

You can:
| Name | Prostaglandin E2 receptor EP4 subtype |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | PTGER4 |
| Synonym | Prostanoid EP4 receptor PGE2 receptor EP4 subtype PGE receptor EP4 subtype EP4 receptor EP2 |
| Disease | Ulcerative colitis Glaucoma Inflammatory disease Migraine Osteoarthritis [ Show all ] |
| Length | 488 |
| Amino acid sequence | MSTPGVNSSASLSPDRLNSPVTIPAVMFIFGVVGNLVAIVVLCKSRKEQKETTFYTLVCGLAVTDLLGTLLVSPVTIATYMKGQWPGGQPLCEYSTFILLFFSLSGLSIICAMSVERYLAINHAYFYSHYVDKRLAGLTLFAVYASNVLFCALPNMGLGSSRLQYPDTWCFIDWTTNVTAHAAYSYMYAGFSSFLILATVLCNVLVCGALLRMHRQFMRRTSLGTEQHHAAAAASVASRGHPAASPALPRLSDFRRRRSFRRIAGAEIQMVILLIATSLVVLICSIPLVVRVFVNQLYQPSLEREVSKNPDLQAIRIASVNPILDPWIYILLRKTVLSKAIEKIKCLFCRIGGSRRERSGQHCSDSQRTSSAMSGHSRSFISRELKEISSTSQTLLPDLSLPDLSENGLGGRNLLPGVPGMGLAQEDTTSLRTLRISETSDSSQGQDSESVLLVDEAGGSGRAGPAPKGSSLQVTFPSETLNLSEKCI |
| UniProt | P35408 |
| Protein Data Bank | 5ywy, 5yhl |
| GPCR-HGmod model | P35408 |
| 3D structure model | This structure is from PDB ID 5ywy. |
| BioLiP | BL0434347, BL0434289 |
| Therapeutic Target Database | T18876 |
| ChEMBL | CHEMBL1836 |
| IUPHAR | 343 |
| DrugBank | BE0003522 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 431 | 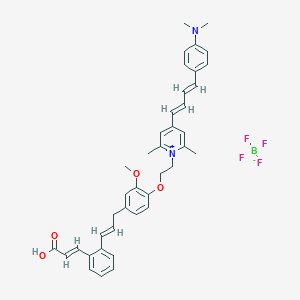 CHEMBL2164612 CHEMBL2164612 | C40H43BF4N2O4 | 702.598 | 10 / 1 | N/A | No |
| 854 | 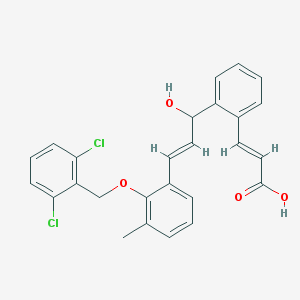 CHEMBL180742 CHEMBL180742 | C26H22Cl2O4 | 469.358 | 4 / 2 | 6.4 | No |
| 535934 | 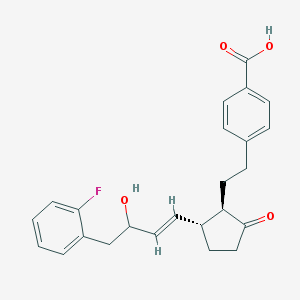 CHEMBL3904946 CHEMBL3904946 | C24H25FO4 | 396.458 | 5 / 2 | 3.9 | Yes |
| 557339 | 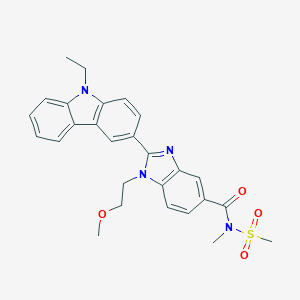 SCHEMBL17938360 SCHEMBL17938360 | C27H28N4O4S | 504.605 | 5 / 0 | 3.6 | No |
| 1597 | 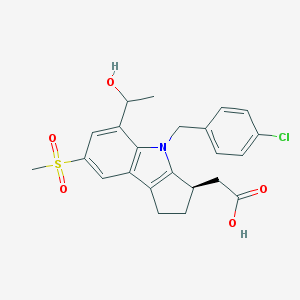 CHEMBL207203 CHEMBL207203 | C23H24ClNO5S | 461.957 | 5 / 2 | 3.0 | Yes |
| 1628 | 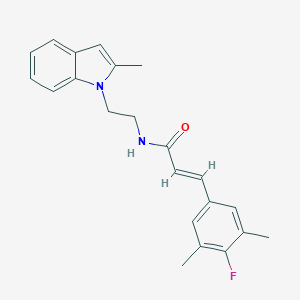 CHEMBL3264203 CHEMBL3264203 | C22H23FN2O | 350.437 | 2 / 1 | 4.6 | Yes |
| 1822 | 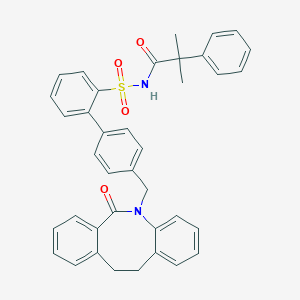 CHEMBL314533 CHEMBL314533 | C38H34N2O4S | 614.76 | 4 / 1 | 7.6 | No |
| 557351 | 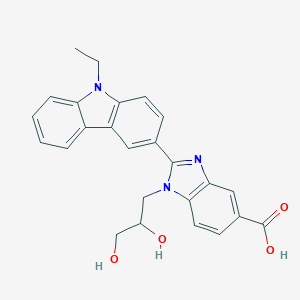 SCHEMBL17944464 SCHEMBL17944464 | C25H23N3O4 | 429.476 | 5 / 3 | 3.0 | Yes |
| 3050 | 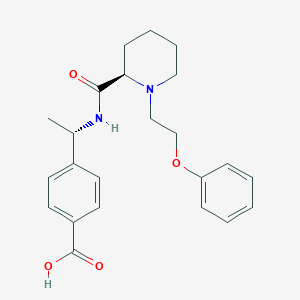 CHEMBL3115074 CHEMBL3115074 | C23H28N2O4 | 396.487 | 5 / 2 | 1.3 | Yes |
| 557409 | 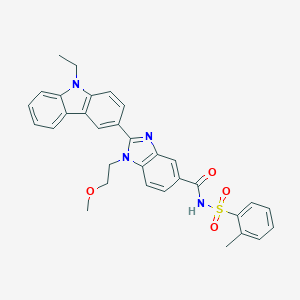 SCHEMBL17938377 SCHEMBL17938377 | C32H30N4O4S | 566.676 | 5 / 1 | 5.4 | No |
| 4546 | 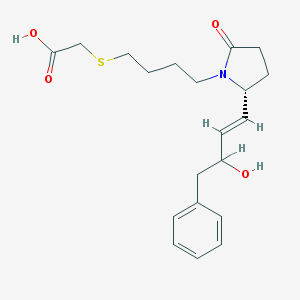 CHEMBL430124 CHEMBL430124 | C20H27NO4S | 377.499 | 5 / 2 | 2.4 | Yes |
| 521601 | 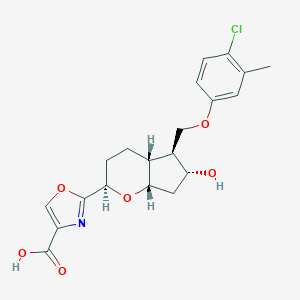 CHEMBL3793009 CHEMBL3793009 | C20H22ClNO6 | 407.847 | 7 / 2 | 3.2 | Yes |
| 517345 | 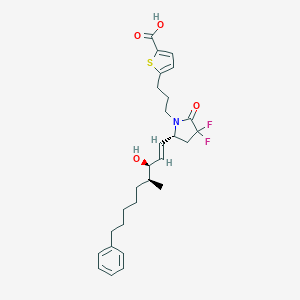 CHEMBL3893847 CHEMBL3893847 | C28H35F2NO4S | 519.648 | 7 / 2 | 6.8 | No |
| 517347 | 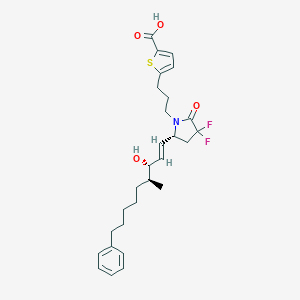 CHEMBL3968443 CHEMBL3968443 | C28H35F2NO4S | 519.648 | 7 / 2 | 6.8 | No |
| 557418 | 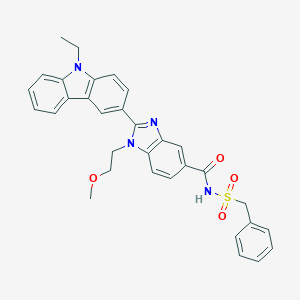 SCHEMBL15774531 SCHEMBL15774531 | C32H30N4O4S | 566.676 | 5 / 1 | 4.9 | No |
| 536081 | 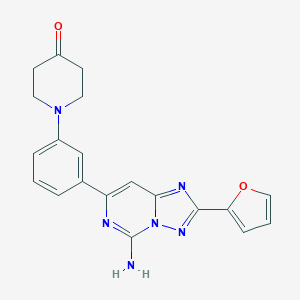 CHEMBL354419 CHEMBL354419 | C20H18N6O2 | 374.404 | 7 / 1 | 1.8 | Yes |
| 6564 | 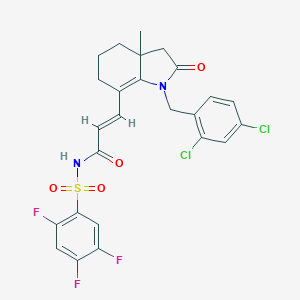 CHEMBL479439 CHEMBL479439 | C25H21Cl2F3N2O4S | 573.408 | 7 / 1 | 4.8 | No |
| 6657 | 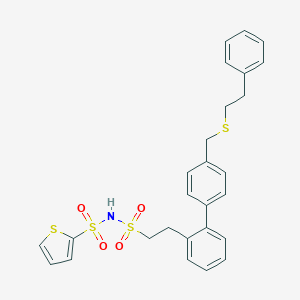 CHEMBL313266 CHEMBL313266 | C27H27NO4S4 | 557.756 | 7 / 1 | 6.5 | No |
| 521662 | 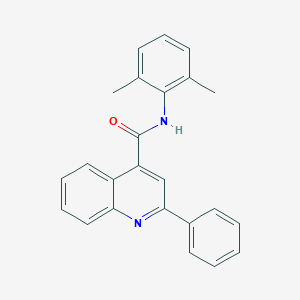 N-(2,6-dimethylphenyl)-2-phenylquinoline-4-carboxamide N-(2,6-dimethylphenyl)-2-phenylquinoline-4-carboxamide | C24H20N2O | 352.437 | 2 / 1 | 5.4 | No |
| 8446 | 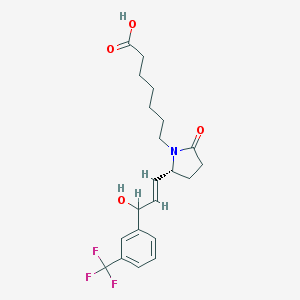 CHEMBL417171 CHEMBL417171 | C21H26F3NO4 | 413.437 | 7 / 2 | 3.1 | Yes |
| 9197 | 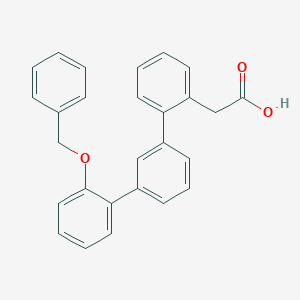 CHEMBL123844 CHEMBL123844 | C27H22O3 | 394.47 | 3 / 1 | 5.3 | No |
| 553296 | 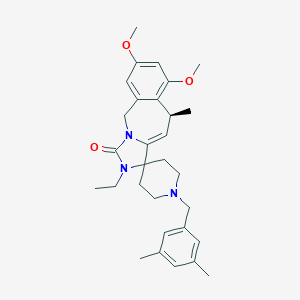 ER 819762 ER 819762 | C30H39N3O3 | 489.66 | 4 / 0 | 4.5 | Yes |
| 557558 | 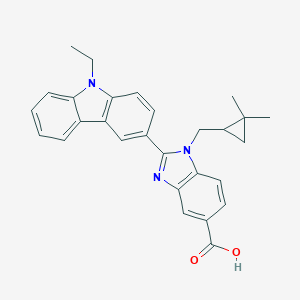 1-[(2,2-Dimethylcyclopropyl)methyl]-2-(9-ethyl-9H-carbazol-3-yl)-1H-benzimidazole-5-carboxylic acid 1-[(2,2-Dimethylcyclopropyl)methyl]-2-(9-ethyl-9H-carbazol-3-yl)-1H-benzimidazole-5-carboxylic acid | C28H27N3O2 | 437.543 | 3 / 1 | 5.9 | No |
| 9788 | 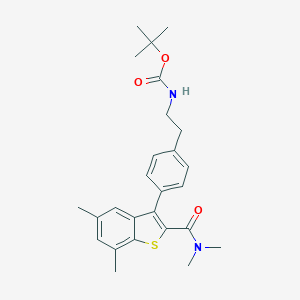 CHEMBL1644015 CHEMBL1644015 | C26H32N2O3S | 452.613 | 4 / 1 | 6.1 | No |
| 10348 | 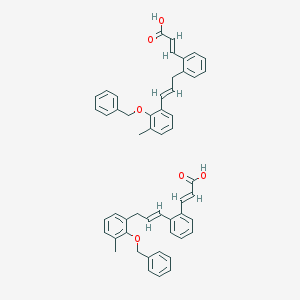 CID 52944193 CID 52944193 | C52H48O6 | 768.95 | 6 / 2 | N/A | No |
| 10673 | 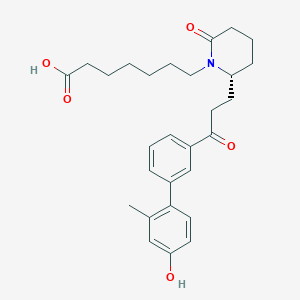 CHEMBL191874 CHEMBL191874 | C28H35NO5 | 465.59 | 5 / 2 | 4.6 | Yes |
| 11637 | 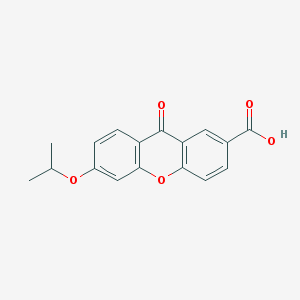 AH 6809 AH 6809 | C17H14O5 | 298.294 | 5 / 1 | 3.9 | Yes |
| 12368 | 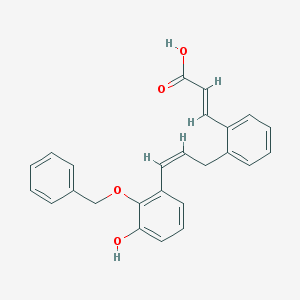 BDBM50370468 BDBM50370468 | C25H22O4 | 386.447 | 4 / 2 | 5.5 | No |
| 553319 | 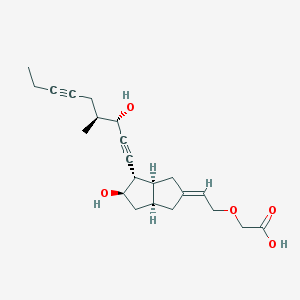 Cicaprost Cicaprost | C22H30O5 | 374.477 | 5 / 3 | 2.1 | Yes |
| 12775 | 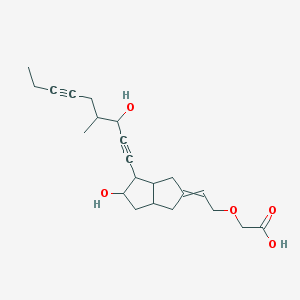 CID 53394056 CID 53394056 | C22H30O5 | 374.477 | 5 / 3 | 2.1 | Yes |
| 557650 | 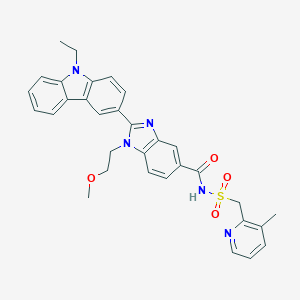 SCHEMBL15774302 SCHEMBL15774302 | C32H31N5O4S | 581.691 | 6 / 1 | 4.3 | No |
| 13142 | 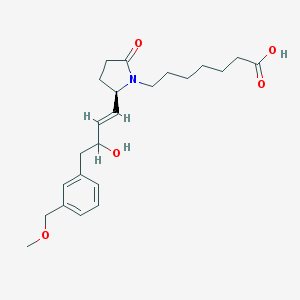 CHEMBL275245 CHEMBL275245 | C23H33NO5 | 403.519 | 5 / 2 | 2.2 | Yes |
| 464463 | 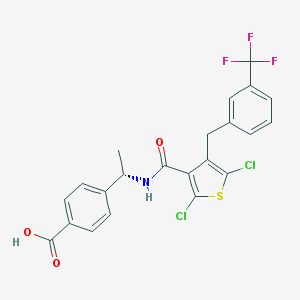 SCHEMBL2350626 SCHEMBL2350626 | C22H16Cl2F3NO3S | 502.329 | 7 / 2 | 7.0 | No |
| 13562 | 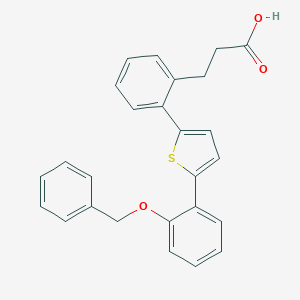 CHEMBL125110 CHEMBL125110 | C26H22O3S | 414.519 | 4 / 1 | 6.1 | No |
| 464548 | 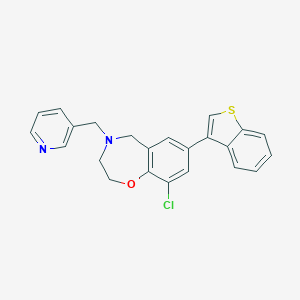 CHEMBL3586353 CHEMBL3586353 | C23H19ClN2OS | 406.928 | 4 / 0 | 5.2 | No |
| 13896 | 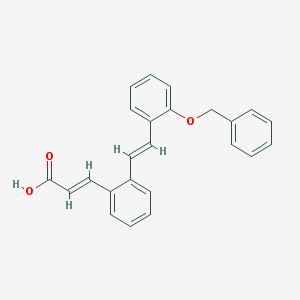 CHEMBL181035 CHEMBL181035 | C24H20O3 | 356.421 | 3 / 1 | 5.6 | No |
| 14340 | 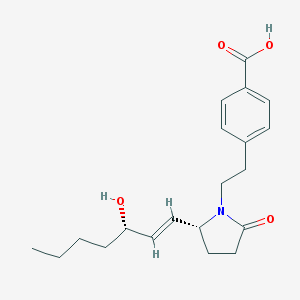 CHEMBL251294 CHEMBL251294 | C20H27NO4 | 345.439 | 4 / 2 | 2.8 | Yes |
| 14356 | 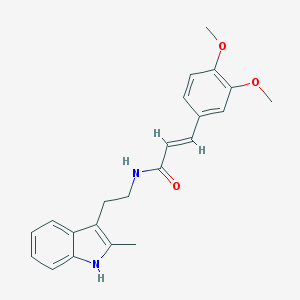 CHEMBL3264208 CHEMBL3264208 | C22H24N2O3 | 364.445 | 3 / 2 | 4.1 | Yes |
| 14515 | 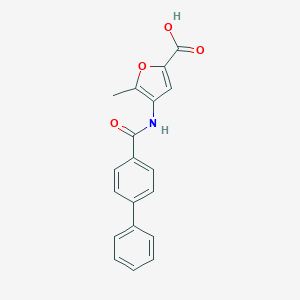 CHEMBL3260433 CHEMBL3260433 | C19H15NO4 | 321.332 | 4 / 2 | 3.7 | Yes |
| 536377 | 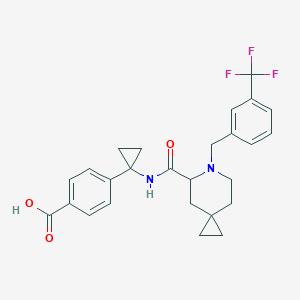 CHEMBL3896466 CHEMBL3896466 | C26H27F3N2O3 | 472.508 | 7 / 2 | 2.6 | Yes |
| 442250 | 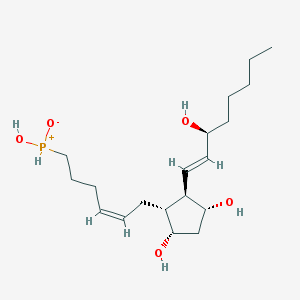 [(5Z,8R,9S,11R,12R,13E,15S)-9,11,15-Trihydroxy-1-norprosta-5,13-diene-2-yl]phosphinic acid [(5Z,8R,9S,11R,12R,13E,15S)-9,11,15-Trihydroxy-1-norprosta-5,13-diene-2-yl]phosphinic acid | C19H35O5P | 374.458 | 5 / 4 | 1.9 | Yes |
| 15305 | 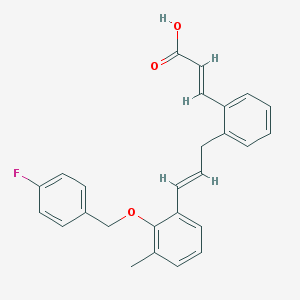 CHEMBL180089 CHEMBL180089 | C26H23FO3 | 402.465 | 4 / 1 | 6.4 | No |
| 16337 | 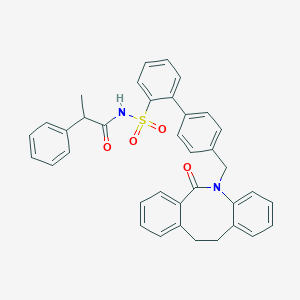 CHEMBL90491 CHEMBL90491 | C37H32N2O4S | 600.733 | 4 / 1 | 7.3 | No |
| 16446 | 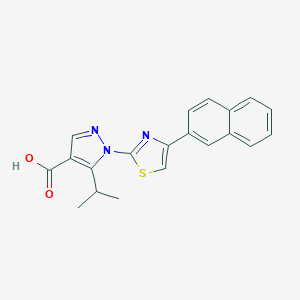 CHEMBL3260420 CHEMBL3260420 | C20H17N3O2S | 363.435 | 5 / 1 | 4.9 | Yes |
| 17052 | 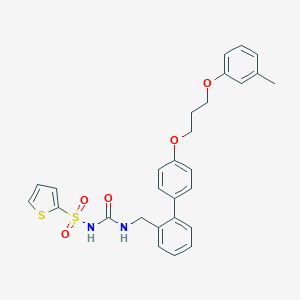 CHEMBL87797 CHEMBL87797 | C28H28N2O5S2 | 536.661 | 6 / 2 | 6.0 | No |
| 17136 | 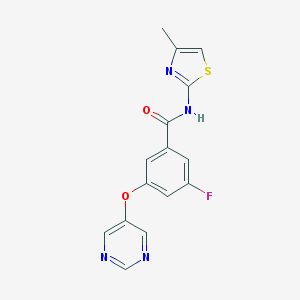 CHEMBL2440659 CHEMBL2440659 | C15H11FN4O2S | 330.337 | 7 / 1 | 2.4 | Yes |
| 459366 | 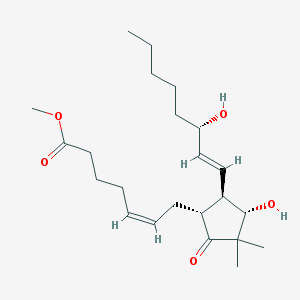 SCHEMBL2195516 SCHEMBL2195516 | C23H38O5 | 394.552 | 5 / 2 | 4.1 | Yes |
| 17940 | 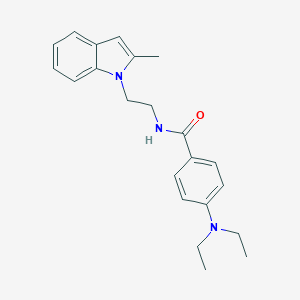 CHEMBL3286635 CHEMBL3286635 | C22H27N3O | 349.478 | 2 / 1 | 4.2 | Yes |
| 459373 | 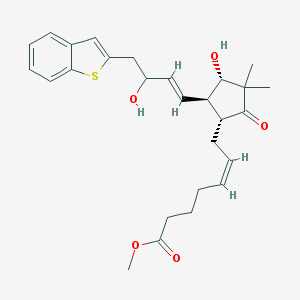 SCHEMBL2191405 SCHEMBL2191405 | C27H34O5S | 470.624 | 6 / 2 | 4.4 | Yes |
| 557807 | 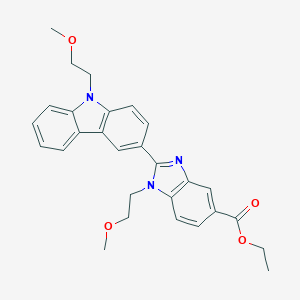 SCHEMBL17938384 SCHEMBL17938384 | C28H29N3O4 | 471.557 | 5 / 0 | 4.3 | Yes |
| 18202 | 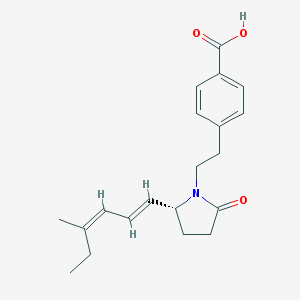 CHEMBL251710 CHEMBL251710 | C20H25NO3 | 327.424 | 3 / 1 | 3.9 | Yes |
| 536455 | 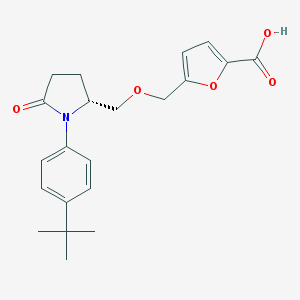 SCHEMBL1310801 SCHEMBL1310801 | C21H25NO5 | 371.433 | 5 / 1 | 3.4 | Yes |
| 557886 | 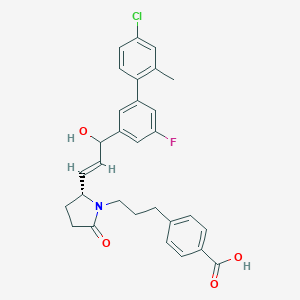 SCHEMBL18357252 SCHEMBL18357252 | C30H29ClFNO4 | 522.013 | 5 / 2 | 5.7 | No |
| 517426 | 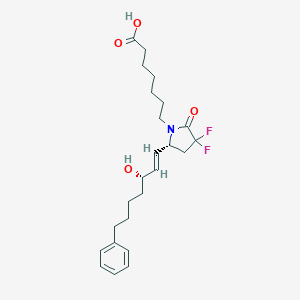 CHEMBL3959605 CHEMBL3959605 | C24H33F2NO4 | 437.528 | 6 / 2 | 4.5 | Yes |
| 20057 | 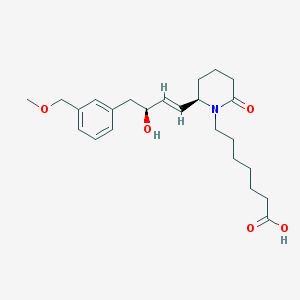 CHEMBL413563 CHEMBL413563 | C24H35NO5 | 417.546 | 5 / 2 | 2.6 | Yes |
| 536485 | 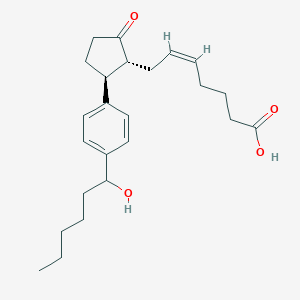 SCHEMBL2011904 SCHEMBL2011904 | C24H34O4 | 386.532 | 4 / 2 | 4.4 | Yes |
| 20284 | 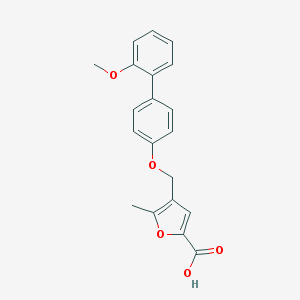 CHEMBL3260439 CHEMBL3260439 | C20H18O5 | 338.359 | 5 / 1 | 4.3 | Yes |
| 517430 | 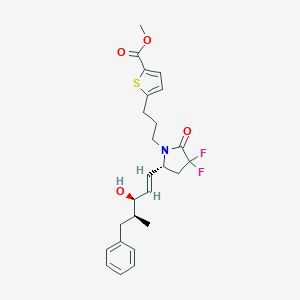 CHEMBL3978072 CHEMBL3978072 | C25H29F2NO4S | 477.567 | 7 / 1 | 5.2 | No |
| 517431 | 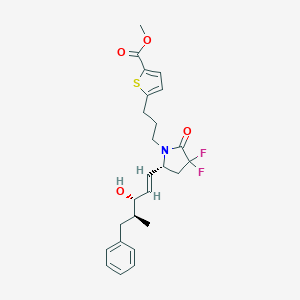 CHEMBL3911081 CHEMBL3911081 | C25H29F2NO4S | 477.567 | 7 / 1 | 5.2 | No |
| 536535 | 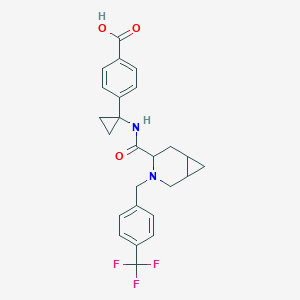 CHEMBL3953493 CHEMBL3953493 | C25H25F3N2O3 | 458.481 | 7 / 2 | 1.9 | Yes |
| 536539 | 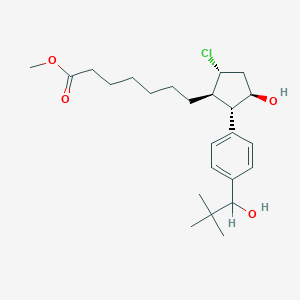 SCHEMBL5203473 SCHEMBL5203473 | C24H37ClO4 | 425.006 | 4 / 2 | 5.4 | No |
| 22146 | 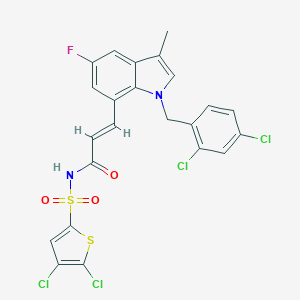 DG-041 DG-041 | C23H15Cl4FN2O3S2 | 592.302 | 5 / 1 | 7.8 | No |
| 522154 | 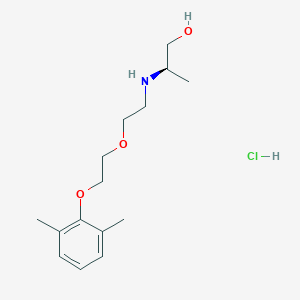 CHEMBL3775436 CHEMBL3775436 | C15H26ClNO3 | 303.827 | 4 / 3 | N/A | N/A |
| 522200 | 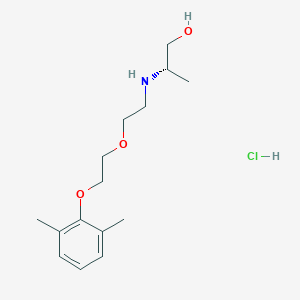 CHEMBL3775769 CHEMBL3775769 | C15H26ClNO3 | 303.827 | 4 / 3 | N/A | N/A |
| 536544 | 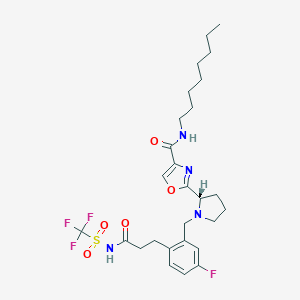 CHEMBL3896035 CHEMBL3896035 | C27H36F4N4O5S | 604.662 | 11 / 2 | 5.6 | No |
| 22800 | 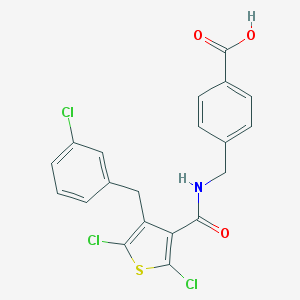 CHEMBL601299 CHEMBL601299 | C20H14Cl3NO3S | 454.746 | 4 / 2 | 6.4 | No |
| 23036 | 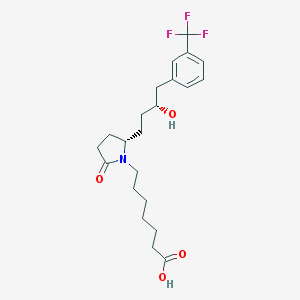 CHEMBL46395 CHEMBL46395 | C22H30F3NO4 | 429.48 | 7 / 2 | 3.9 | Yes |
| 553365 | 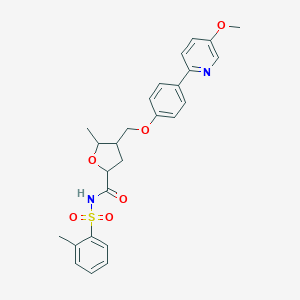 EP4 antagonist, BTG EP4 antagonist, BTG | C26H28N2O6S | 496.578 | 7 / 1 | 4.0 | Yes |
| 459419 | 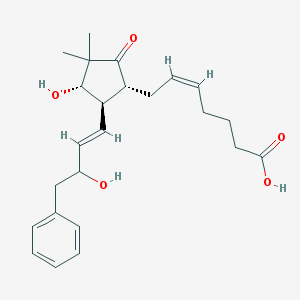 SCHEMBL2194130 SCHEMBL2194130 | C24H32O5 | 400.515 | 5 / 3 | 3.0 | Yes |
| 553379 | 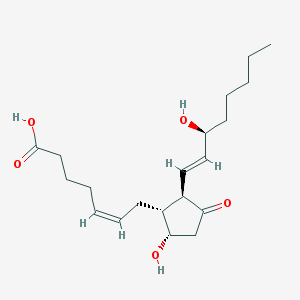 prostaglandin D2 prostaglandin D2 | C20H32O5 | 352.471 | 5 / 3 | 2.6 | Yes |
| 24172 | 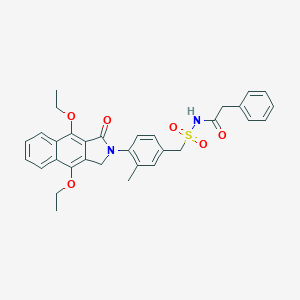 CHEMBL255527 CHEMBL255527 | C32H32N2O6S | 572.676 | 6 / 1 | 5.1 | No |
| 24228 | 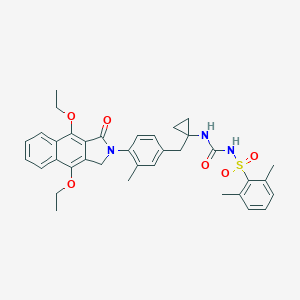 CHEMBL1669026 CHEMBL1669026 | C36H39N3O6S | 641.783 | 6 / 2 | 7.0 | No |
| 442611 | 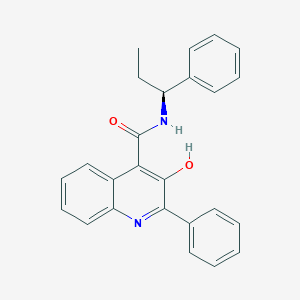 Talnetant Talnetant | C25H22N2O2 | 382.463 | 3 / 2 | 5.7 | No |
| 465992 | 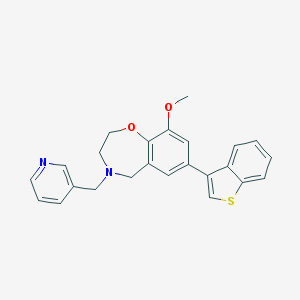 CHEMBL3586309 CHEMBL3586309 | C24H22N2O2S | 402.512 | 5 / 0 | 4.5 | Yes |
| 536642 | 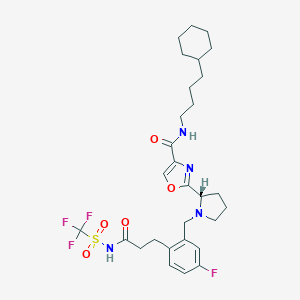 SCHEMBL671511 SCHEMBL671511 | C29H38F4N4O5S | 630.7 | 11 / 2 | 6.2 | No |
| 536662 | 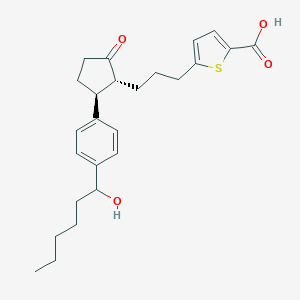 SCHEMBL2014800 SCHEMBL2014800 | C25H32O4S | 428.587 | 5 / 2 | 5.5 | No |
| 536671 | 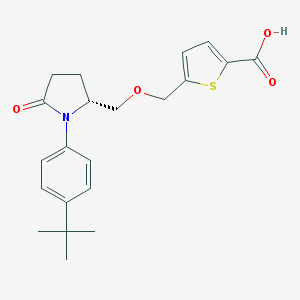 SCHEMBL1309617 SCHEMBL1309617 | C21H25NO4S | 387.494 | 5 / 1 | 4.0 | Yes |
| 26669 | 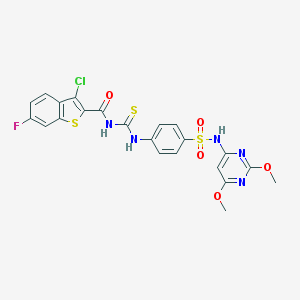 MLS000553986 MLS000553986 | C22H17ClFN5O5S3 | 582.036 | 11 / 3 | 5.3 | No |
| 26696 | 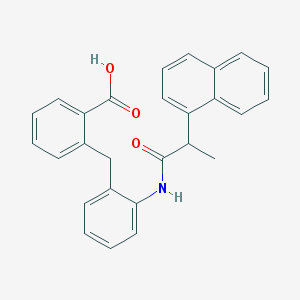 ONO-AE2-227 ONO-AE2-227 | C27H23NO3 | 409.485 | 3 / 2 | 6.0 | No |
| 27562 | 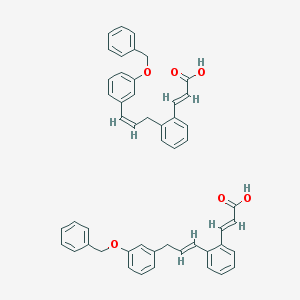 CID 52945421 CID 52945421 | C50H44O6 | 740.896 | 6 / 2 | N/A | No |
| 558120 | 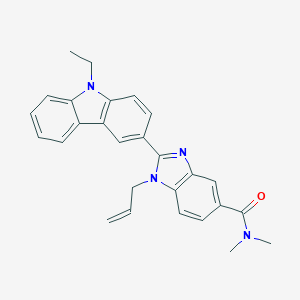 SCHEMBL15774525 SCHEMBL15774525 | C27H26N4O | 422.532 | 2 / 0 | 5.0 | Yes |
| 28502 | 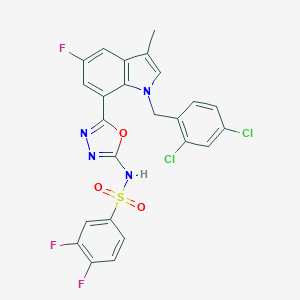 CHEMBL1080281 CHEMBL1080281 | C24H15Cl2F3N4O3S | 567.364 | 9 / 1 | 5.9 | No |
| 548199 | 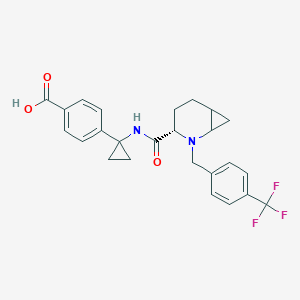 CHEMBL3938316 CHEMBL3938316 | C25H25F3N2O3 | 458.481 | 7 / 2 | 2.1 | Yes |
| 558124 | 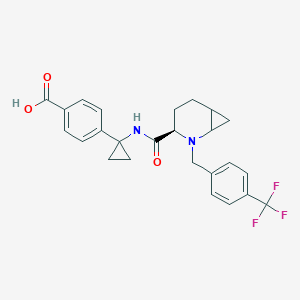 SCHEMBL16584716 SCHEMBL16584716 | C25H25F3N2O3 | 458.481 | 7 / 2 | 2.1 | Yes |
| 28688 | 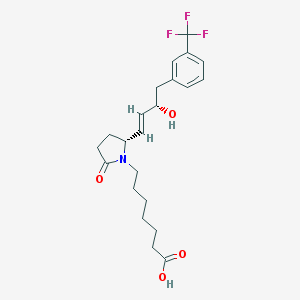 CHEMBL45008 CHEMBL45008 | C22H28F3NO4 | 427.464 | 7 / 2 | 3.4 | Yes |
| 28693 | 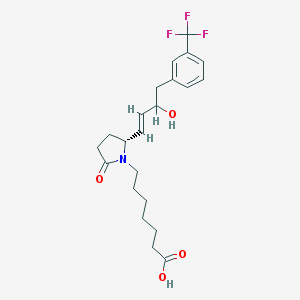 CHEMBL440474 CHEMBL440474 | C22H28F3NO4 | 427.464 | 7 / 2 | 3.4 | Yes |
| 522421 | 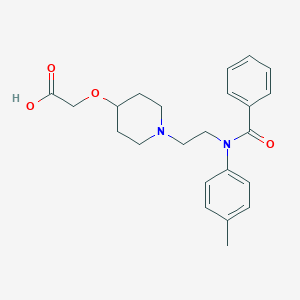 CHEMBL3793911 CHEMBL3793911 | C23H28N2O4 | 396.487 | 5 / 1 | 1.1 | Yes |
| 522434 | 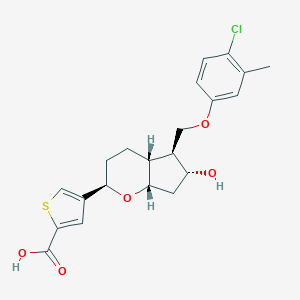 CHEMBL3794185 CHEMBL3794185 | C21H23ClO5S | 422.92 | 6 / 2 | 4.4 | Yes |
| 29712 | 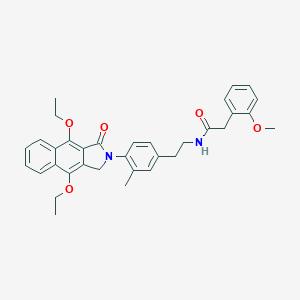 CHEMBL1669023 CHEMBL1669023 | C34H36N2O5 | 552.671 | 5 / 1 | 5.9 | No |
| 30173 | 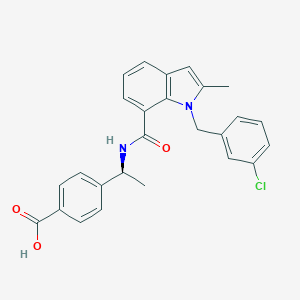 CHEMBL1084047 CHEMBL1084047 | C26H23ClN2O3 | 446.931 | 3 / 2 | 5.3 | No |
| 30440 | 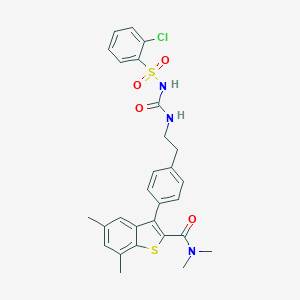 CHEMBL1644005 CHEMBL1644005 | C28H28ClN3O4S2 | 570.119 | 5 / 2 | 6.2 | No |
| 522479 | 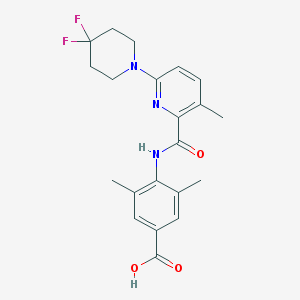 CHEMBL3793002 CHEMBL3793002 | C21H23F2N3O3 | 403.43 | 7 / 2 | 4.3 | Yes |
| 30899 | 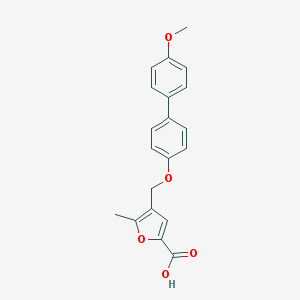 CHEMBL3260437 CHEMBL3260437 | C20H18O5 | 338.359 | 5 / 1 | 4.3 | Yes |
| 31004 | 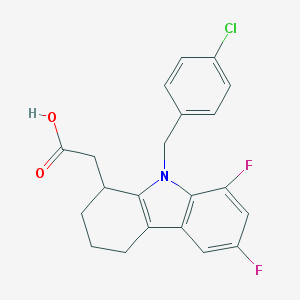 CHEMBL208260 CHEMBL208260 | C21H18ClF2NO2 | 389.827 | 4 / 1 | 4.9 | Yes |
| 31569 | 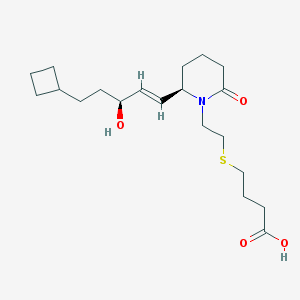 CHEMBL372894 CHEMBL372894 | C20H33NO4S | 383.547 | 5 / 2 | 2.7 | Yes |
| 548246 | 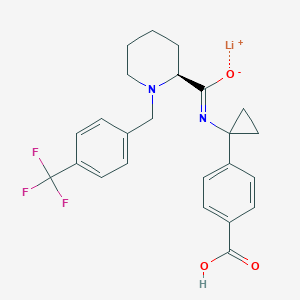 CHEMBL3964180 CHEMBL3964180 | C24H24F3LiN2O3 | 452.402 | 8 / 1 | N/A | N/A |
| 558248 | 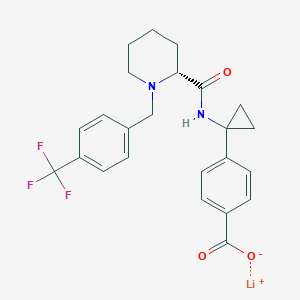 SCHEMBL16584587 SCHEMBL16584587 | C24H24F3LiN2O3 | 452.402 | 7 / 1 | N/A | N/A |
| 459502 | 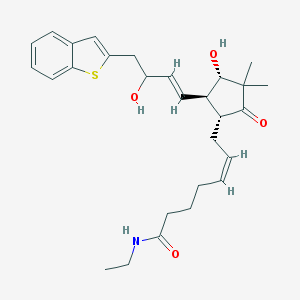 SCHEMBL2190873 SCHEMBL2190873 | C28H37NO4S | 483.667 | 5 / 3 | 4.2 | Yes |
| 32541 | 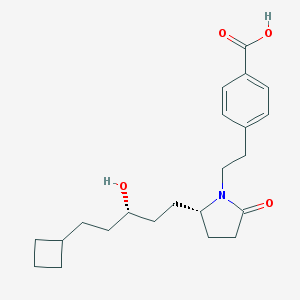 CHEMBL223744 CHEMBL223744 | C22H31NO4 | 373.493 | 4 / 2 | 3.8 | Yes |
| 32687 | 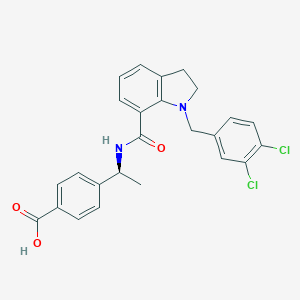 CHEMBL1645134 CHEMBL1645134 | C25H22Cl2N2O3 | 469.362 | 4 / 2 | 5.5 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218