

You can:
| Name | Metabotropic glutamate receptor 7 |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Grm7 |
| Synonym | GLUR7 glutamate receptor GPRC1G mGlu7 receptor mGlu7a receptor [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 915 |
| Amino acid sequence | MVQLGKLLRVLTLMKFPCCVLEVLLCVLAAAARGQEMYAPHSIRIEGDVTLGGLFPVHAKGPSGVPCGDIKRENGIHRLEAMLYALDQINSDPNLLPNVTLGARILDTCSRDTYALEQSLTFVQALIQKDTSDVRCTNGEPPVFVKPEKVVGVIGASGSSVSIMVANILRLFQIPQISYASTAPELSDDRRYDFFSRVVPPDSFQAQAMVDIVKALGWNYVSTLASEGSYGEKGVESFTQISKEAGGLCIAQSVRIPQERKDRTIDFDRIIKQLLDTPNSRAVVIFANDEDIKQILAAAKRADQVGHFLWVGSDSWGSKINPLHQHEDIAEGAITIQPKRATVEGFDAYFTSRTLENNRRNVWFAEYWEENFNCKLTISGSKKEDTDRKCTGQERIGKDSNYEQEGKVQFVIDAVYAMAHALHHMNKDLCADYRGVCPEMEQAGGKKLLKYIRHVNFNGSAGTPVMFNKNGDAPGRYDIFQYQTTNTTNPGYRLIGQWTDELQLNIEDMQWGKGVREIPSSVCTLPCKPGQRKKTQKGTPCCWTCEPCDGYQYQFDEMTCQHCPYDQRPNENRTGCQNIPIIKLEWHSPWAVIPVFLAMLGIIATIFVMATFIRYNDTPIVRASGRELSYVLLTGIFLCYIITFLMIAKPDVAVCSFRRVFLGLGMCISYAALLTKTNRIYRIFEQGKKSVTAPRLISPTSQLAITSSLISVQLLGVFIWFGVDPPNIIIDYDEHKTMNPEQARGVLKCDITDLQIICSLGYSILLMVTCTVYAIKTRGVPENFNEAKPIGFTMYTTCIVWLAFIPIFFGTAQSAEKLYIQTTTLTISMNLSASVALGMLYMPKVYIIIFHPELNVQKRKRSFKAVVTAATMSSRLSHKPSDRPNGEAKTELCENVDPNSPAAKKKYVSYNNLVI |
| UniProt | P35400 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3879 |
| IUPHAR | 295 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 1109 | 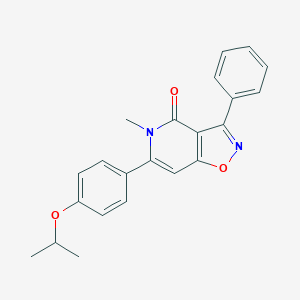 CHEMBL604891 CHEMBL604891 | C22H20N2O3 | 360.413 | 4 / 0 | 4.1 | Yes |
| 557354 | 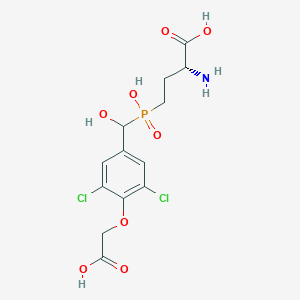 US9212196, Derivative 9 US9212196, Derivative 9 | C13H16Cl2NO8P | 416.144 | 9 / 5 | -2.6 | Yes |
| 535967 | 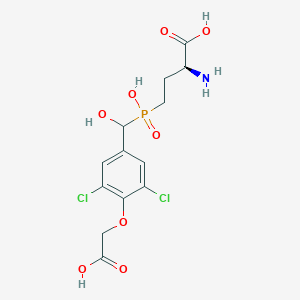 CHEMBL3974450 CHEMBL3974450 | C13H16Cl2NO8P | 416.144 | 9 / 5 | -2.6 | Yes |
| 557606 | 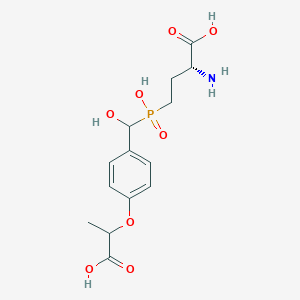 US9212196, Derivative 29 US9212196, Derivative 29 | C14H20NO8P | 361.287 | 9 / 5 | -3.4 | Yes |
| 536281 | 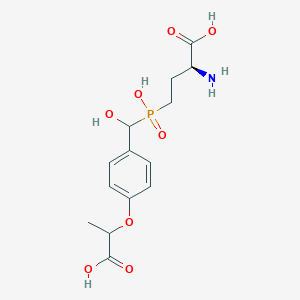 CHEMBL3967637 CHEMBL3967637 | C14H20NO8P | 361.287 | 9 / 5 | -3.4 | Yes |
| 12907 | 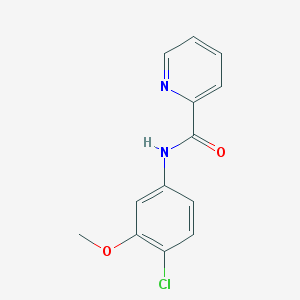 VU 0361737 VU 0361737 | C13H11ClN2O2 | 262.693 | 3 / 1 | 2.6 | Yes |
| 13321 | 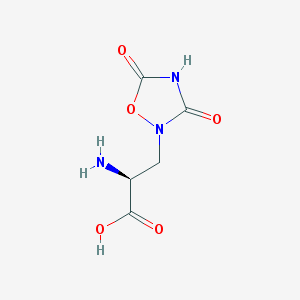 QUISQUALIC ACID QUISQUALIC ACID | C5H7N3O5 | 189.127 | 6 / 3 | -3.9 | Yes |
| 21874 | 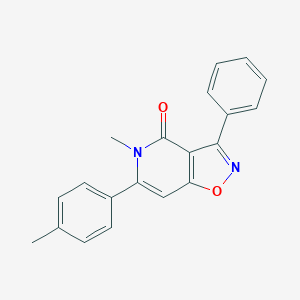 CHEMBL595825 CHEMBL595825 | C20H16N2O2 | 316.36 | 3 / 0 | 3.7 | Yes |
| 22843 | 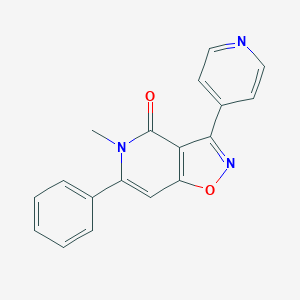 CHEMBL592956 CHEMBL592956 | C18H13N3O2 | 303.321 | 4 / 0 | 2.2 | Yes |
| 536573 | 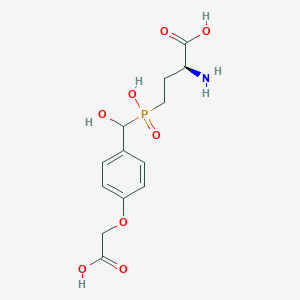 LSP4-2022 LSP4-2022 | C13H18NO8P | 347.26 | 9 / 5 | -3.8 | Yes |
| 522266 | 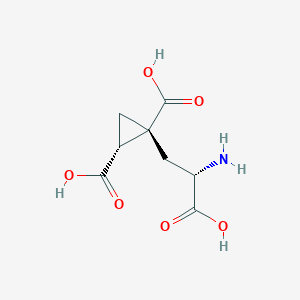 CHEMBL3786667 CHEMBL3786667 | C8H11NO6 | 217.177 | 7 / 4 | -4.0 | Yes |
| 522273 | 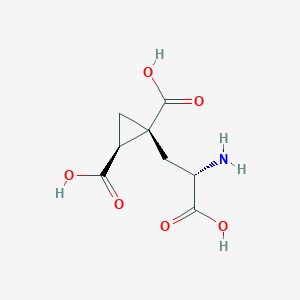 CHEMBL3787264 CHEMBL3787264 | C8H11NO6 | 217.177 | 7 / 4 | -4.0 | Yes |
| 522277 | 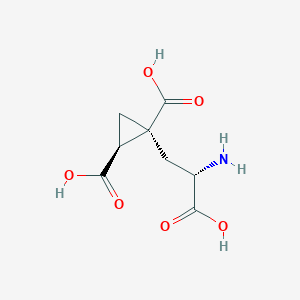 CHEMBL3786026 CHEMBL3786026 | C8H11NO6 | 217.177 | 7 / 4 | -4.0 | Yes |
| 522289 | 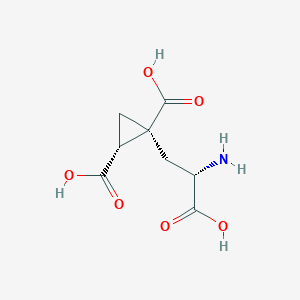 CHEMBL3786938 CHEMBL3786938 | C8H11NO6 | 217.177 | 7 / 4 | -4.0 | Yes |
| 25564 | 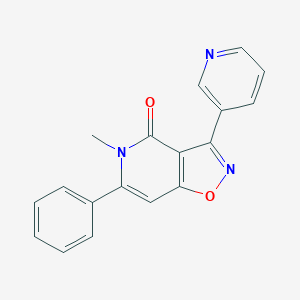 CHEMBL596274 CHEMBL596274 | C18H13N3O2 | 303.321 | 4 / 0 | 2.2 | Yes |
| 33518 | 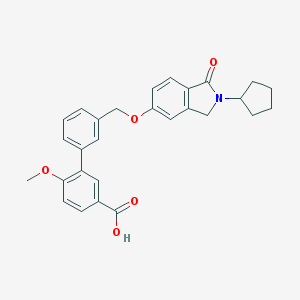 CHEMBL2179631 CHEMBL2179631 | C28H27NO5 | 457.526 | 5 / 1 | 4.8 | Yes |
| 33670 | 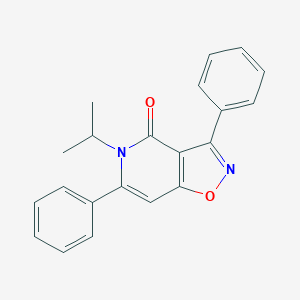 CHEMBL603652 CHEMBL603652 | C21H18N2O2 | 330.387 | 3 / 0 | 4.1 | Yes |
| 36690 | 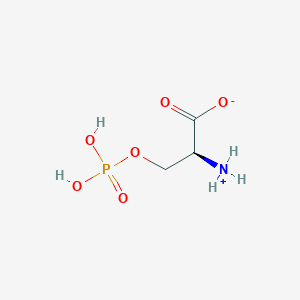 CID 57689797 CID 57689797 | C3H8NO6P | 185.072 | 6 / 3 | -4.5 | Yes |
| 558421 | 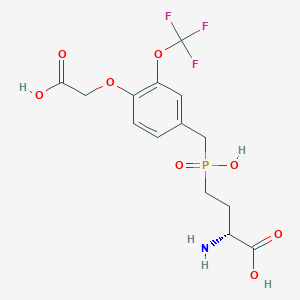 US9212196, Derivative 32 US9212196, Derivative 32 | C14H17F3NO8P | 415.258 | 12 / 4 | -2.1 | No |
| 536937 | 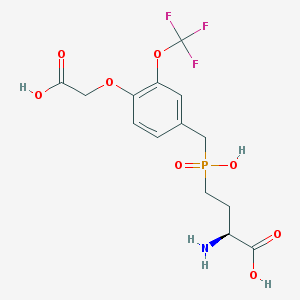 CHEMBL3910941 CHEMBL3910941 | C14H17F3NO8P | 415.258 | 12 / 4 | -2.1 | No |
| 536961 | 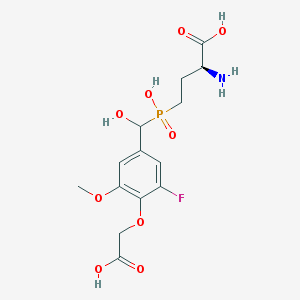 CHEMBL3898023 CHEMBL3898023 | C14H19FNO9P | 395.276 | 11 / 5 | -3.8 | No |
| 558442 | 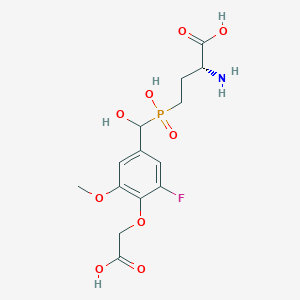 US9212196, Derivative 6 US9212196, Derivative 6 | C14H19FNO9P | 395.276 | 11 / 5 | -3.8 | No |
| 42858 | 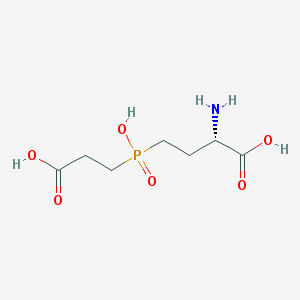 CHEMBL1089515 CHEMBL1089515 | C7H14NO6P | 239.164 | 7 / 4 | -4.8 | Yes |
| 537083 | 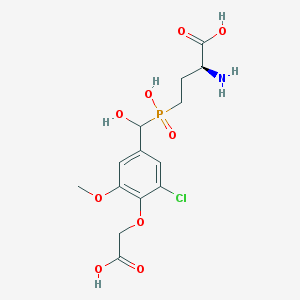 CHEMBL3969216 CHEMBL3969216 | C14H19ClNO9P | 411.728 | 10 / 5 | -3.2 | Yes |
| 558601 | 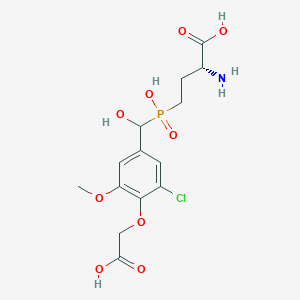 US9212196, Derivative 7 US9212196, Derivative 7 | C14H19ClNO9P | 411.728 | 10 / 5 | -3.2 | Yes |
| 553469 | 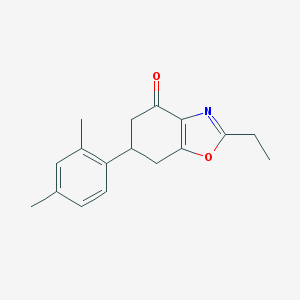 ADX-71743 ADX-71743 | C17H19NO2 | 269.344 | 3 / 0 | 3.8 | Yes |
| 519956 | 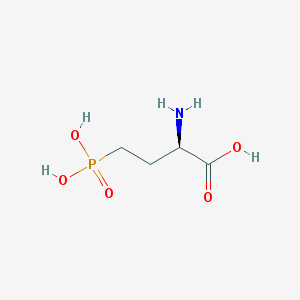 D-AP4 D-AP4 | C4H10NO5P | 183.1 | 6 / 4 | -5.5 | Yes |
| 57282 | 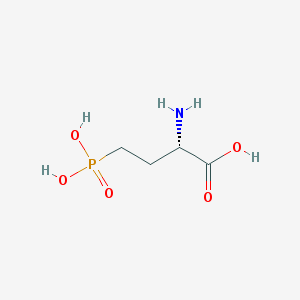 L-AP4 L-AP4 | C4H10NO5P | 183.1 | 6 / 4 | -5.5 | Yes |
| 537499 | 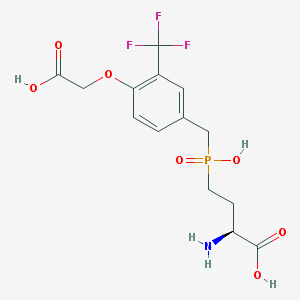 CHEMBL3980977 CHEMBL3980977 | C14H17F3NO7P | 399.259 | 11 / 4 | -2.4 | No |
| 559219 | 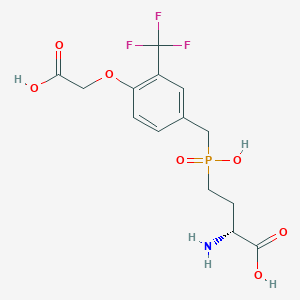 US9212196, Derivative 33 US9212196, Derivative 33 | C14H17F3NO7P | 399.259 | 11 / 4 | -2.4 | No |
| 63772 | 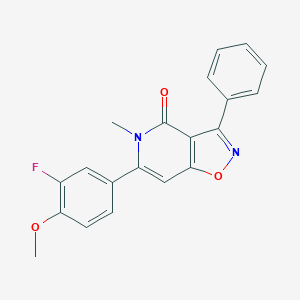 CHEMBL594648 CHEMBL594648 | C20H15FN2O3 | 350.349 | 5 / 0 | 3.4 | Yes |
| 66844 | 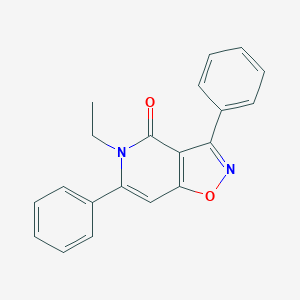 CHEMBL596305 CHEMBL596305 | C20H16N2O2 | 316.36 | 3 / 0 | 3.7 | Yes |
| 559477 | 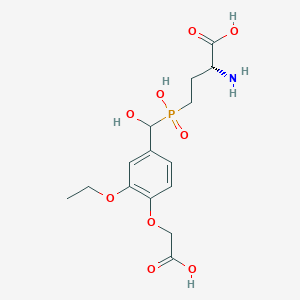 US9212196, Derivative 19 US9212196, Derivative 19 | C15H22NO9P | 391.313 | 10 / 5 | -3.5 | Yes |
| 537687 | 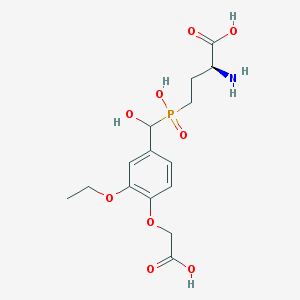 CHEMBL3941586 CHEMBL3941586 | C15H22NO9P | 391.313 | 10 / 5 | -3.5 | Yes |
| 82273 | 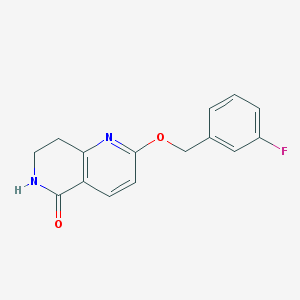 CHEMBL3298273 CHEMBL3298273 | C15H13FN2O2 | 272.279 | 4 / 1 | 2.1 | Yes |
| 85069 | 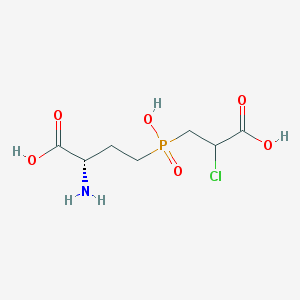 CHEMBL1076865 CHEMBL1076865 | C7H13ClNO6P | 273.606 | 7 / 4 | -4.1 | Yes |
| 97189 | 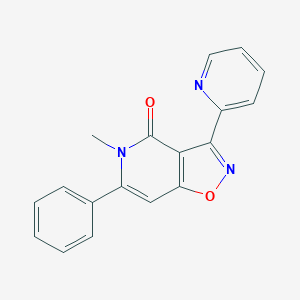 CHEMBL596275 CHEMBL596275 | C18H13N3O2 | 303.321 | 4 / 0 | 2.3 | Yes |
| 109307 | 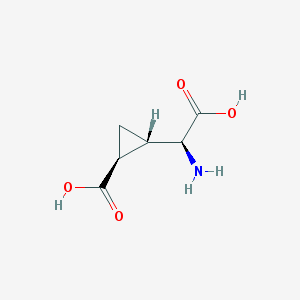 (2S,1'S,2'S)-2-(carboxycyclopropyl)glycine (2S,1'S,2'S)-2-(carboxycyclopropyl)glycine | C6H9NO4 | 159.141 | 5 / 3 | -3.4 | Yes |
| 538735 | 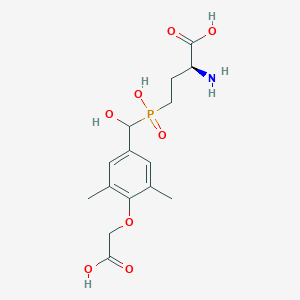 CHEMBL3912541 CHEMBL3912541 | C15H22NO8P | 375.314 | 9 / 5 | -3.1 | Yes |
| 560799 | 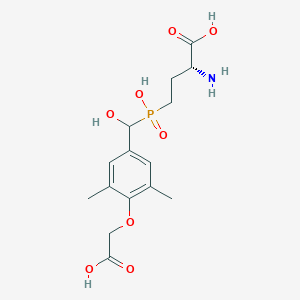 US9212196, Derivative 10 US9212196, Derivative 10 | C15H22NO8P | 375.314 | 9 / 5 | -3.1 | Yes |
| 538881 | 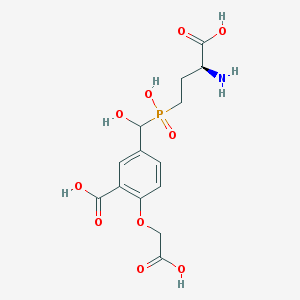 CHEMBL3927149 CHEMBL3927149 | C14H18NO10P | 391.269 | 11 / 6 | -4.3 | No |
| 560946 | 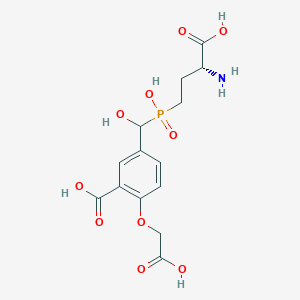 US9212196, Derivative 23 US9212196, Derivative 23 | C14H18NO10P | 391.269 | 11 / 6 | -4.3 | No |
| 538897 | 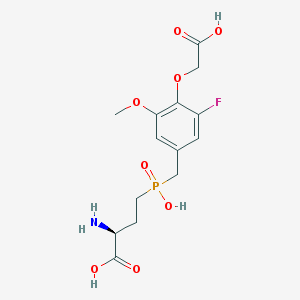 CHEMBL3913188 CHEMBL3913188 | C14H19FNO8P | 379.277 | 10 / 4 | -3.2 | Yes |
| 560961 | 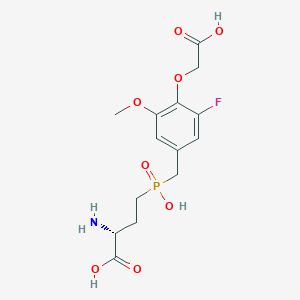 US9212196, Derivative 34 US9212196, Derivative 34 | C14H19FNO8P | 379.277 | 10 / 4 | -3.2 | Yes |
| 128447 | 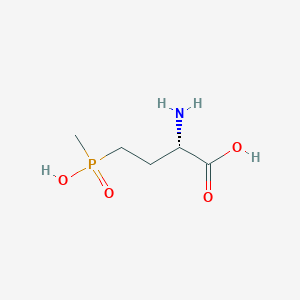 L-PHOSPHINOTHRICIN L-PHOSPHINOTHRICIN | C5H12NO4P | 181.128 | 5 / 3 | -5.0 | Yes |
| 130325 | 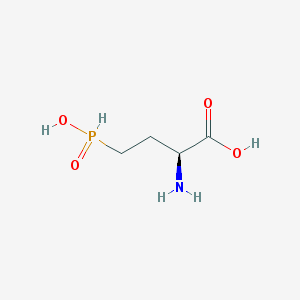 demethylphosphinothricin demethylphosphinothricin | C4H10NO4P | 167.101 | 5 / 3 | -5.1 | Yes |
| 135007 | 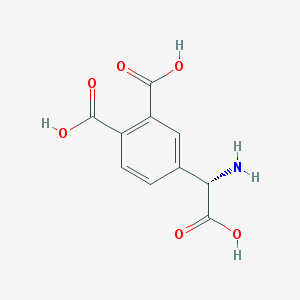 (S)-3,4-DCPG (S)-3,4-DCPG | C10H9NO6 | 239.183 | 7 / 4 | -2.7 | Yes |
| 135675 | 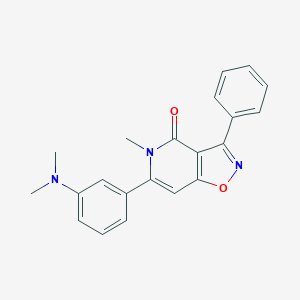 CHEMBL594223 CHEMBL594223 | C21H19N3O2 | 345.402 | 4 / 0 | 3.4 | Yes |
| 561524 | 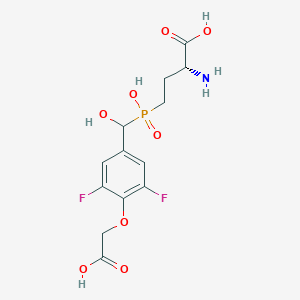 US9212196, Derivative 28 US9212196, Derivative 28 | C13H16F2NO8P | 383.241 | 11 / 5 | -3.6 | No |
| 539421 | 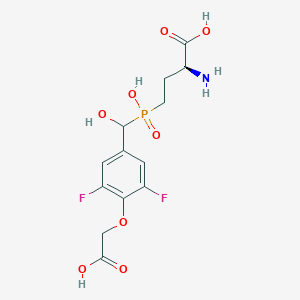 CHEMBL3896442 CHEMBL3896442 | C13H16F2NO8P | 383.241 | 11 / 5 | -3.6 | No |
| 539454 | 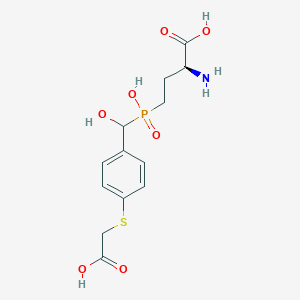 CHEMBL3905370 CHEMBL3905370 | C13H18NO7PS | 363.321 | 9 / 5 | -3.3 | Yes |
| 561554 | 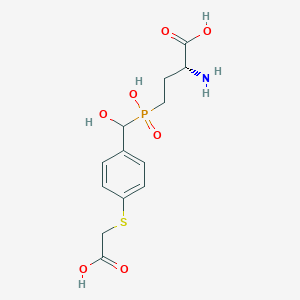 US9212196, Derivative 26 US9212196, Derivative 26 | C13H18NO7PS | 363.321 | 9 / 5 | -3.3 | Yes |
| 539799 | 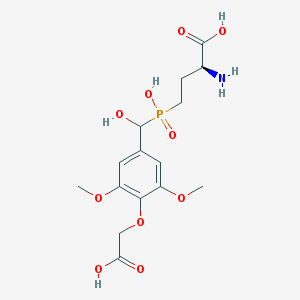 CHEMBL3982807 CHEMBL3982807 | C15H22NO10P | 407.312 | 11 / 5 | -3.9 | No |
| 561931 | 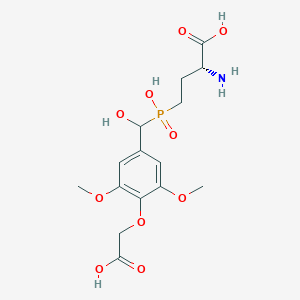 US9212196, Derivative 4 US9212196, Derivative 4 | C15H22NO10P | 407.312 | 11 / 5 | -3.9 | No |
| 158290 | 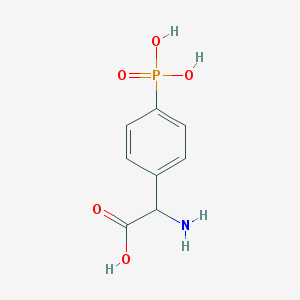 (RS)-PPG (RS)-PPG | C8H10NO5P | 231.144 | 6 / 4 | -3.7 | Yes |
| 184105 | 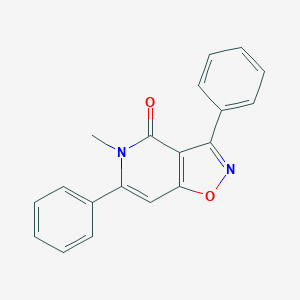 CHEMBL595840 CHEMBL595840 | C19H14N2O2 | 302.333 | 3 / 0 | 3.3 | Yes |
| 185818 | 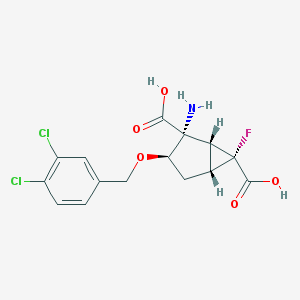 MGS-0039 MGS-0039 | C15H14Cl2FNO5 | 378.177 | 7 / 3 | -0.7 | Yes |
| 187013 | 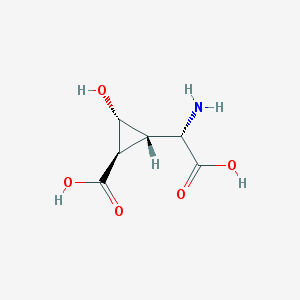 CHEMBL140197 CHEMBL140197 | C6H9NO5 | 175.14 | 6 / 4 | -3.9 | Yes |
| 189225 | 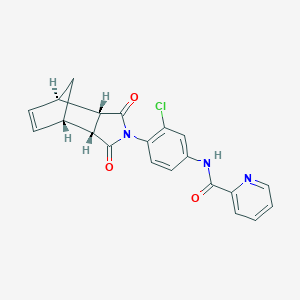 MLS003171619 MLS003171619 | C21H16ClN3O3 | 393.827 | 4 / 1 | 2.6 | Yes |
| 189233 | 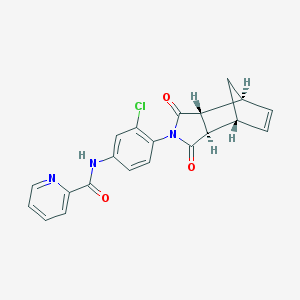 CHEMBL2373733 CHEMBL2373733 | C21H16ClN3O3 | 393.827 | 4 / 1 | 2.6 | Yes |
| 540924 | 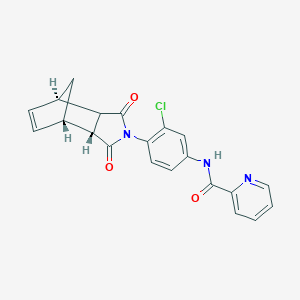 ML182 ML182 | C21H16ClN3O3 | 393.827 | 4 / 1 | 2.6 | Yes |
| 541005 | 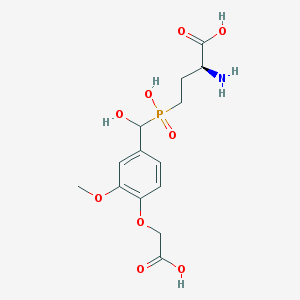 CHEMBL3935255 CHEMBL3935255 | C14H20NO9P | 377.286 | 10 / 5 | -3.9 | Yes |
| 563327 | 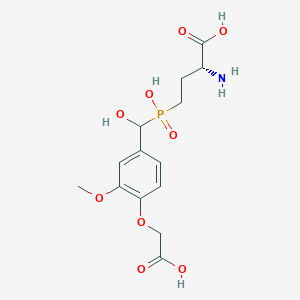 US9212196, Derivative 3 US9212196, Derivative 3 | C14H20NO9P | 377.286 | 10 / 5 | -3.9 | Yes |
| 195616 | 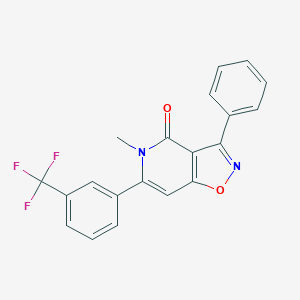 CHEMBL594448 CHEMBL594448 | C20H13F3N2O2 | 370.331 | 6 / 0 | 4.2 | Yes |
| 198457 | 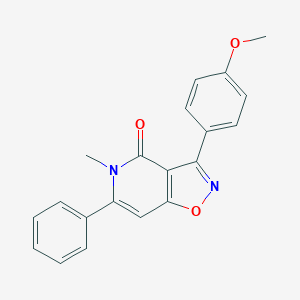 CHEMBL592954 CHEMBL592954 | C20H16N2O3 | 332.359 | 4 / 0 | 3.3 | Yes |
| 201027 | 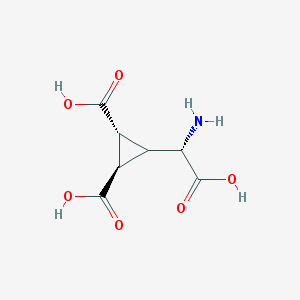 Dcg-IV Dcg-IV | C7H9NO6 | 203.15 | 7 / 4 | -4.3 | Yes |
| 213637 | 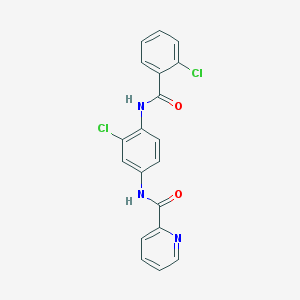 VU0366037-2 VU0366037-2 | C19H13Cl2N3O2 | 386.232 | 3 / 2 | 4.1 | Yes |
| 550720 | 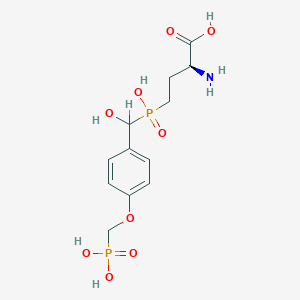 CHEMBL3894544 CHEMBL3894544 | C12H19NO9P2 | 383.23 | 10 / 6 | -5.2 | No |
| 564461 | 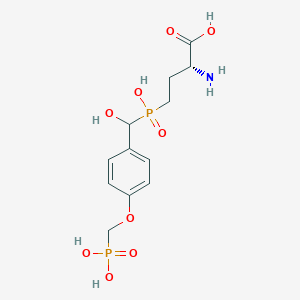 US9212196, Derivative 14 US9212196, Derivative 14 | C12H19NO9P2 | 383.23 | 10 / 6 | -5.2 | No |
| 230092 | 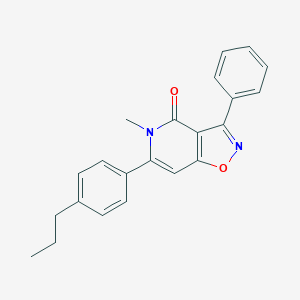 CHEMBL595388 CHEMBL595388 | C22H20N2O2 | 344.414 | 3 / 0 | 4.6 | Yes |
| 542100 | 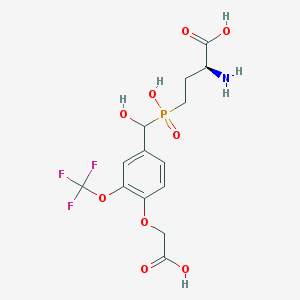 CHEMBL3890165 CHEMBL3890165 | C14H17F3NO9P | 431.257 | 13 / 5 | -2.7 | No |
| 564551 | 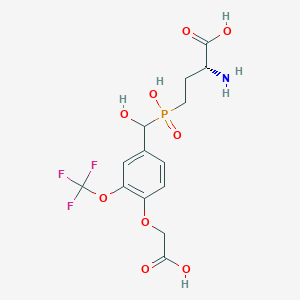 US9212196, Derivative 20 US9212196, Derivative 20 | C14H17F3NO9P | 431.257 | 13 / 5 | -2.7 | No |
| 235587 | 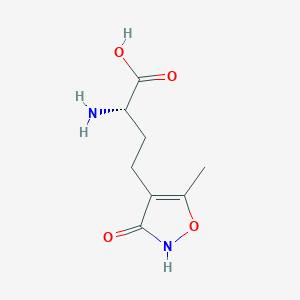 CHEMBL315268 CHEMBL315268 | C8H12N2O4 | 200.194 | 5 / 3 | -2.9 | Yes |
| 235602 | 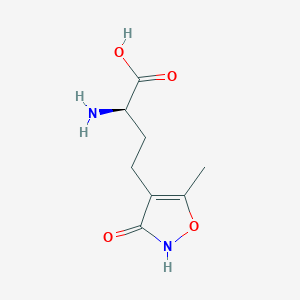 CHEMBL262416 CHEMBL262416 | C8H12N2O4 | 200.194 | 5 / 3 | -2.9 | Yes |
| 542298 | 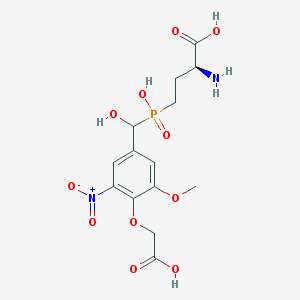 CHEMBL3966924 CHEMBL3966924 | C14H19N2O11P | 422.283 | 12 / 5 | -4.0 | No |
| 564753 | 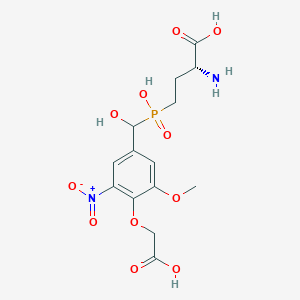 US9212196, Derivative 5 US9212196, Derivative 5 | C14H19N2O11P | 422.283 | 12 / 5 | -4.0 | No |
| 251317 | 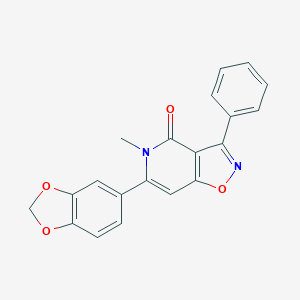 CHEMBL399603 CHEMBL399603 | C20H14N2O4 | 346.342 | 5 / 0 | 3.1 | Yes |
| 257353 | 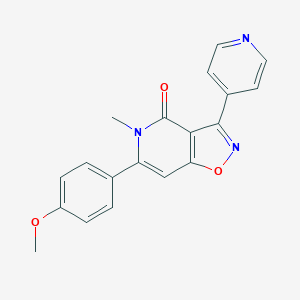 MMPIP MMPIP | C19H15N3O3 | 333.347 | 5 / 0 | 2.2 | Yes |
| 554557 | 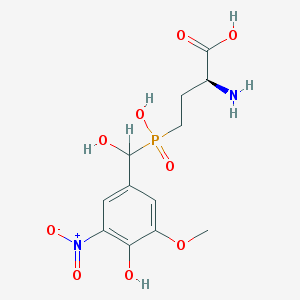 LSP1-2111 LSP1-2111 | C12H17N2O9P | 364.247 | 10 / 5 | -3.4 | Yes |
| 543033 | 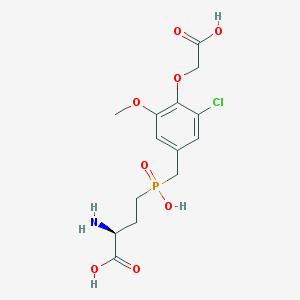 CHEMBL3909874 CHEMBL3909874 | C14H19ClNO8P | 395.729 | 9 / 4 | -2.7 | Yes |
| 565521 | 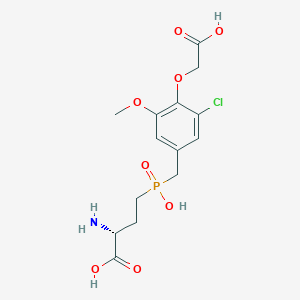 US9212196, Derivative 35 US9212196, Derivative 35 | C14H19ClNO8P | 395.729 | 9 / 4 | -2.7 | Yes |
| 266003 | 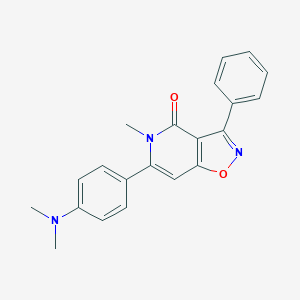 CHEMBL592957 CHEMBL592957 | C21H19N3O2 | 345.402 | 4 / 0 | 3.4 | Yes |
| 274917 | 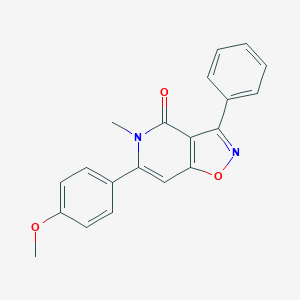 CHEMBL399602 CHEMBL399602 | C20H16N2O3 | 332.359 | 4 / 0 | 3.3 | Yes |
| 566052 | 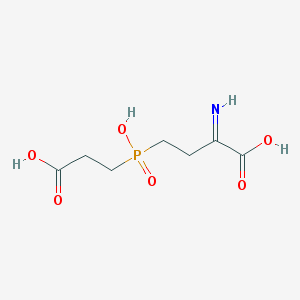 BDBM50314796 BDBM50314796 | C7H12NO6P | 237.148 | 7 / 4 | -2.1 | Yes |
| 292325 | 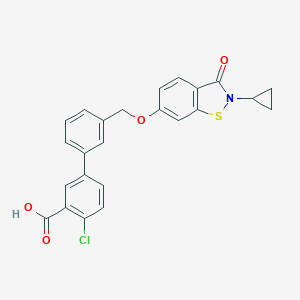 CHEMBL2179629 CHEMBL2179629 | C24H18ClNO4S | 451.921 | 5 / 1 | 5.3 | No |
| 310565 | 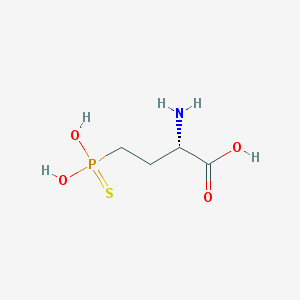 CHEMBL243925 CHEMBL243925 | C4H10NO4PS | 199.161 | 6 / 4 | -3.8 | Yes |
| 310889 | 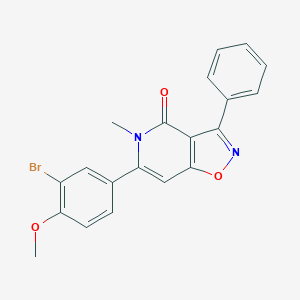 CHEMBL606137 CHEMBL606137 | C20H15BrN2O3 | 411.255 | 4 / 0 | 4.0 | Yes |
| 323916 | 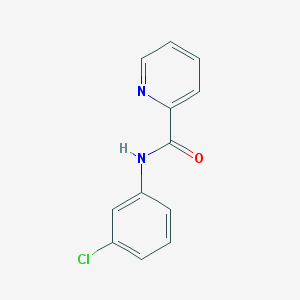 61350-00-3 61350-00-3 | C12H9ClN2O | 232.667 | 2 / 1 | 2.6 | Yes |
| 329964 | 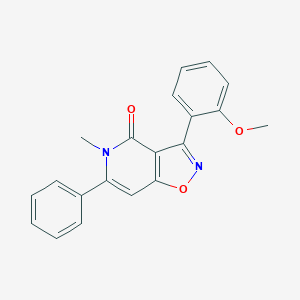 479077-09-3 479077-09-3 | C20H16N2O3 | 332.359 | 4 / 0 | 3.3 | Yes |
| 567704 | 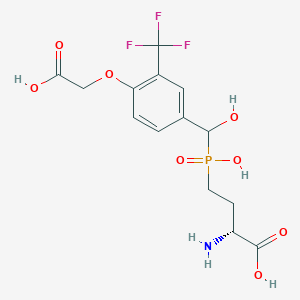 US9212196, Derivative 21 US9212196, Derivative 21 | C14H17F3NO8P | 415.258 | 12 / 5 | -3.0 | No |
| 544956 | 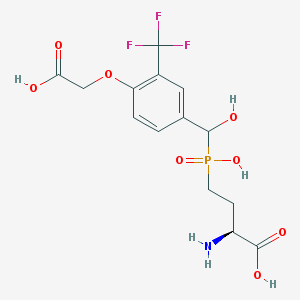 CHEMBL3918221 CHEMBL3918221 | C14H17F3NO8P | 415.258 | 12 / 5 | -3.0 | No |
| 545044 | 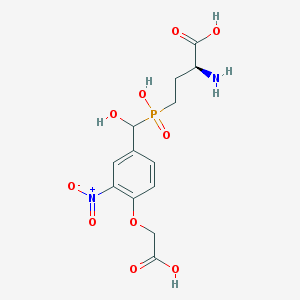 CHEMBL3896848 CHEMBL3896848 | C13H17N2O10P | 392.257 | 11 / 5 | -4.0 | No |
| 567759 | 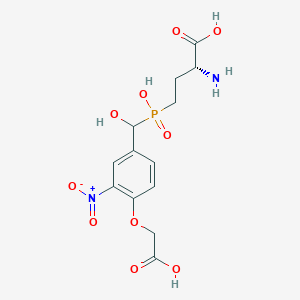 US9212196, Derivative 24 US9212196, Derivative 24 | C13H17N2O10P | 392.257 | 11 / 5 | -4.0 | No |
| 545308 | 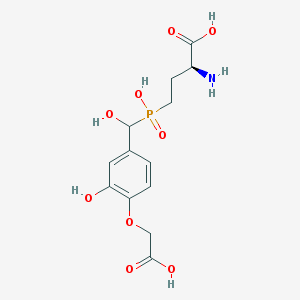 CHEMBL3895668 CHEMBL3895668 | C13H18NO9P | 363.259 | 10 / 6 | -4.2 | No |
| 568057 | 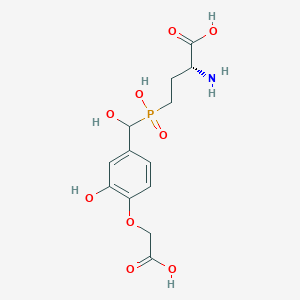 US9212196, Derivative 22 US9212196, Derivative 22 | C13H18NO9P | 363.259 | 10 / 6 | -4.2 | No |
| 348639 | 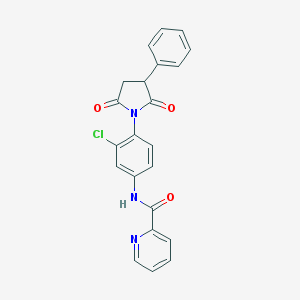 MLS003171622 MLS003171622 | C22H16ClN3O3 | 405.838 | 4 / 1 | 3.1 | Yes |
| 352370 | 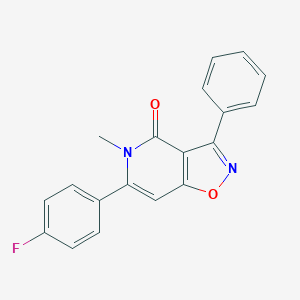 CHEMBL595826 CHEMBL595826 | C19H13FN2O2 | 320.323 | 4 / 0 | 3.4 | Yes |
| 455825 | 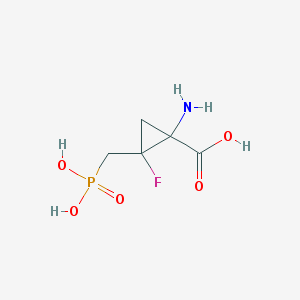 CHEMBL3427499 CHEMBL3427499 | C5H9FNO5P | 213.101 | 7 / 4 | -5.0 | Yes |
| 358355 | 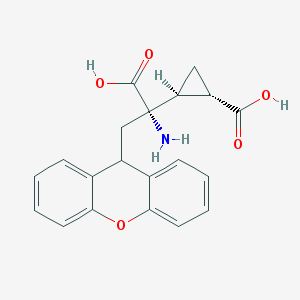 201943-63-7 201943-63-7 | C20H19NO5 | 353.374 | 6 / 3 | -0.2 | Yes |
| 373658 | 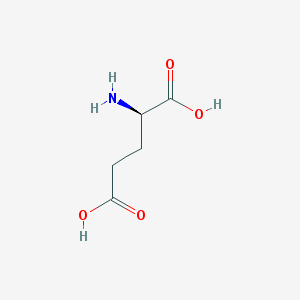 D-glutamic acid D-glutamic acid | C5H9NO4 | 147.13 | 5 / 3 | -3.7 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218