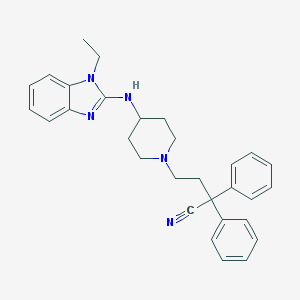 CHEMBL294867 CHEMBL294867 | C30H33N5 | 463.629 | 4 / 1 | 5.9 | No |
| 38264 | 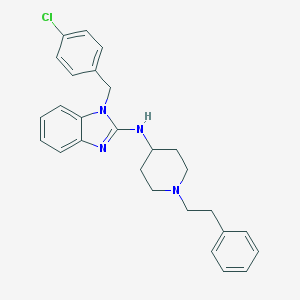 CHEMBL299282 CHEMBL299282 | C27H29ClN4 | 445.007 | 3 / 1 | 6.5 | No |
| 443203 | 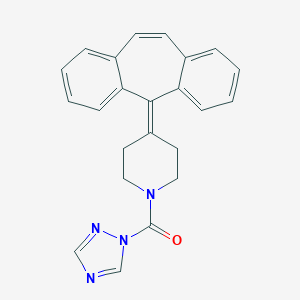 CHEMBL3402827 CHEMBL3402827 | C23H20N4O | 368.44 | 3 / 0 | 3.9 | Yes |
| 38674 | 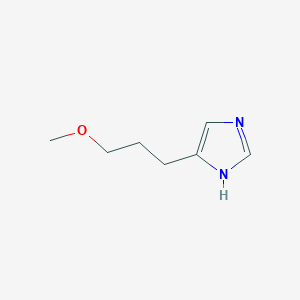 SCHEMBL7977259 SCHEMBL7977259 | C7H12N2O | 140.186 | 2 / 1 | 0.5 | Yes |
| 38705 | 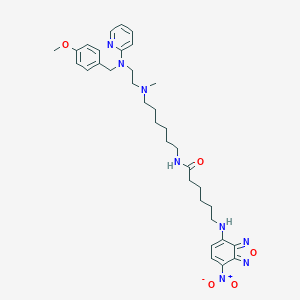 CHEMBL277848 CHEMBL277848 | C34H46N8O5 | 646.793 | 11 / 2 | 5.3 | No |
| 38730 | 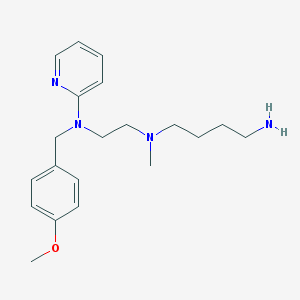 CHEMBL21520 CHEMBL21520 | C20H30N4O | 342.487 | 5 / 1 | 3.0 | Yes |
| 38885 | 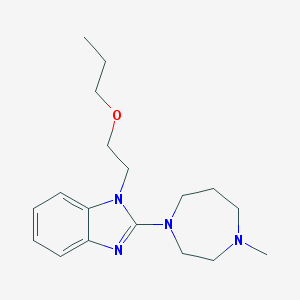 CHEMBL18234 CHEMBL18234 | C18H28N4O | 316.449 | 4 / 0 | 2.7 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218