

You can:
| Name | Cannabinoid receptor 2 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | CNR2 |
| Synonym | Peripheral cannabinoid receptor rCB2 hCB2 cannabinoid receptor 2 (macrophage) cannabinoid receptor 2 (spleen) [ Show all ] |
| Disease | Immune disorder Inflammatory bowel disease Inflammatory disease Neuropathic pain Osteoporosis [ Show all ] |
| Length | 360 |
| Amino acid sequence | MEECWVTEIANGSKDGLDSNPMKDYMILSGPQKTAVAVLCTLLGLLSALENVAVLYLILSSHQLRRKPSYLFIGSLAGADFLASVVFACSFVNFHVFHGVDSKAVFLLKIGSVTMTFTASVGSLLLTAIDRYLCLRYPPSYKALLTRGRALVTLGIMWVLSALVSYLPLMGWTCCPRPCSELFPLIPNDYLLSWLLFIAFLFSGIIYTYGHVLWKAHQHVASLSGHQDRQVPGMARMRLDVRLAKTLGLVLAVLLICWFPVLALMAHSLATTLSDQVKKAFAFCSMLCLINSMVNPVIYALRSGEIRSSAHHCLAHWKKCVRGLGSEAKEEAPRSSVTETEADGKITPWPDSRDLDLSDC |
| UniProt | P34972 |
| Protein Data Bank | 5zty |
| GPCR-HGmod model | P34972 |
| 3D structure model | This structure is from PDB ID 5zty. |
| BioLiP | BL0438927 |
| Therapeutic Target Database | T37693 |
| ChEMBL | CHEMBL253 |
| IUPHAR | 57 |
| DrugBank | BE0000095 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 16 | 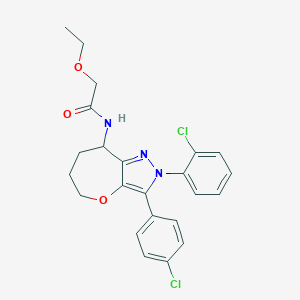 CHEMBL2063242 CHEMBL2063242 | C23H23Cl2N3O3 | 460.355 | 4 / 1 | 4.7 | Yes |
| 26 | 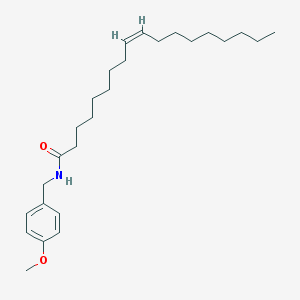 N-(4-methoxybenzyl)oleamide N-(4-methoxybenzyl)oleamide | C26H43NO2 | 401.635 | 2 / 1 | 8.5 | No |
| 235 | 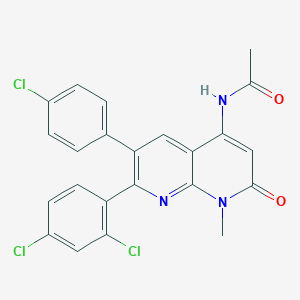 CHEMBL204391 CHEMBL204391 | C23H16Cl3N3O2 | 472.75 | 3 / 1 | 4.8 | Yes |
| 278 | 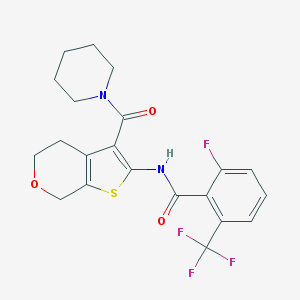 CHEMBL2205617 CHEMBL2205617 | C21H20F4N2O3S | 456.456 | 8 / 1 | 4.3 | Yes |
| 441683 | 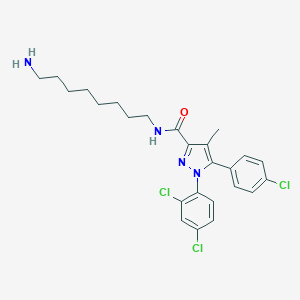 CHEMBL3322349 CHEMBL3322349 | C25H29Cl3N4O | 507.884 | 3 / 2 | 7.0 | No |
| 456 | 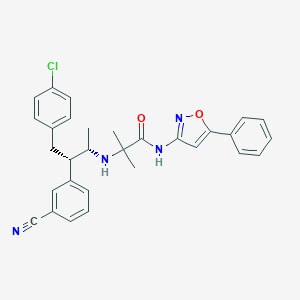 CHEMBL572080 CHEMBL572080 | C30H29ClN4O2 | 513.038 | 5 / 2 | 6.2 | No |
| 465 | 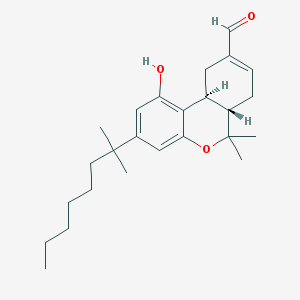 CHEMBL294472 CHEMBL294472 | C25H36O3 | 384.56 | 3 / 1 | 7.0 | No |
| 466 | 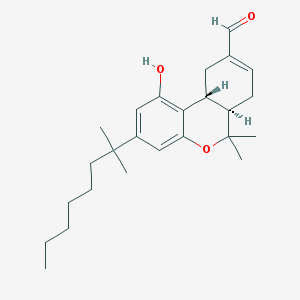 CHEMBL59540 CHEMBL59540 | C25H36O3 | 384.56 | 3 / 1 | 7.0 | No |
| 505 | 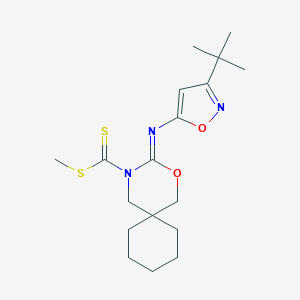 CHEMBL481508 CHEMBL481508 | C18H27N3O2S2 | 381.553 | 6 / 0 | 5.8 | No |
| 558 | 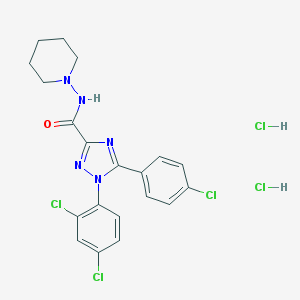 CHEMBL558564 CHEMBL558564 | C20H20Cl5N5O | 523.664 | 4 / 3 | N/A | No |
| 533915 | 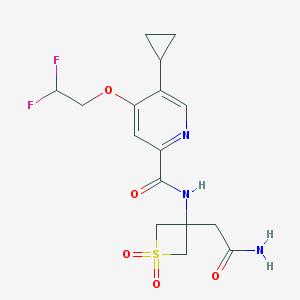 CHEMBL3937055 CHEMBL3937055 | C16H19F2N3O5S | 403.401 | 8 / 2 | 0.6 | Yes |
| 588 | 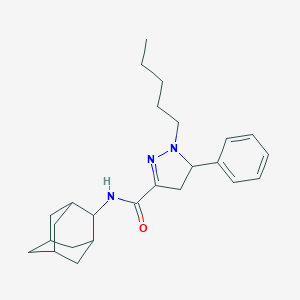 CHEMBL1223776 CHEMBL1223776 | C25H35N3O | 393.575 | 3 / 1 | 5.4 | No |
| 606 | 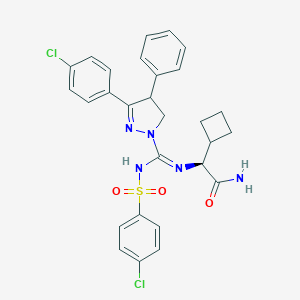 CHEMBL2153673 CHEMBL2153673 | C28H27Cl2N5O3S | 584.516 | 5 / 2 | 5.6 | No |
| 519701 | 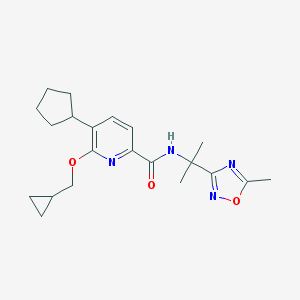 CHEMBL3924062 CHEMBL3924062 | C21H28N4O3 | 384.48 | 6 / 1 | 4.2 | Yes |
| 666 | 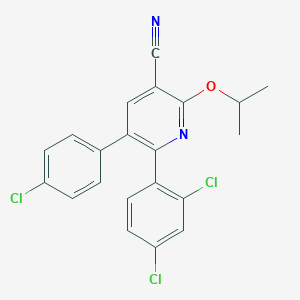 CHEMBL392836 CHEMBL392836 | C21H15Cl3N2O | 417.714 | 3 / 0 | 6.8 | No |
| 723 | 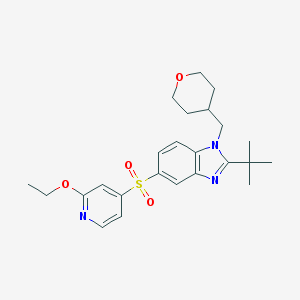 CHEMBL411504 CHEMBL411504 | C24H31N3O4S | 457.589 | 6 / 0 | 4.1 | Yes |
| 519703 | 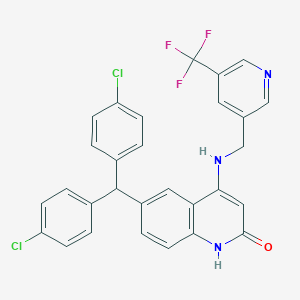 CHEMBL3915356 CHEMBL3915356 | C29H20Cl2F3N3O | 554.394 | 6 / 2 | 7.0 | No |
| 835 | 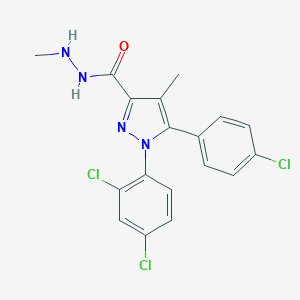 CHEMBL88548 CHEMBL88548 | C18H15Cl3N4O | 409.695 | 3 / 2 | 5.2 | No |
| 858 | 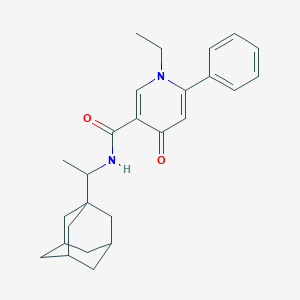 CHEMBL1641939 CHEMBL1641939 | C26H32N2O2 | 404.554 | 3 / 1 | 5.7 | No |
| 535936 | 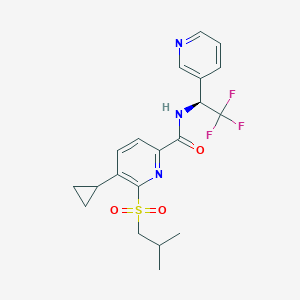 CHEMBL3936958 CHEMBL3936958 | C20H22F3N3O3S | 441.469 | 8 / 1 | 3.8 | Yes |
| 535938 | 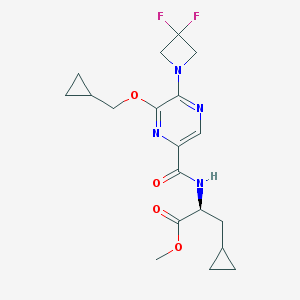 CHEMBL3903525 CHEMBL3903525 | C19H24F2N4O4 | 410.422 | 9 / 1 | 2.8 | Yes |
| 1066 | 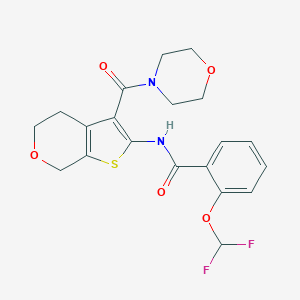 CHEMBL2205606 CHEMBL2205606 | C20H20F2N2O5S | 438.446 | 8 / 1 | 3.1 | Yes |
| 459243 | 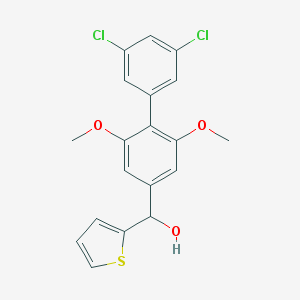 CHEMBL1076680 CHEMBL1076680 | C19H16Cl2O3S | 395.294 | 4 / 1 | 5.3 | No |
| 1171 | 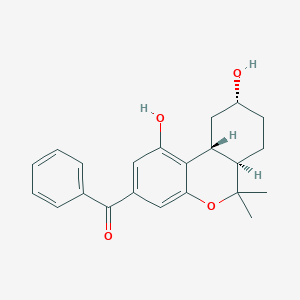 CHEMBL2064052 CHEMBL2064052 | C22H24O4 | 352.43 | 4 / 2 | 3.8 | Yes |
| 1181 | 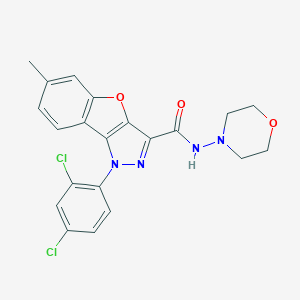 CHEMBL3290443 CHEMBL3290443 | C21H18Cl2N4O3 | 445.3 | 5 / 1 | 4.8 | Yes |
| 1312 | 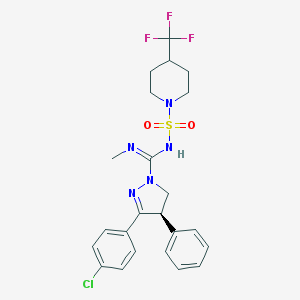 CHEMBL1080908 CHEMBL1080908 | C23H25ClF3N5O2S | 527.991 | 8 / 1 | 4.8 | No |
| 1315 | 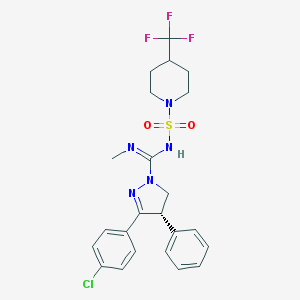 CHEMBL1080545 CHEMBL1080545 | C23H25ClF3N5O2S | 527.991 | 8 / 1 | 4.8 | No |
| 1316 | 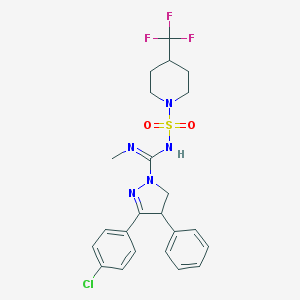 CHEMBL1081213 CHEMBL1081213 | C23H25ClF3N5O2S | 527.991 | 8 / 1 | 4.8 | No |
| 1427 | 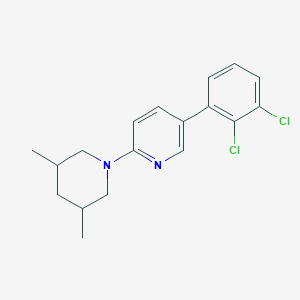 CHEMBL570193 CHEMBL570193 | C18H20Cl2N2 | 335.272 | 2 / 0 | 5.9 | No |
| 519707 | 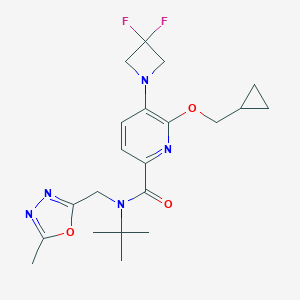 CHEMBL3908425 CHEMBL3908425 | C21H27F2N5O3 | 435.476 | 9 / 0 | 3.0 | Yes |
| 1552 | 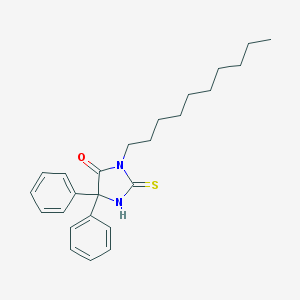 875014-22-5 875014-22-5 | C25H32N2OS | 408.604 | 2 / 1 | 7.4 | No |
| 1604 | 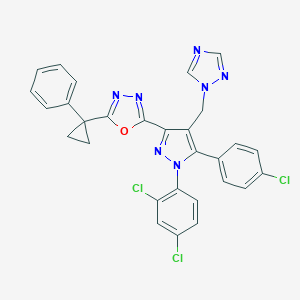 CHEMBL488787 CHEMBL488787 | C29H20Cl3N7O | 588.877 | 6 / 0 | 6.9 | No |
| 1672 | 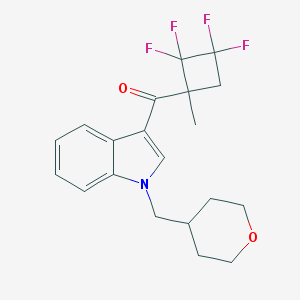 CHEMBL584528 CHEMBL584528 | C20H21F4NO2 | 383.387 | 6 / 0 | 4.0 | Yes |
| 1682 | 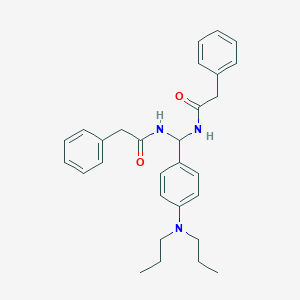 CHEMBL2179751 CHEMBL2179751 | C29H35N3O2 | 457.618 | 3 / 2 | 5.6 | No |
| 535963 | 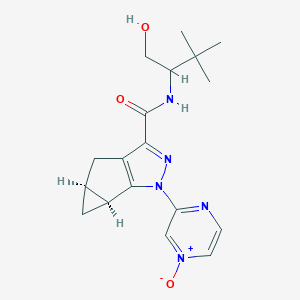 CHEMBL3900220 CHEMBL3900220 | C18H23N5O3 | 357.414 | 5 / 2 | 0.3 | Yes |
| 2006 | 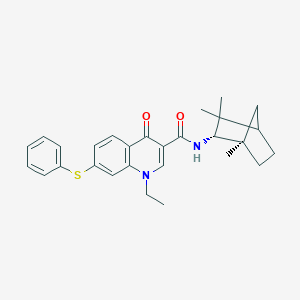 CHEMBL2152810 CHEMBL2152810 | C28H32N2O2S | 460.636 | 4 / 1 | 6.6 | No |
| 2024 | 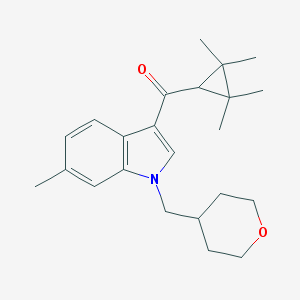 CHEMBL271839 CHEMBL271839 | C23H31NO2 | 353.506 | 2 / 0 | 4.7 | Yes |
| 535970 | 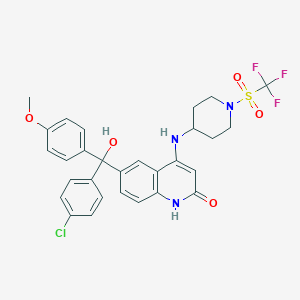 CHEMBL3925964 CHEMBL3925964 | C29H27ClF3N3O5S | 622.056 | 10 / 3 | 5.0 | No |
| 2042 | 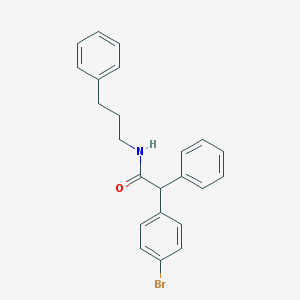 CHEMBL478643 CHEMBL478643 | C23H22BrNO | 408.339 | 1 / 1 | 5.8 | No |
| 535976 | 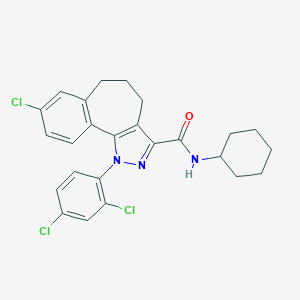 CHEMBL225622 CHEMBL225622 | C25H24Cl3N3O | 488.837 | 2 / 1 | 7.7 | No |
| 2092 | 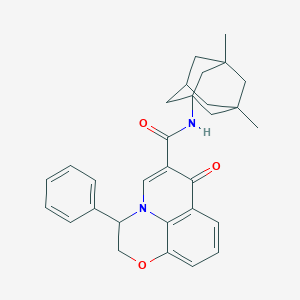 CHEMBL2163952 CHEMBL2163952 | C30H32N2O3 | 468.597 | 4 / 1 | 5.9 | No |
| 547921 | 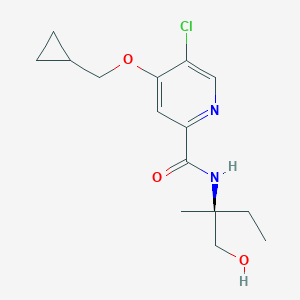 CHEMBL3911884 CHEMBL3911884 | C15H21ClN2O3 | 312.794 | 4 / 2 | 2.3 | Yes |
| 557363 | 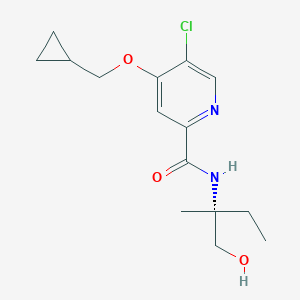 SCHEMBL15790159 SCHEMBL15790159 | C15H21ClN2O3 | 312.794 | 4 / 2 | 2.3 | Yes |
| 535977 | 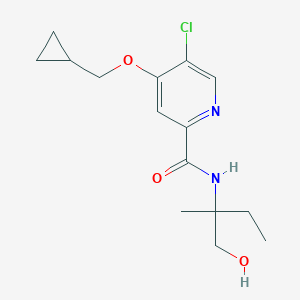 CHEMBL3911641 CHEMBL3911641 | C15H21ClN2O3 | 312.794 | 4 / 2 | 2.3 | Yes |
| 2349 | 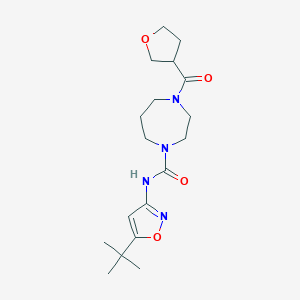 CHEMBL1762420 CHEMBL1762420 | C18H28N4O4 | 364.446 | 5 / 1 | 1.2 | Yes |
| 441767 | 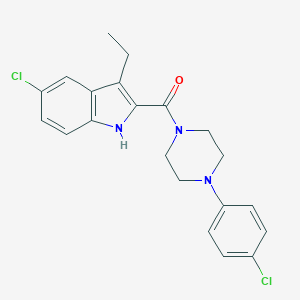 CHEMBL3401612 CHEMBL3401612 | C21H21Cl2N3O | 402.319 | 2 / 1 | 5.5 | No |
| 2364 | 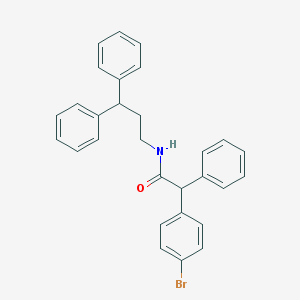 CHEMBL475910 CHEMBL475910 | C29H26BrNO | 484.437 | 1 / 1 | 7.3 | No |
| 2382 | 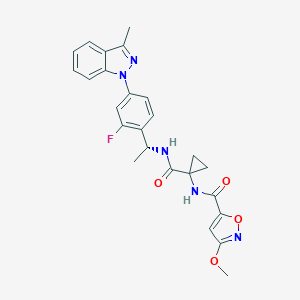 CHEMBL1801072 CHEMBL1801072 | C25H24FN5O4 | 477.496 | 7 / 2 | 3.7 | Yes |
| 535983 | 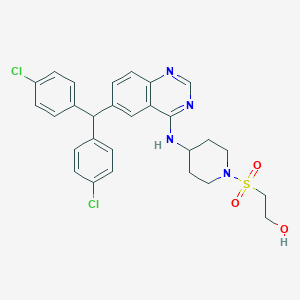 SCHEMBL17557589 SCHEMBL17557589 | C28H28Cl2N4O3S | 571.517 | 7 / 2 | 5.6 | No |
| 3007 | 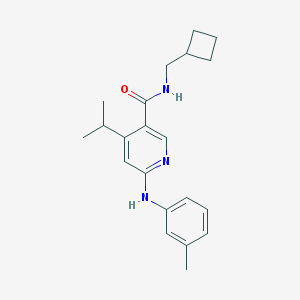 CHEMBL464916 CHEMBL464916 | C21H27N3O | 337.467 | 3 / 2 | 4.8 | Yes |
| 3010 | 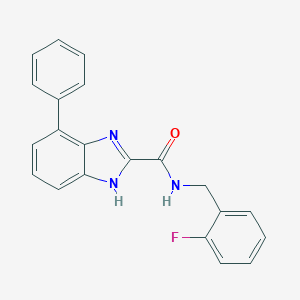 CHEMBL3114514 CHEMBL3114514 | C21H16FN3O | 345.377 | 3 / 2 | 4.3 | Yes |
| 463240 | 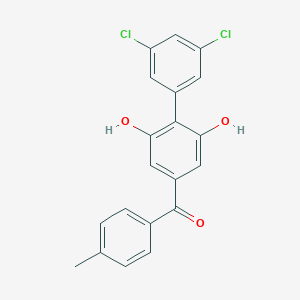 CHEMBL3605035 CHEMBL3605035 | C20H14Cl2O3 | 373.229 | 3 / 2 | 5.8 | No |
| 3066 | 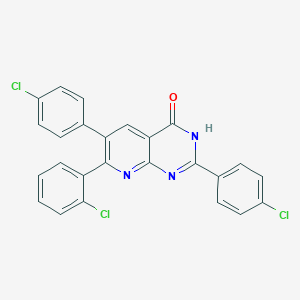 CHEMBL511558 CHEMBL511558 | C25H14Cl3N3O | 478.757 | 3 / 1 | 6.8 | No |
| 535986 | 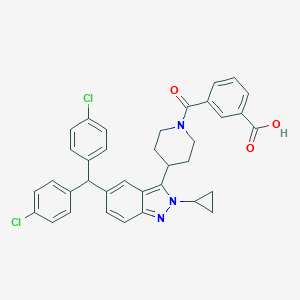 BDBM118149 BDBM118149 | C36H31Cl2N3O3 | 624.562 | 4 / 1 | 7.9 | No |
| 3072 | 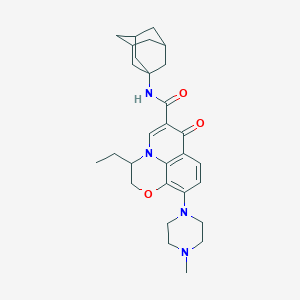 CHEMBL2163572 CHEMBL2163572 | C29H38N4O3 | 490.648 | 6 / 1 | 4.2 | Yes |
| 3089 | 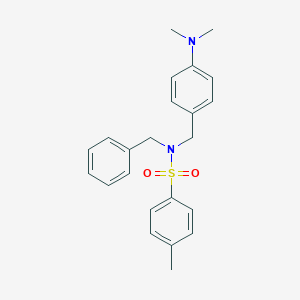 CHEMBL2336550 CHEMBL2336550 | C23H26N2O2S | 394.533 | 4 / 0 | 4.5 | Yes |
| 459252 | 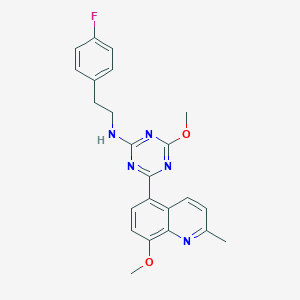 CHEMBL3669096 CHEMBL3669096 | C23H22FN5O2 | 419.46 | 8 / 1 | 5.0 | Yes |
| 3336 | 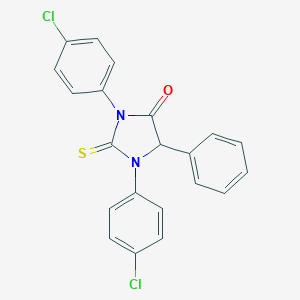 CID 11668762 CID 11668762 | C21H14Cl2N2OS | 413.316 | 2 / 0 | 6.0 | No |
| 536001 | 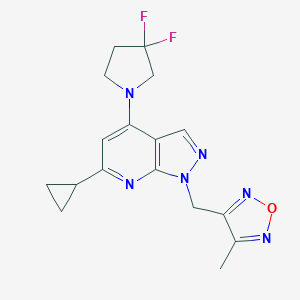 SCHEMBL16023414 SCHEMBL16023414 | C17H18F2N6O | 360.369 | 8 / 0 | 2.2 | Yes |
| 3475 | 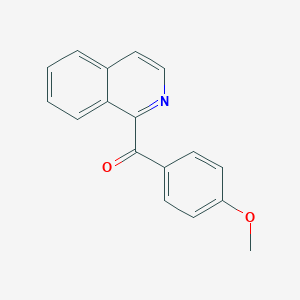 AC1LEVMS AC1LEVMS | C17H13NO2 | 263.296 | 3 / 0 | 3.7 | Yes |
| 536003 | 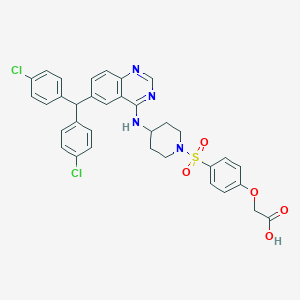 SCHEMBL17557608 SCHEMBL17557608 | C34H30Cl2N4O5S | 677.597 | 9 / 2 | 7.5 | No |
| 3518 | 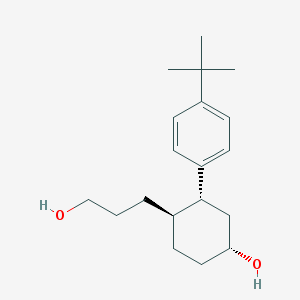 CHEMBL263632 CHEMBL263632 | C19H30O2 | 290.447 | 2 / 2 | 3.8 | Yes |
| 3543 | 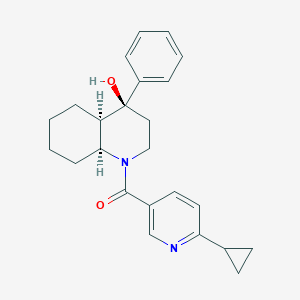 CHEMBL1765270 CHEMBL1765270 | C24H28N2O2 | 376.5 | 3 / 1 | 3.4 | Yes |
| 3563 | 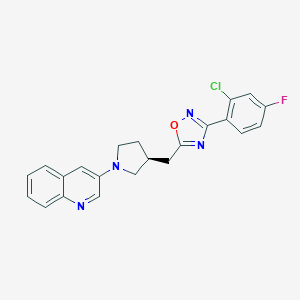 CHEMBL515648 CHEMBL515648 | C22H18ClFN4O | 408.861 | 6 / 0 | 5.3 | No |
| 3566 | 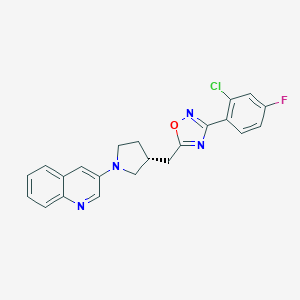 CHEMBL478202 CHEMBL478202 | C22H18ClFN4O | 408.861 | 6 / 0 | 5.3 | No |
| 3586 | 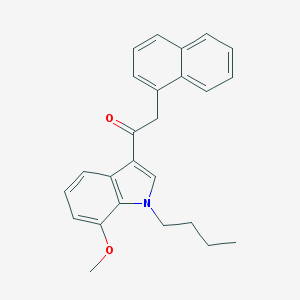 CHEMBL2380424 CHEMBL2380424 | C25H25NO2 | 371.48 | 2 / 0 | 5.7 | No |
| 536006 | 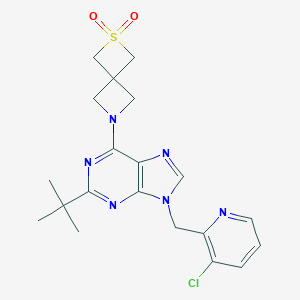 SCHEMBL16222656 SCHEMBL16222656 | C20H23ClN6O2S | 446.954 | 7 / 0 | 2.4 | Yes |
| 536011 | 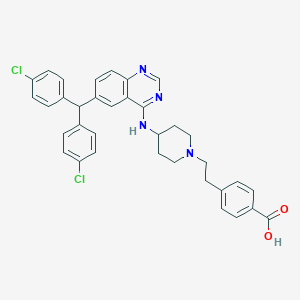 SCHEMBL17557600 SCHEMBL17557600 | C35H32Cl2N4O2 | 611.567 | 6 / 2 | 6.3 | No |
| 3736 | 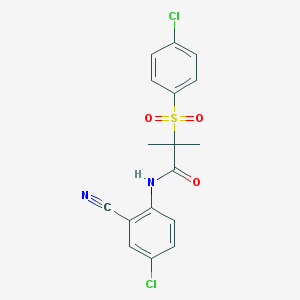 CHEMBL449341 CHEMBL449341 | C17H14Cl2N2O3S | 397.27 | 4 / 1 | 4.2 | Yes |
| 3923 | 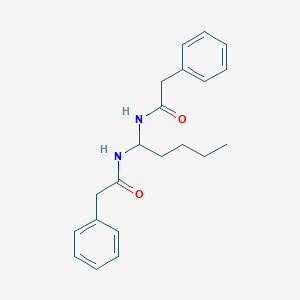 CHEMBL2179729 CHEMBL2179729 | C21H26N2O2 | 338.451 | 2 / 2 | 4.0 | Yes |
| 3970 | 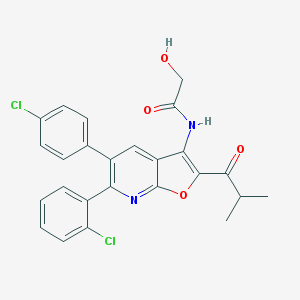 CHEMBL1091832 CHEMBL1091832 | C25H20Cl2N2O4 | 483.345 | 5 / 2 | 6.3 | No |
| 4003 | 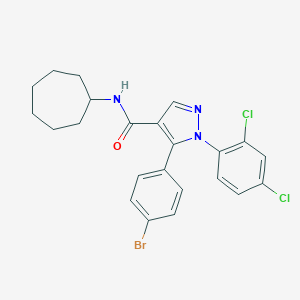 CHEMBL518505 CHEMBL518505 | C23H22BrCl2N3O | 507.253 | 2 / 1 | 7.1 | No |
| 4066 | 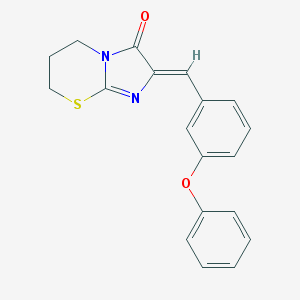 AC1LP5MH AC1LP5MH | C19H16N2O2S | 336.409 | 4 / 0 | 4.1 | Yes |
| 536033 | 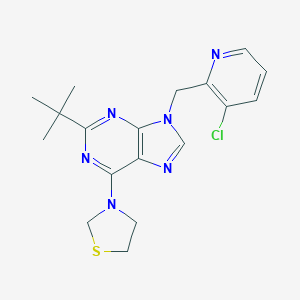 SCHEMBL16211797 SCHEMBL16211797 | C18H21ClN6S | 388.918 | 6 / 0 | 3.9 | Yes |
| 4146 | 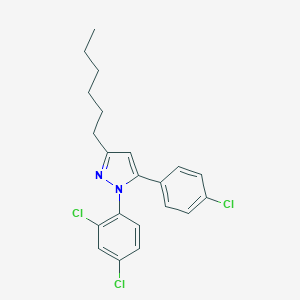 CHEMBL2347042 CHEMBL2347042 | C21H21Cl3N2 | 407.763 | 1 / 0 | 8.3 | No |
| 441879 | 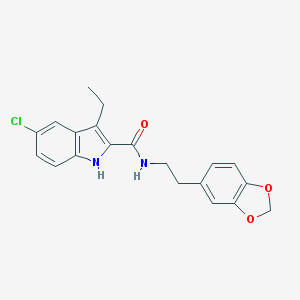 CHEMBL3401608 CHEMBL3401608 | C20H19ClN2O3 | 370.833 | 3 / 2 | 4.9 | Yes |
| 4187 | 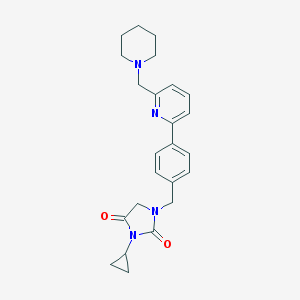 CID 56595575 CID 56595575 | C24H28N4O2 | 404.514 | 4 / 0 | 2.7 | Yes |
| 4194 | 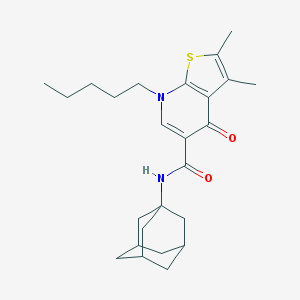 CHEMBL2316259 CHEMBL2316259 | C25H34N2O2S | 426.619 | 4 / 1 | 6.4 | No |
| 4261 | 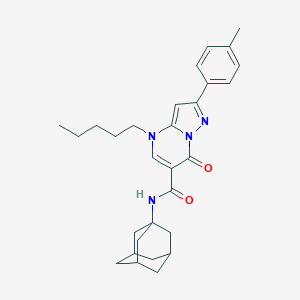 CHEMBL2381791 CHEMBL2381791 | C29H36N4O2 | 472.633 | 4 / 1 | 6.4 | No |
| 4365 | 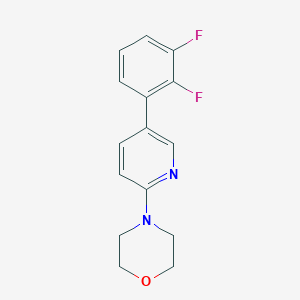 CHEMBL576709 CHEMBL576709 | C15H14F2N2O | 276.287 | 5 / 0 | 2.8 | Yes |
| 4423 | 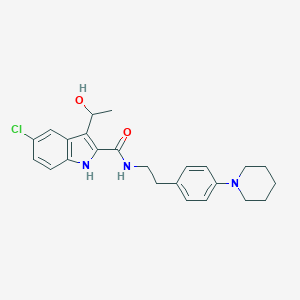 CHEMBL2071068 CHEMBL2071068 | C24H28ClN3O2 | 425.957 | 3 / 3 | 4.7 | Yes |
| 536045 | 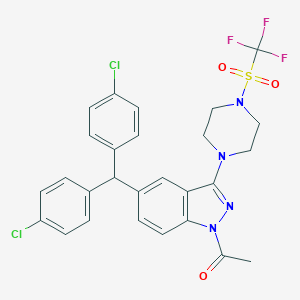 SCHEMBL18548029 SCHEMBL18548029 | C27H23Cl2F3N4O3S | 611.461 | 9 / 0 | 6.8 | No |
| 4456 | 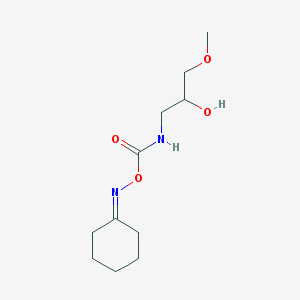 CHEMBL2172461 CHEMBL2172461 | C11H20N2O4 | 244.291 | 5 / 2 | 0.4 | Yes |
| 4554 | 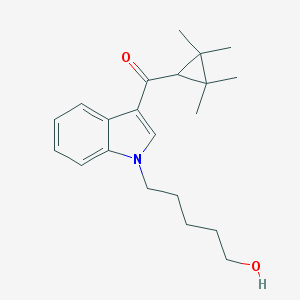 UNII-4MC625LVID UNII-4MC625LVID | C21H29NO2 | 327.468 | 2 / 1 | 4.1 | Yes |
| 441891 | 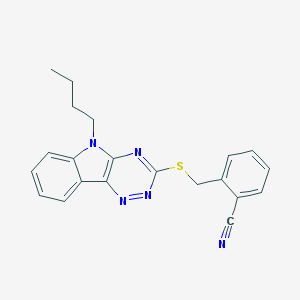 CHEMBL3353830 CHEMBL3353830 | C21H19N5S | 373.478 | 5 / 0 | 4.3 | Yes |
| 547950 | 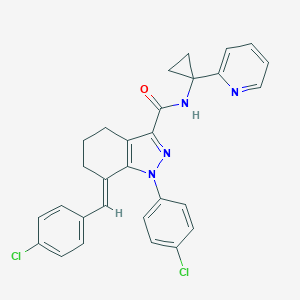 CHEMBL3930886 CHEMBL3930886 | C29H24Cl2N4O | 515.438 | 3 / 1 | 6.3 | No |
| 557415 | 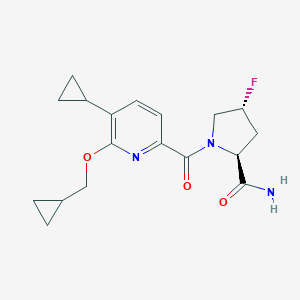 SCHEMBL15776122 SCHEMBL15776122 | C18H22FN3O3 | 347.39 | 5 / 1 | 2.1 | Yes |
| 547952 | 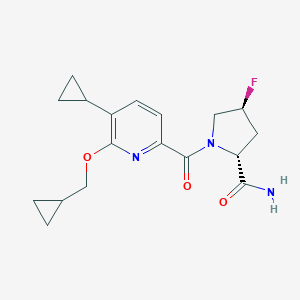 CHEMBL3891188 CHEMBL3891188 | C18H22FN3O3 | 347.39 | 5 / 1 | 2.1 | Yes |
| 536058 | 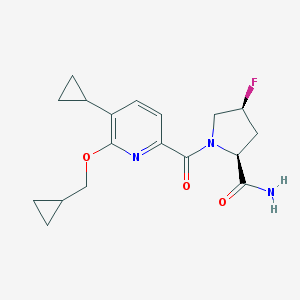 CHEMBL3898811 CHEMBL3898811 | C18H22FN3O3 | 347.39 | 5 / 1 | 2.1 | Yes |
| 536061 | 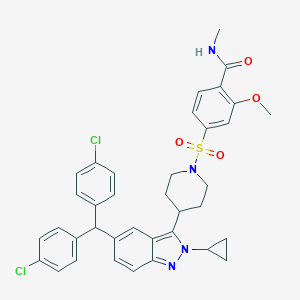 SCHEMBL18548061 SCHEMBL18548061 | C37H36Cl2N4O4S | 703.679 | 6 / 1 | 7.3 | No |
| 4861 | 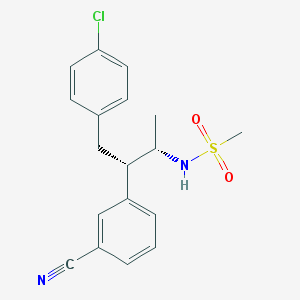 CHEMBL397690 CHEMBL397690 | C18H19ClN2O2S | 362.872 | 4 / 1 | 3.8 | Yes |
| 536064 | 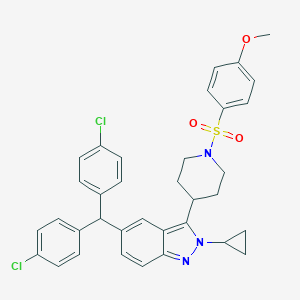 SCHEMBL18547958 SCHEMBL18547958 | C35H33Cl2N3O3S | 646.627 | 5 / 0 | 8.0 | No |
| 4991 | 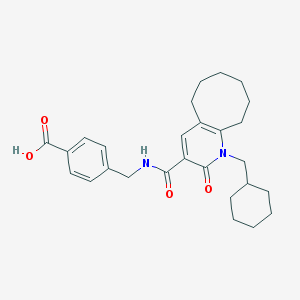 CHEMBL2023578 CHEMBL2023578 | C27H34N2O4 | 450.579 | 4 / 2 | 5.7 | No |
| 5072 | 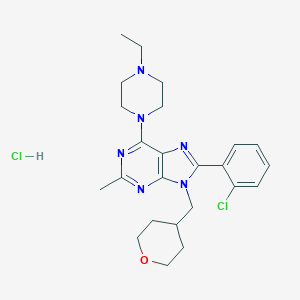 SCHEMBL2485239 SCHEMBL2485239 | C24H32Cl2N6O | 491.461 | 6 / 1 | N/A | N/A |
| 5073 | 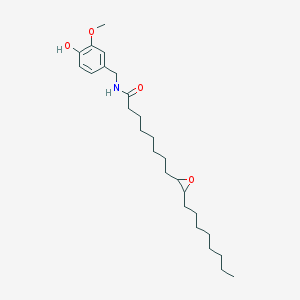 PhAR, Cycloproyl PhAR, Cycloproyl | C26H43NO4 | 433.633 | 4 / 2 | 7.0 | No |
| 5107 | 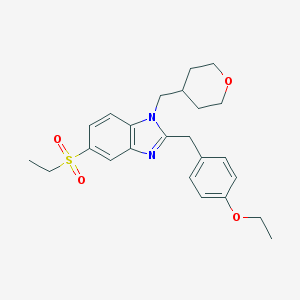 CHEMBL256781 CHEMBL256781 | C24H30N2O4S | 442.574 | 5 / 0 | 3.9 | Yes |
| 5118 | 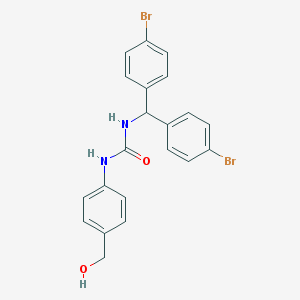 CHEMBL225414 CHEMBL225414 | C21H18Br2N2O2 | 490.195 | 2 / 3 | 4.6 | Yes |
| 5259 | 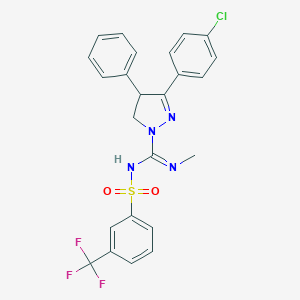 CHEMBL157591 CHEMBL157591 | C24H20ClF3N4O2S | 520.955 | 7 / 1 | 5.5 | No |
| 557425 | 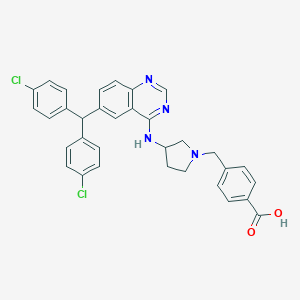 SCHEMBL17557708 SCHEMBL17557708 | C33H28Cl2N4O2 | 583.513 | 6 / 2 | 5.5 | No |
| 5331 | 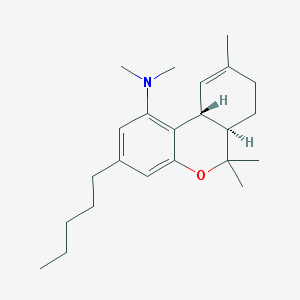 CHEMBL600426 CHEMBL600426 | C23H35NO | 341.539 | 2 / 0 | 7.4 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218