

You can:
| Name | Prostacyclin receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | PTGIR |
| Synonym | prostanoid IP receptor prostaglandin I2 receptor prostaglandin I2 (prostacyclin) receptor (IP) prostacyclin receptor PGI2 receptor [ Show all ] |
| Disease | Solid tumours Pulmonary hypertension Medical abortion Pulmonary arterial hypertension Hypertension [ Show all ] |
| Length | 386 |
| Amino acid sequence | MADSCRNLTYVRGSVGPATSTLMFVAGVVGNGLALGILSARRPARPSAFAVLVTGLAATDLLGTSFLSPAVFVAYARNSSLLGLARGGPALCDAFAFAMTFFGLASMLILFAMAVERCLALSHPYLYAQLDGPRCARLALPAIYAFCVLFCALPLLGLGQHQQYCPGSWCFLRMRWAQPGGAAFSLAYAGLVALLVAAIFLCNGSVTLSLCRMYRQQKRHQGSLGPRPRTGEDEVDHLILLALMTVVMAVCSLPLTIRCFTQAVAPDSSSEMGDLLAFRFYAFNPILDPWVFILFRKAVFQRLKLWVCCLCLGPAHGDSQTPLSQLASGRRDPRAPSAPVGKEGSCVPLSAWGEGQVEPLPPTQQSSGSAVGTSSKAEASVACSLC |
| UniProt | P43119 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P43119 |
| 3D structure model | This predicted structure model is from GPCR-EXP P43119. |
| BioLiP | N/A |
| Therapeutic Target Database | T99954 |
| ChEMBL | CHEMBL1995 |
| IUPHAR | 345 |
| DrugBank | BE0000475 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 831 | 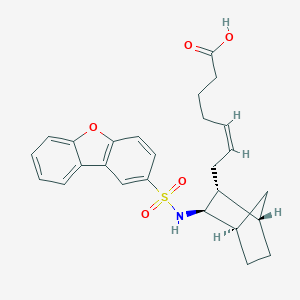 CHEMBL309835 CHEMBL309835 | C26H29NO5S | 467.58 | 6 / 2 | 5.4 | No |
| 1594 | 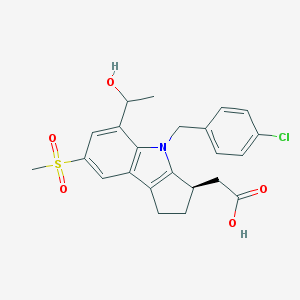 CHEMBL207203 CHEMBL207203 | C23H24ClNO5S | 461.957 | 5 / 2 | 3.0 | Yes |
| 1629 | 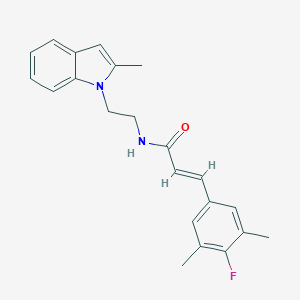 CHEMBL3264203 CHEMBL3264203 | C22H23FN2O | 350.437 | 2 / 1 | 4.6 | Yes |
| 1820 | 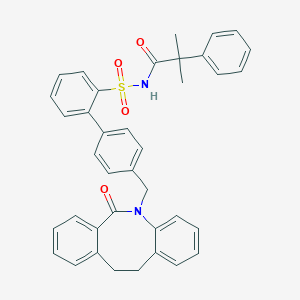 CHEMBL314533 CHEMBL314533 | C38H34N2O4S | 614.76 | 4 / 1 | 7.6 | No |
| 3287 | 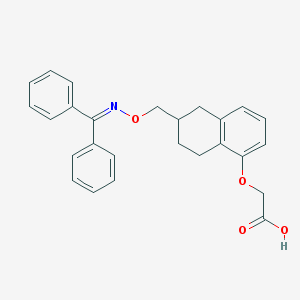 CHEMBL417069 CHEMBL417069 | C26H25NO4 | 415.489 | 5 / 1 | 6.0 | No |
| 4549 | 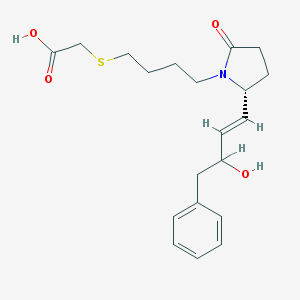 CHEMBL430124 CHEMBL430124 | C20H27NO4S | 377.499 | 5 / 2 | 2.4 | Yes |
| 521596 | 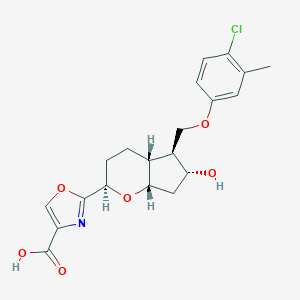 CHEMBL3793009 CHEMBL3793009 | C20H22ClNO6 | 407.847 | 7 / 2 | 3.2 | Yes |
| 521622 | 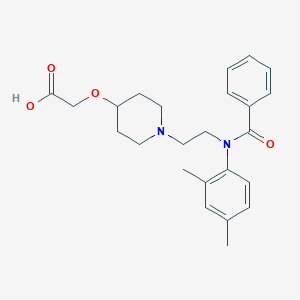 CHEMBL3793957 CHEMBL3793957 | C24H30N2O4 | 410.514 | 5 / 1 | 1.4 | Yes |
| 536083 | 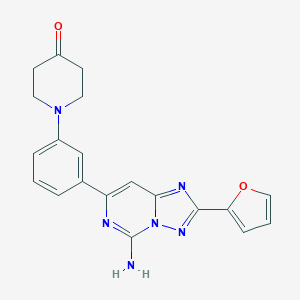 CHEMBL354419 CHEMBL354419 | C20H18N6O2 | 374.404 | 7 / 1 | 1.8 | Yes |
| 9392 | 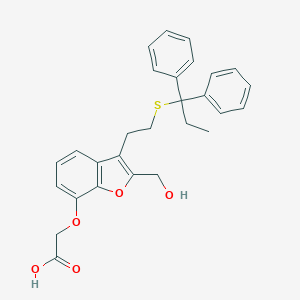 CHEMBL189694 CHEMBL189694 | C28H28O5S | 476.587 | 6 / 2 | 5.9 | No |
| 11638 | 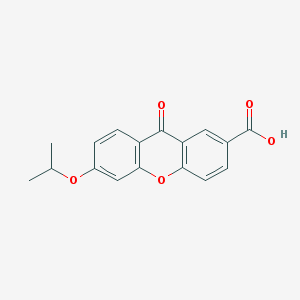 AH 6809 AH 6809 | C17H14O5 | 298.294 | 5 / 1 | 3.9 | Yes |
| 553318 | 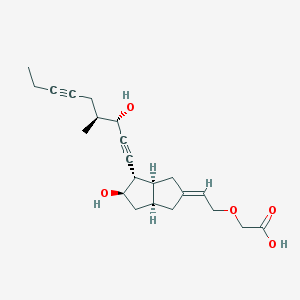 Cicaprost Cicaprost | C22H30O5 | 374.477 | 5 / 3 | 2.1 | Yes |
| 12779 | 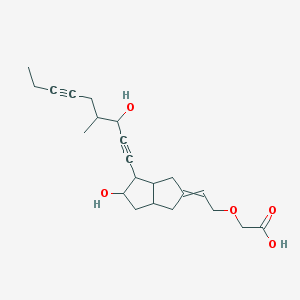 CID 53394056 CID 53394056 | C22H30O5 | 374.477 | 5 / 3 | 2.1 | Yes |
| 13140 | 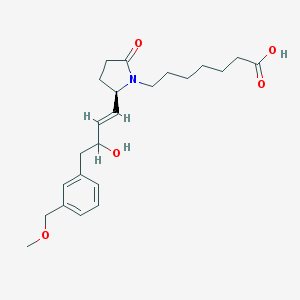 CHEMBL275245 CHEMBL275245 | C23H33NO5 | 403.519 | 5 / 2 | 2.2 | Yes |
| 13889 | 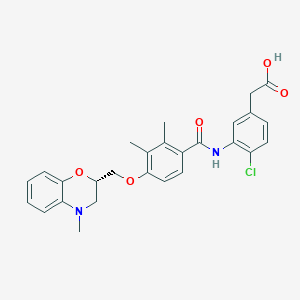 CHEMBL1915676 CHEMBL1915676 | C27H27ClN2O5 | 494.972 | 6 / 2 | 5.2 | No |
| 14357 | 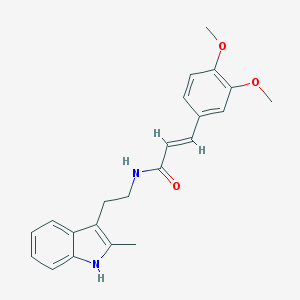 CHEMBL3264208 CHEMBL3264208 | C22H24N2O3 | 364.445 | 3 / 2 | 4.1 | Yes |
| 442249 | 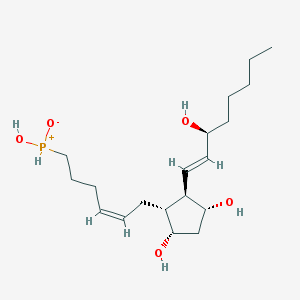 [(5Z,8R,9S,11R,12R,13E,15S)-9,11,15-Trihydroxy-1-norprosta-5,13-diene-2-yl]phosphinic acid [(5Z,8R,9S,11R,12R,13E,15S)-9,11,15-Trihydroxy-1-norprosta-5,13-diene-2-yl]phosphinic acid | C19H35O5P | 374.458 | 5 / 4 | 1.9 | Yes |
| 15993 | 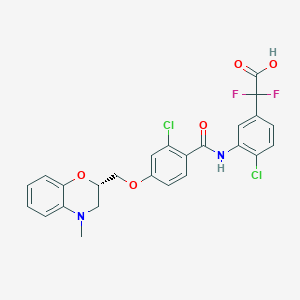 CHEMBL1915866 CHEMBL1915866 | C25H20Cl2F2N2O5 | 537.341 | 8 / 2 | 5.8 | No |
| 16329 | 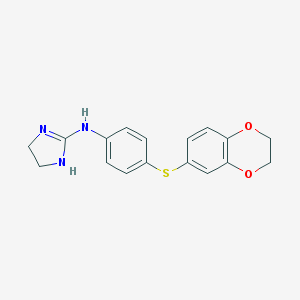 CHEMBL9649 CHEMBL9649 | C17H17N3O2S | 327.402 | 4 / 2 | 2.4 | Yes |
| 16330 | 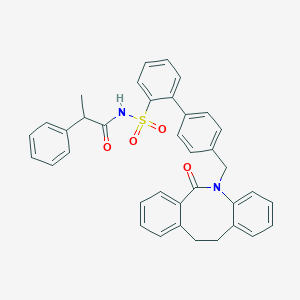 CHEMBL90491 CHEMBL90491 | C37H32N2O4S | 600.733 | 4 / 1 | 7.3 | No |
| 17937 | 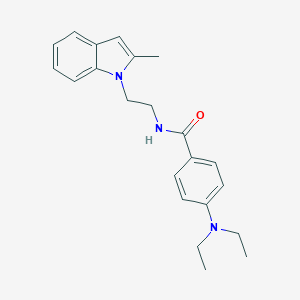 CHEMBL3286635 CHEMBL3286635 | C22H27N3O | 349.478 | 2 / 1 | 4.2 | Yes |
| 18388 | 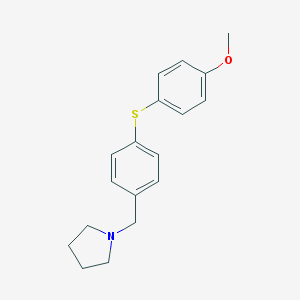 CHEMBL9367 CHEMBL9367 | C18H21NOS | 299.432 | 3 / 0 | 4.3 | Yes |
| 18599 | 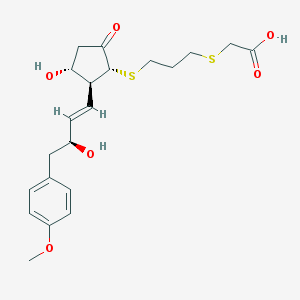 CHEMBL305126 CHEMBL305126 | C21H28O6S2 | 440.569 | 8 / 3 | 2.3 | Yes |
| 442474 | 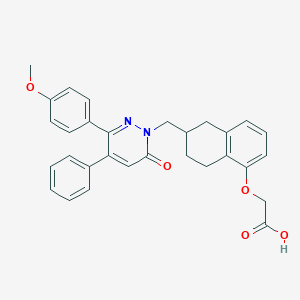 CHEMBL3398215 CHEMBL3398215 | C30H28N2O5 | 496.563 | 6 / 1 | 5.2 | No |
| 21214 | 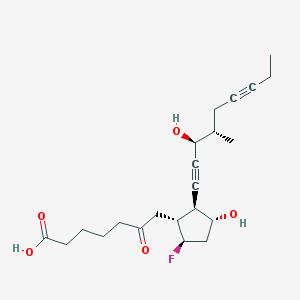 CHEMBL138520 CHEMBL138520 | C22H31FO5 | 394.483 | 6 / 3 | 2.2 | Yes |
| 442512 | 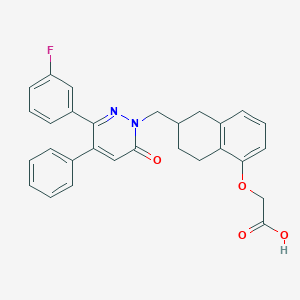 CHEMBL3398214 CHEMBL3398214 | C29H25FN2O4 | 484.527 | 6 / 1 | 5.3 | No |
| 21697 | 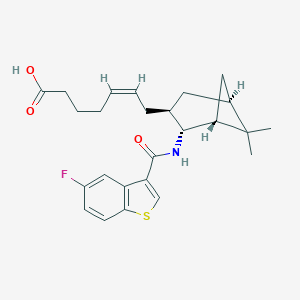 CHEMBL116836 CHEMBL116836 | C25H30FNO3S | 443.577 | 5 / 2 | 5.7 | No |
| 22145 | 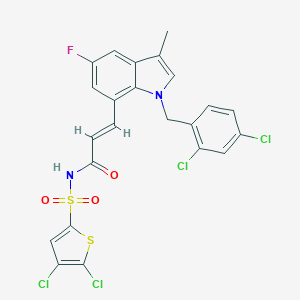 DG-041 DG-041 | C23H15Cl4FN2O3S2 | 592.302 | 5 / 1 | 7.8 | No |
| 522127 | 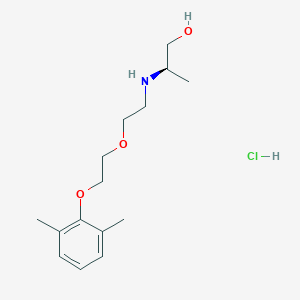 CHEMBL3775436 CHEMBL3775436 | C15H26ClNO3 | 303.827 | 4 / 3 | N/A | N/A |
| 522173 | 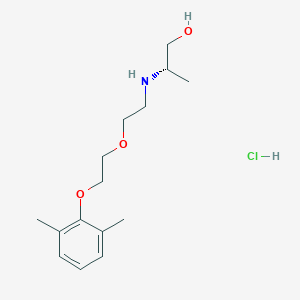 CHEMBL3775769 CHEMBL3775769 | C15H26ClNO3 | 303.827 | 4 / 3 | N/A | N/A |
| 22195 | 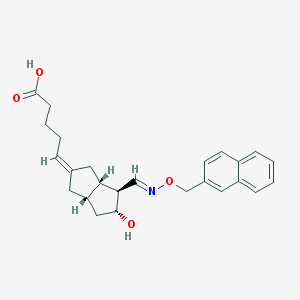 CHEMBL62977 CHEMBL62977 | C25H29NO4 | 407.51 | 5 / 2 | 3.8 | Yes |
| 22197 | 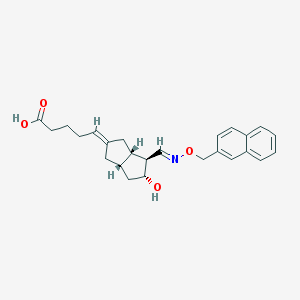 CHEMBL60613 CHEMBL60613 | C25H29NO4 | 407.51 | 5 / 2 | 3.8 | Yes |
| 536550 | 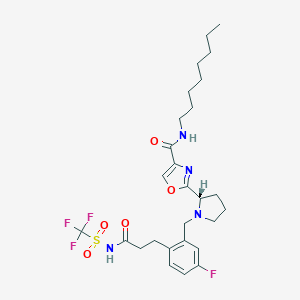 CHEMBL3896035 CHEMBL3896035 | C27H36F4N4O5S | 604.662 | 11 / 2 | 5.6 | No |
| 22692 | 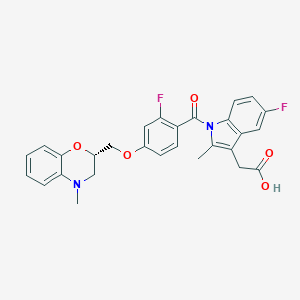 CHEMBL1813277 CHEMBL1813277 | C28H24F2N2O5 | 506.506 | 8 / 1 | 5.2 | No |
| 22807 | 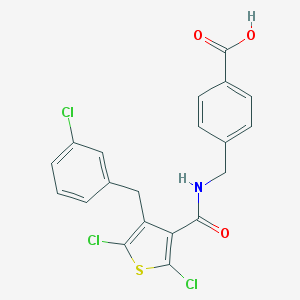 CHEMBL601299 CHEMBL601299 | C20H14Cl3NO3S | 454.746 | 4 / 2 | 6.4 | No |
| 536646 | 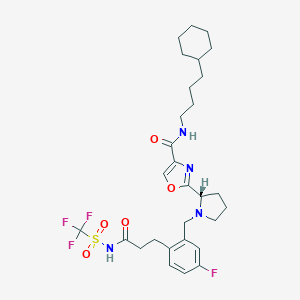 SCHEMBL671511 SCHEMBL671511 | C29H38F4N4O5S | 630.7 | 11 / 2 | 6.2 | No |
| 25481 | 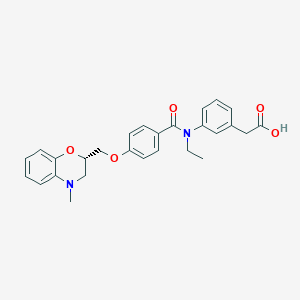 CHEMBL1819607 CHEMBL1819607 | C27H28N2O5 | 460.53 | 6 / 1 | 4.4 | Yes |
| 26668 | 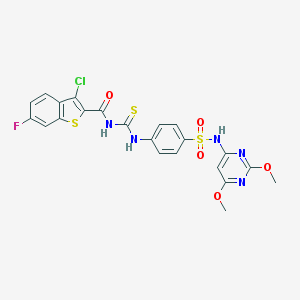 MLS000553986 MLS000553986 | C22H17ClFN5O5S3 | 582.036 | 11 / 3 | 5.3 | No |
| 26683 | 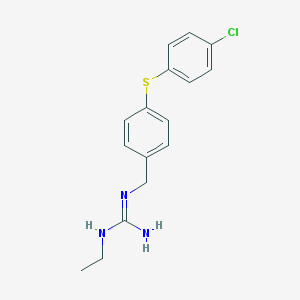 CHEMBL10074 CHEMBL10074 | C16H18ClN3S | 319.851 | 2 / 2 | 3.8 | Yes |
| 27216 | 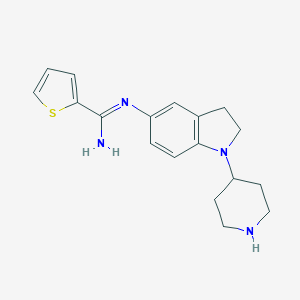 CHEMBL2203713 CHEMBL2203713 | C18H22N4S | 326.462 | 4 / 2 | 2.9 | Yes |
| 522423 | 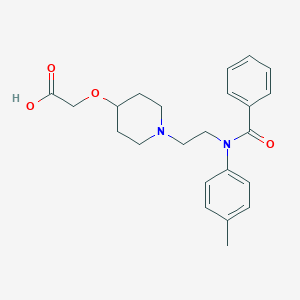 CHEMBL3793911 CHEMBL3793911 | C23H28N2O4 | 396.487 | 5 / 1 | 1.1 | Yes |
| 522433 | 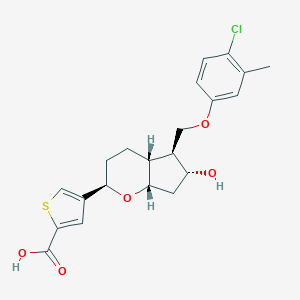 CHEMBL3794185 CHEMBL3794185 | C21H23ClO5S | 422.92 | 6 / 2 | 4.4 | Yes |
| 553406 | 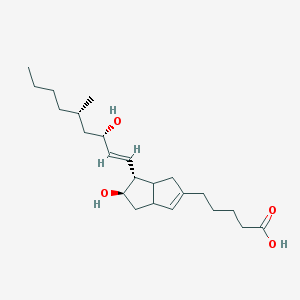 Tei 9063 Tei 9063 | C23H38O4 | 378.553 | 4 / 3 | 4.4 | Yes |
| 31001 | 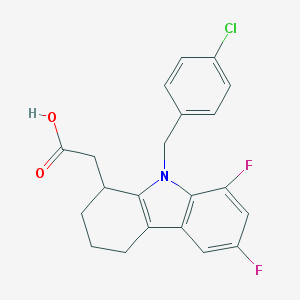 CHEMBL208260 CHEMBL208260 | C21H18ClF2NO2 | 389.827 | 4 / 1 | 4.9 | Yes |
| 32351 | 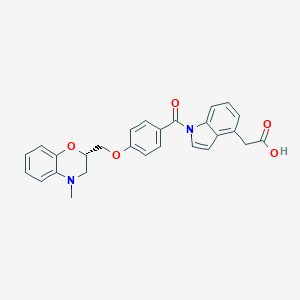 CHEMBL1915669 CHEMBL1915669 | C27H24N2O5 | 456.498 | 6 / 1 | 4.6 | Yes |
| 32615 | 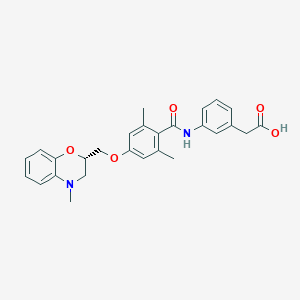 CHEMBL1915672 CHEMBL1915672 | C27H28N2O5 | 460.53 | 6 / 2 | 4.5 | Yes |
| 34236 | 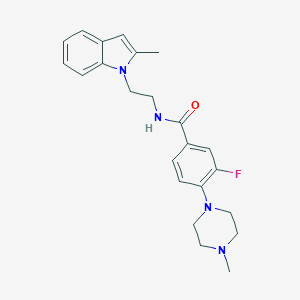 CHEMBL3286655 CHEMBL3286655 | C23H27FN4O | 394.494 | 4 / 1 | 3.3 | Yes |
| 34825 | 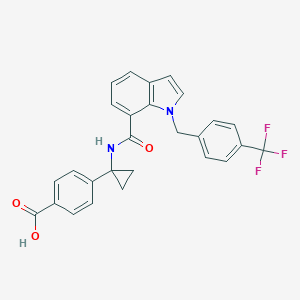 UNII-86KF5VSV88 UNII-86KF5VSV88 | C27H21F3N2O3 | 478.471 | 6 / 2 | 5.1 | No |
| 35050 | 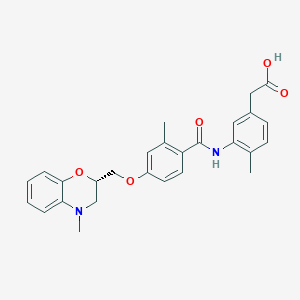 CHEMBL1819611 CHEMBL1819611 | C27H28N2O5 | 460.53 | 6 / 2 | 4.5 | Yes |
| 35476 | 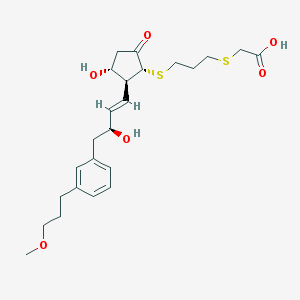 CHEMBL303532 CHEMBL303532 | C24H34O6S2 | 482.65 | 8 / 3 | 2.8 | Yes |
| 35562 | 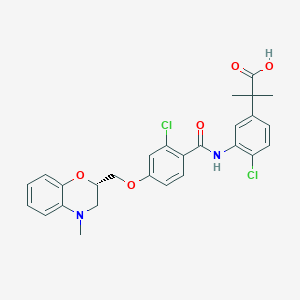 CHEMBL1915861 CHEMBL1915861 | C27H26Cl2N2O5 | 529.414 | 6 / 2 | 6.0 | No |
| 443151 | 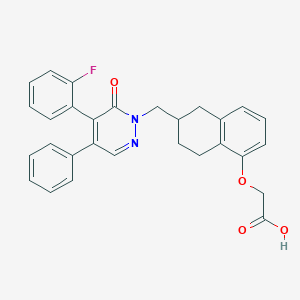 CHEMBL3398230 CHEMBL3398230 | C29H25FN2O4 | 484.527 | 6 / 1 | 5.0 | Yes |
| 38277 | 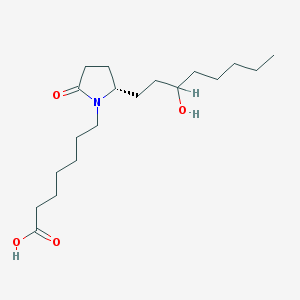 CHEMBL428524 CHEMBL428524 | C19H35NO4 | 341.492 | 4 / 2 | 3.4 | Yes |
| 38280 | 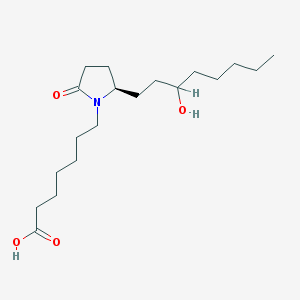 CHEMBL14286 CHEMBL14286 | C19H35NO4 | 341.492 | 4 / 2 | 3.4 | Yes |
| 38293 | 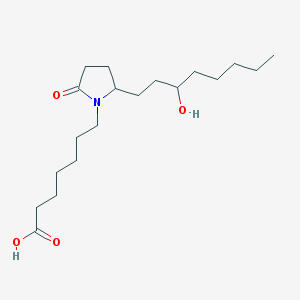 64054-40-6 64054-40-6 | C19H35NO4 | 341.492 | 4 / 2 | 3.4 | Yes |
| 38591 | 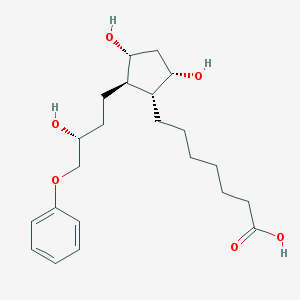 CHEMBL425681 CHEMBL425681 | C22H34O6 | 394.508 | 6 / 4 | 3.2 | Yes |
| 39098 | 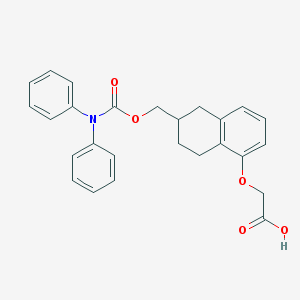 CHEMBL197319 CHEMBL197319 | C26H25NO5 | 431.488 | 5 / 1 | 5.4 | No |
| 40034 | 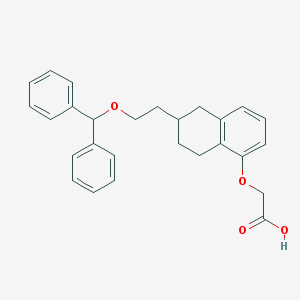 CHEMBL274393 CHEMBL274393 | C27H28O4 | 416.517 | 4 / 1 | 6.0 | No |
| 41240 | 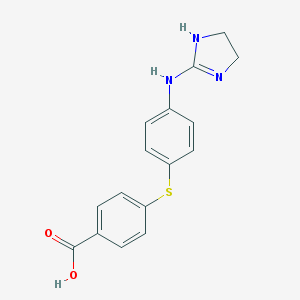 CHEMBL9541 CHEMBL9541 | C16H15N3O2S | 313.375 | 4 / 3 | 2.7 | Yes |
| 41376 | 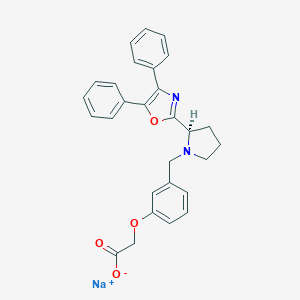 CHEMBL439357 CHEMBL439357 | C28H25N2NaO4 | 476.508 | 6 / 0 | N/A | N/A |
| 41379 | 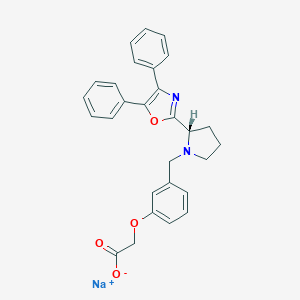 CHEMBL438092 CHEMBL438092 | C28H25N2NaO4 | 476.508 | 6 / 0 | N/A | N/A |
| 41635 | 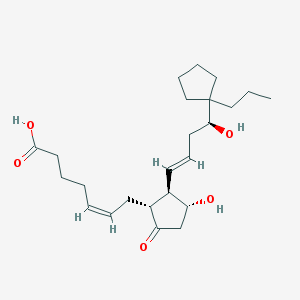 CHEMBL62885 CHEMBL62885 | C24H38O5 | 406.563 | 5 / 3 | 4.1 | Yes |
| 41882 | 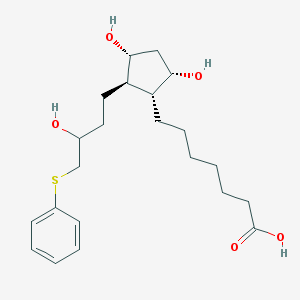 CHEMBL173299 CHEMBL173299 | C22H34O5S | 410.569 | 6 / 4 | 3.8 | Yes |
| 44515 | 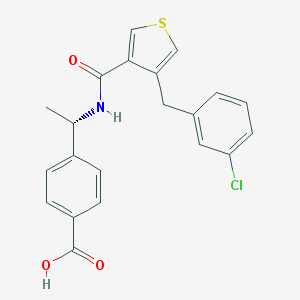 CHEMBL599051 CHEMBL599051 | C21H18ClNO3S | 399.889 | 4 / 2 | 4.9 | Yes |
| 44624 | 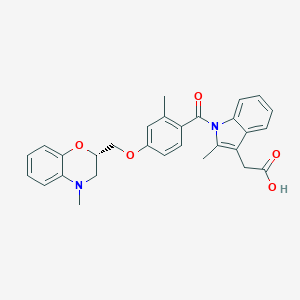 CHEMBL1813120 CHEMBL1813120 | C29H28N2O5 | 484.552 | 6 / 1 | 5.4 | No |
| 45055 | 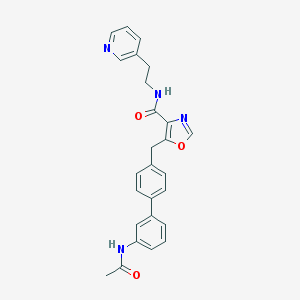 CHEMBL400405 CHEMBL400405 | C26H24N4O3 | 440.503 | 5 / 2 | 3.6 | Yes |
| 47919 | 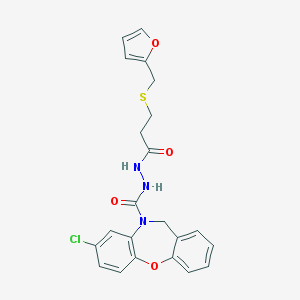 SC-51322 SC-51322 | C22H20ClN3O4S | 457.929 | 5 / 2 | 3.5 | Yes |
| 443592 | 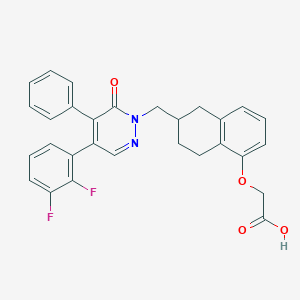 CHEMBL3398228 CHEMBL3398228 | C29H24F2N2O4 | 502.518 | 7 / 1 | 5.1 | No |
| 48279 | 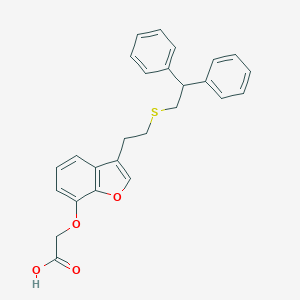 CHEMBL361939 CHEMBL361939 | C26H24O4S | 432.534 | 5 / 1 | 6.2 | No |
| 49361 | 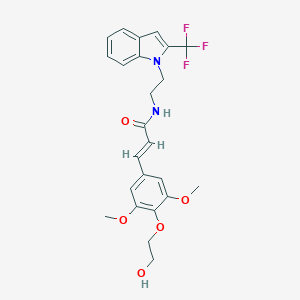 CHEMBL3264231 CHEMBL3264231 | C24H25F3N2O5 | 478.468 | 8 / 2 | 3.5 | Yes |
| 49366 | 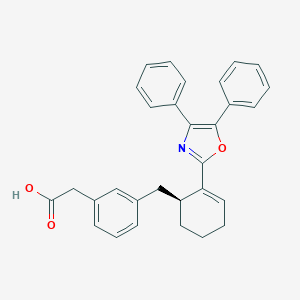 CHEMBL189378 CHEMBL189378 | C30H27NO3 | 449.55 | 4 / 1 | 6.6 | No |
| 49876 | 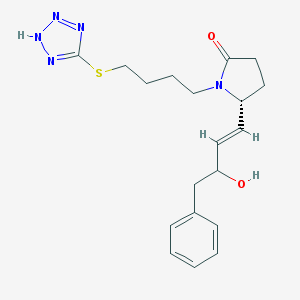 CHEMBL14600 CHEMBL14600 | C19H25N5O2S | 387.502 | 6 / 2 | 2.2 | Yes |
| 553480 | 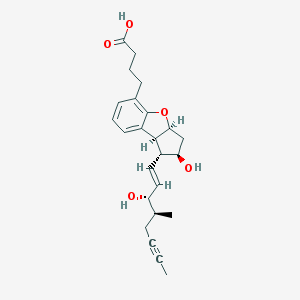 Esuberaprost Esuberaprost | C24H30O5 | 398.499 | 5 / 3 | 3.2 | Yes |
| 553482 | 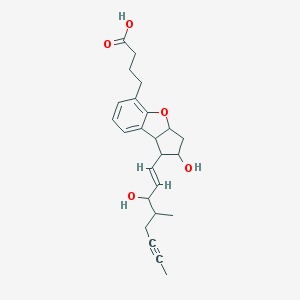 beraprost beraprost | C24H30O5 | 398.499 | 5 / 3 | 3.2 | Yes |
| 50306 | 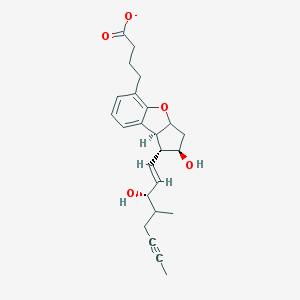 CHEMBL94694 CHEMBL94694 | C24H29O5- | 397.491 | 5 / 2 | 3.9 | Yes |
| 50819 | 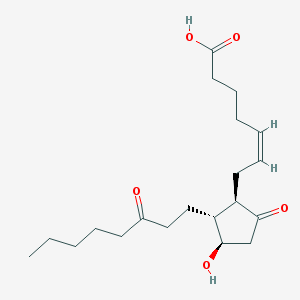 13,14-Dihydro-15-keto-PGE2 13,14-Dihydro-15-keto-PGE2 | C20H32O5 | 352.471 | 5 / 2 | 2.4 | Yes |
| 51188 | 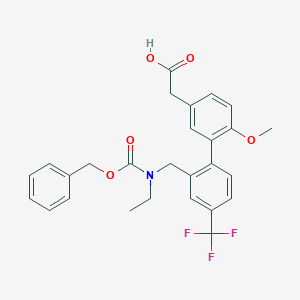 CHEMBL1916700 CHEMBL1916700 | C27H26F3NO5 | 501.502 | 8 / 1 | 5.4 | No |
| 51583 | 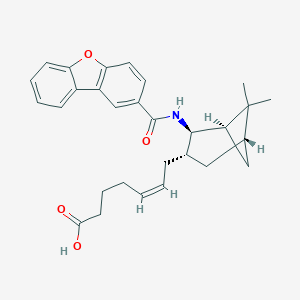 CHEMBL119550 CHEMBL119550 | C29H33NO4 | 459.586 | 4 / 2 | 6.3 | No |
| 52448 | 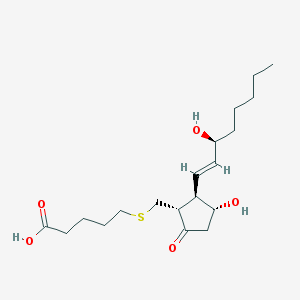 CHEMBL60555 CHEMBL60555 | C19H32O5S | 372.52 | 6 / 3 | 2.2 | Yes |
| 52503 | 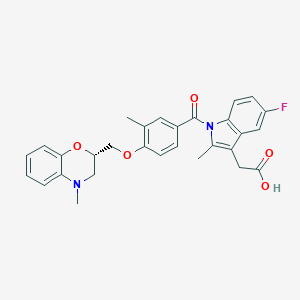 CHEMBL1813278 CHEMBL1813278 | C29H27FN2O5 | 502.542 | 7 / 1 | 5.5 | No |
| 52655 | 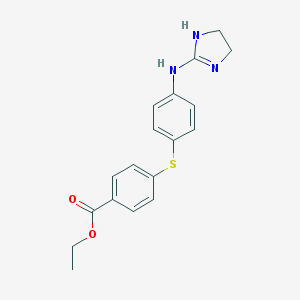 CHEMBL9846 CHEMBL9846 | C18H19N3O2S | 341.429 | 4 / 2 | 3.3 | Yes |
| 537314 | 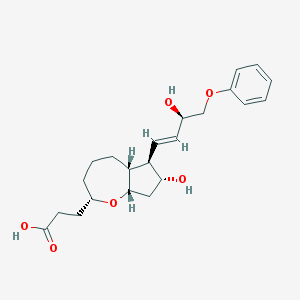 CHEMBL3956817 CHEMBL3956817 | C22H30O6 | 390.476 | 6 / 3 | 2.3 | Yes |
| 537317 | 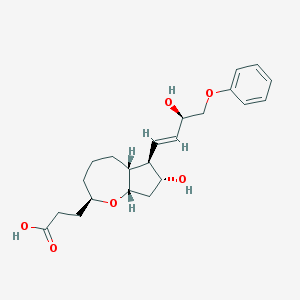 CHEMBL3920347 CHEMBL3920347 | C22H30O6 | 390.476 | 6 / 3 | 2.3 | Yes |
| 443857 | 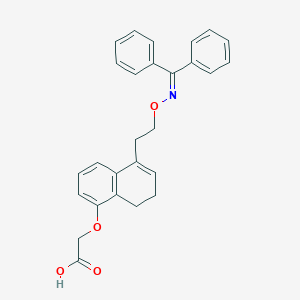 CHEMBL3398237 CHEMBL3398237 | C27H25NO4 | 427.5 | 5 / 1 | 6.0 | No |
| 55072 | 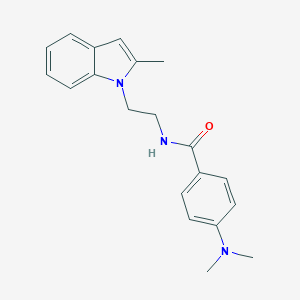 CHEMBL3286634 CHEMBL3286634 | C20H23N3O | 321.424 | 2 / 1 | 3.4 | Yes |
| 55699 | 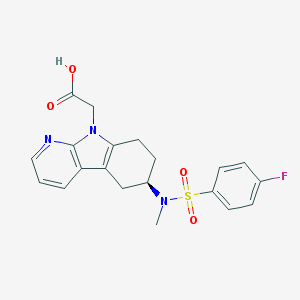 CHEMBL552211 CHEMBL552211 | C20H20FN3O4S | 417.455 | 7 / 1 | 2.7 | Yes |
| 56853 | 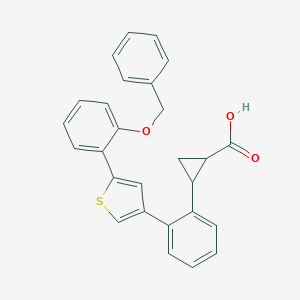 CHEMBL125588 CHEMBL125588 | C27H22O3S | 426.53 | 4 / 1 | 6.1 | No |
| 57129 | 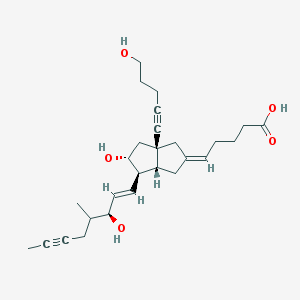 CHEMBL341706 CHEMBL341706 | C27H38O5 | 442.596 | 5 / 4 | 3.0 | Yes |
| 57130 | 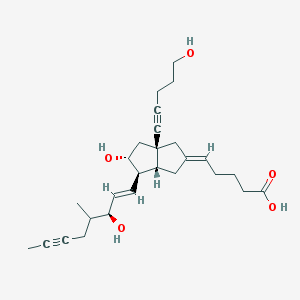 CHEMBL336895 CHEMBL336895 | C27H38O5 | 442.596 | 5 / 4 | 3.0 | Yes |
| 58237 | 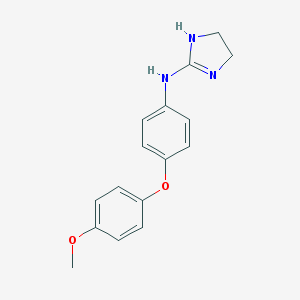 CHEMBL9873 CHEMBL9873 | C16H17N3O2 | 283.331 | 3 / 2 | 2.6 | Yes |
| 60224 | 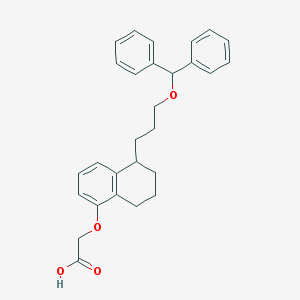 CHEMBL266622 CHEMBL266622 | C28H30O4 | 430.544 | 4 / 1 | 6.4 | No |
| 60368 | 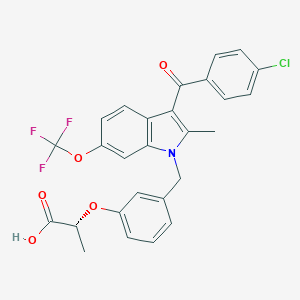 CHEMBL461571 CHEMBL461571 | C27H21ClF3NO5 | 531.912 | 8 / 1 | 7.1 | No |
| 61483 | 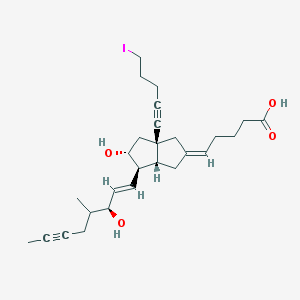 CHEMBL344390 CHEMBL344390 | C27H37IO4 | 552.493 | 4 / 3 | 5.2 | No |
| 61484 | 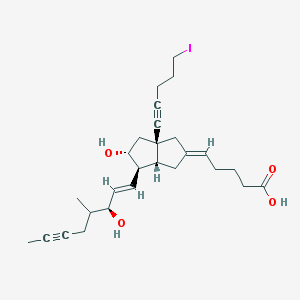 CHEMBL138317 CHEMBL138317 | C27H37IO4 | 552.493 | 4 / 3 | 5.2 | No |
| 61559 | 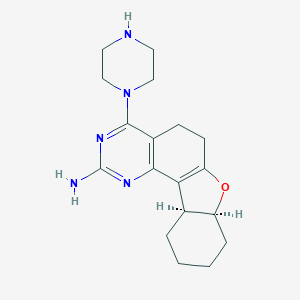 SCHEMBL604437 SCHEMBL604437 | C18H25N5O | 327.432 | 6 / 2 | 1.3 | Yes |
| 63136 | 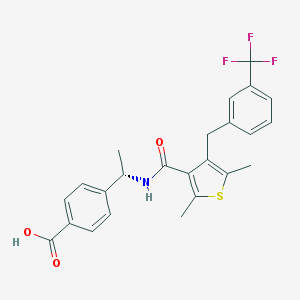 CHEMBL605330 CHEMBL605330 | C24H22F3NO3S | 461.499 | 7 / 2 | 5.9 | No |
| 64445 | 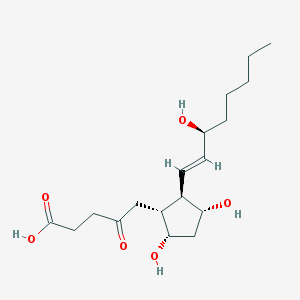 2,3-Dkpgf1alpha 2,3-Dkpgf1alpha | C18H30O6 | 342.432 | 6 / 4 | 0.7 | Yes |
| 65309 | 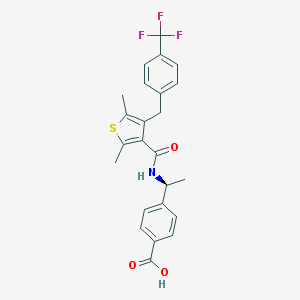 CHEMBL603690 CHEMBL603690 | C24H22F3NO3S | 461.499 | 7 / 2 | 5.9 | No |
| 65567 | 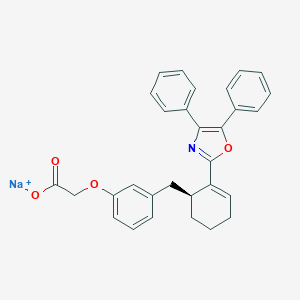 FR-181157 FR-181157 | C30H26NNaO4 | 487.531 | 5 / 0 | N/A | N/A |
| 65748 | 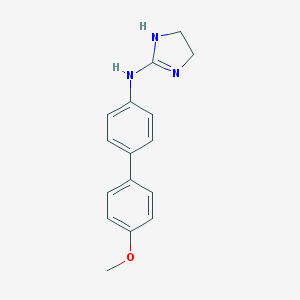 CHEMBL9936 CHEMBL9936 | C16H17N3O | 267.332 | 2 / 2 | 2.5 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218