

You can:
| Name | Adenosine receptor A1 |
|---|---|
| Species | Oryctolagus cuniculus (Rabbit) |
| Gene | ADORA1 |
| Synonym | N/A |
| Disease | N/A for non-human GPCRs |
| Length | 328 |
| Amino acid sequence | MPPSISAFQAAYIGIEVLIALVSVPGNVLVIWAVKVNQALRDATFCFIVSLAVADVAVGALVIPLAILINIGPETYFHTCLMVACPVLILTQSSILALLAIAVDRYLRVKIPLRYKAVVTPRRAAVAIAGCWILSLVVGLTPMFGWNNLREVQRAWAANGSVGEPVIKCEFEKVISMEYMVYFNFFVWVLPPLLLMVLIYLEVFYLIRRQLSKKASASSGDPHKYYGKELKIAKSLALILFLFALSWLPLHILNCVTLFCPSCQKPSILVYTAIFLTHGNSAMNPIVYAFRIHKFRVTFLKIWNDHFRCRPAPAGDGDEDLPEEKPND |
| UniProt | P34970 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3947 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 49482 | 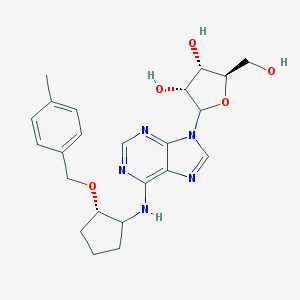 CHEMBL3674590 CHEMBL3674590 | C23H29N5O5 | 455.515 | 9 / 4 | 2.3 | Yes |
| 53518 | 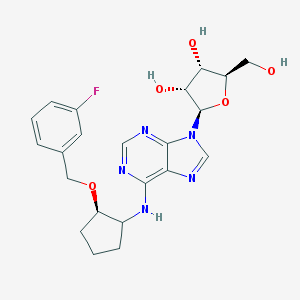 CHEMBL3679384 CHEMBL3679384 | C22H26FN5O5 | 459.478 | 10 / 4 | 2.0 | Yes |
| 53520 | 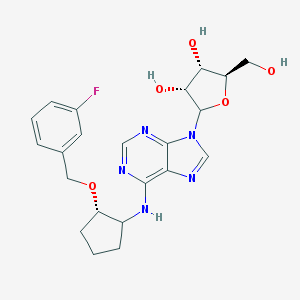 CHEMBL3679377 CHEMBL3679377 | C22H26FN5O5 | 459.478 | 10 / 4 | 2.0 | Yes |
| 62692 | 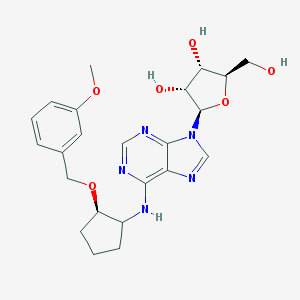 CHEMBL3679388 CHEMBL3679388 | C23H29N5O6 | 471.514 | 10 / 4 | 1.9 | Yes |
| 62694 | 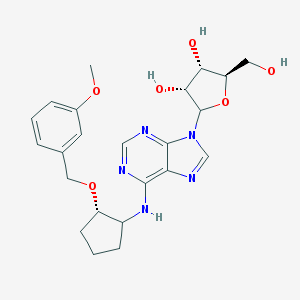 CHEMBL3679381 CHEMBL3679381 | C23H29N5O6 | 471.514 | 10 / 4 | 1.9 | Yes |
| 76822 | 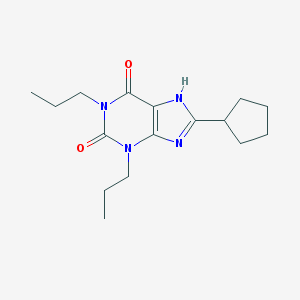 8-Cyclopentyl-1,3-dipropylxanthine 8-Cyclopentyl-1,3-dipropylxanthine | C16H24N4O2 | 304.394 | 3 / 1 | 4.0 | Yes |
| 79288 | 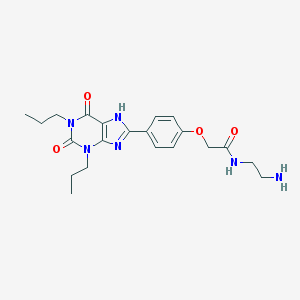 Xanthine amine congener Xanthine amine congener | C21H28N6O4 | 428.493 | 6 / 3 | 2.7 | Yes |
| 89152 | 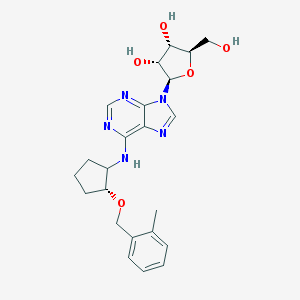 CHEMBL3679383 CHEMBL3679383 | C23H29N5O5 | 455.515 | 9 / 4 | 2.3 | Yes |
| 89153 | 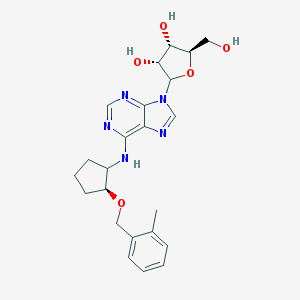 CHEMBL3679376 CHEMBL3679376 | C23H29N5O5 | 455.515 | 9 / 4 | 2.3 | Yes |
| 112748 | 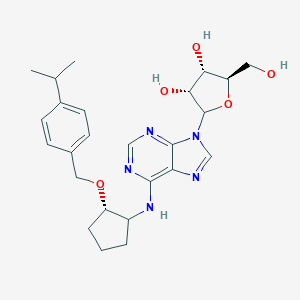 CHEMBL3679379 CHEMBL3679379 | C25H33N5O5 | 483.569 | 9 / 4 | 3.1 | Yes |
| 112750 | 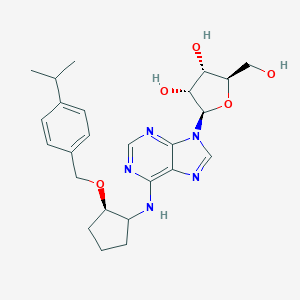 CHEMBL3679386 CHEMBL3679386 | C25H33N5O5 | 483.569 | 9 / 4 | 3.1 | Yes |
| 124388 | 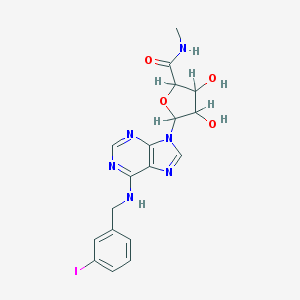 AC1L1GG2 AC1L1GG2 | C18H19IN6O4 | 510.292 | 8 / 4 | 0.9 | No |
| 127524 | 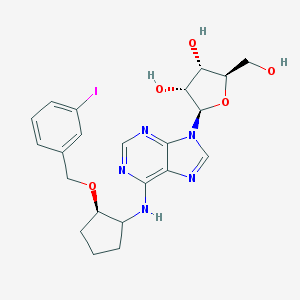 CHEMBL3679387 CHEMBL3679387 | C22H26IN5O5 | 567.384 | 9 / 4 | 2.6 | No |
| 127526 | 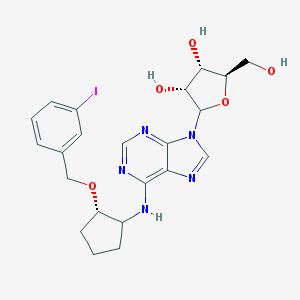 CHEMBL3679380 CHEMBL3679380 | C22H26IN5O5 | 567.384 | 9 / 4 | 2.6 | No |
| 145766 | 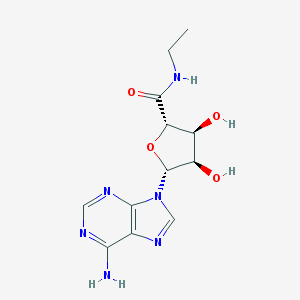 NECA NECA | C12H16N6O4 | 308.298 | 8 / 4 | -0.7 | Yes |
| 145787 | 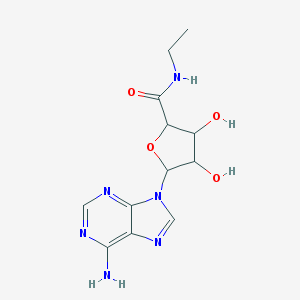 SMR000104521 SMR000104521 | C12H16N6O4 | 308.298 | 8 / 4 | -0.7 | Yes |
| 153698 | 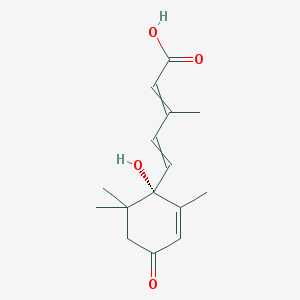 (+)-Abscisic acid (+)-Abscisic acid | C15H20O4 | 264.321 | 4 / 2 | 1.6 | Yes |
| 154976 | 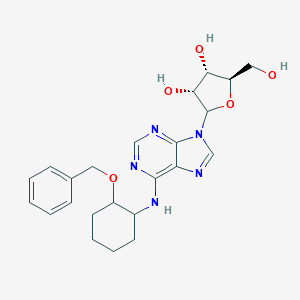 US8501708, 15 US8501708, 15 | C23H29N5O5 | 455.515 | 9 / 4 | 2.3 | Yes |
| 198805 | 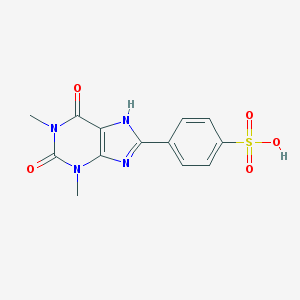 8-(p-Sulfophenyl)theophylline 8-(p-Sulfophenyl)theophylline | C13H12N4O5S | 336.322 | 6 / 2 | 1.6 | Yes |
| 234882 | 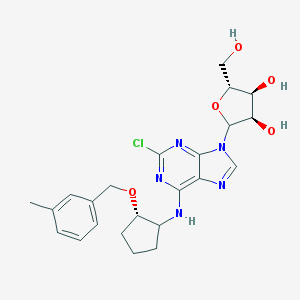 CHEMBL3679382 CHEMBL3679382 | C23H28ClN5O5 | 489.957 | 9 / 4 | 2.4 | Yes |
| 248445 | 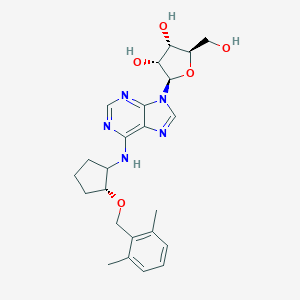 CHEMBL3679389 CHEMBL3679389 | C24H31N5O5 | 469.542 | 9 / 4 | 2.7 | Yes |
| 267299 | 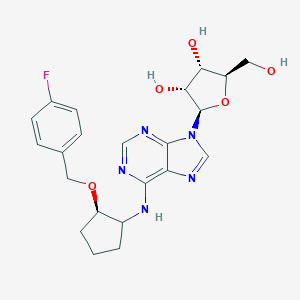 CHEMBL3679385 CHEMBL3679385 | C22H26FN5O5 | 459.478 | 10 / 4 | 2.0 | Yes |
| 267301 | 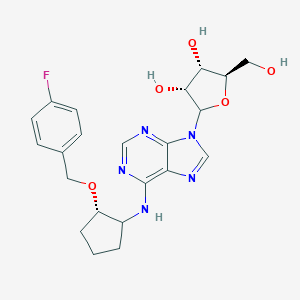 CHEMBL3679378 CHEMBL3679378 | C22H26FN5O5 | 459.478 | 10 / 4 | 2.0 | Yes |
| 294089 | 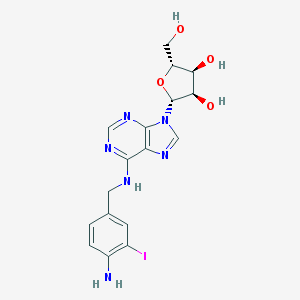 IABA IABA | C17H19IN6O4 | 498.281 | 9 / 5 | 1.1 | Yes |
| 297126 | 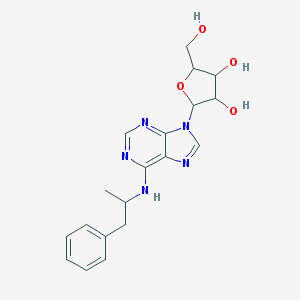 CHEMBL274022 CHEMBL274022 | C19H23N5O4 | 385.424 | 8 / 4 | 2.0 | Yes |
| 311163 | 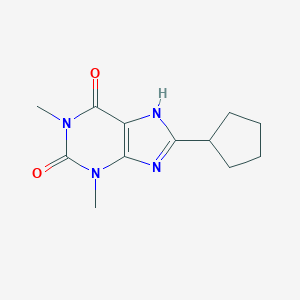 8-Cyclopentyl-1,3-dimethylxanthine 8-Cyclopentyl-1,3-dimethylxanthine | C12H16N4O2 | 248.286 | 3 / 1 | 1.6 | Yes |
| 320779 | 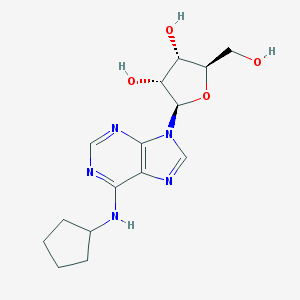 N6-Cyclopentyladenosine N6-Cyclopentyladenosine | C15H21N5O4 | 335.364 | 8 / 4 | 0.9 | Yes |
| 320781 | 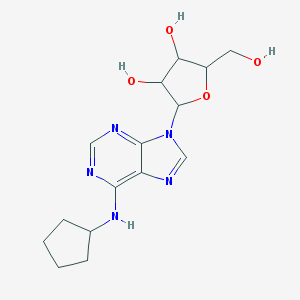 n-cyclopentyl-9-pentofuranosyl-9h-purin-6-amine n-cyclopentyl-9-pentofuranosyl-9h-purin-6-amine | C15H21N5O4 | 335.364 | 8 / 4 | 0.9 | Yes |
| 332230 | 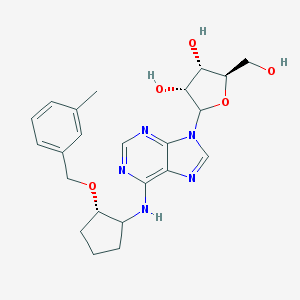 CHEMBL3674592 CHEMBL3674592 | C23H29N5O5 | 455.515 | 9 / 4 | 2.3 | Yes |
| 336089 | 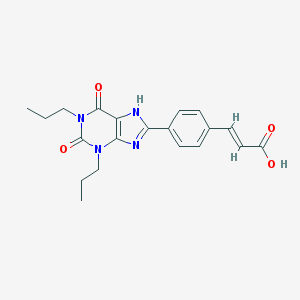 BW-A1433 BW-A1433 | C20H22N4O4 | 382.42 | 5 / 2 | 3.9 | Yes |
| 336099 | 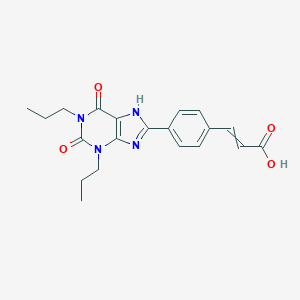 CID 129447 CID 129447 | C20H22N4O4 | 382.42 | 5 / 2 | 3.9 | Yes |
| 399824 | 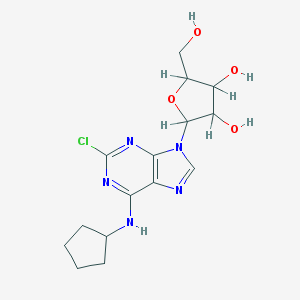 AC1L1BPF AC1L1BPF | C15H20ClN5O4 | 369.806 | 8 / 4 | 1.8 | Yes |
| 400084 | 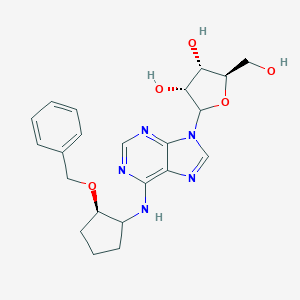 CHEMBL3674589 CHEMBL3674589 | C22H27N5O5 | 441.488 | 9 / 4 | 1.9 | Yes |
| 400603 | 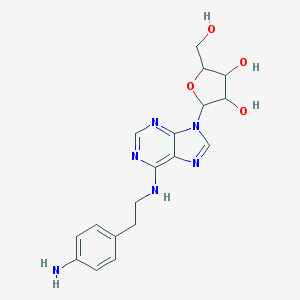 N6-2-(4-Aminophenyl)ethyladenosine N6-2-(4-Aminophenyl)ethyladenosine | C18H22N6O4 | 386.412 | 9 / 5 | 0.9 | Yes |
| 427521 | 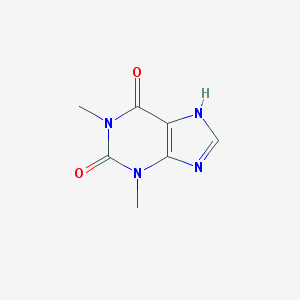 theophylline theophylline | C7H8N4O2 | 180.167 | 3 / 1 | 0.0 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218