

You can:
| Name | C-C chemokine receptor type 10 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | CCR10 |
| Synonym | CCR-10 CCR10 CC-CKR-10 G-protein coupled receptor 2 GPR-2 [ Show all ] |
| Disease | Inflammatory disease |
| Length | 362 |
| Amino acid sequence | MGTEATEQVSWGHYSGDEEDAYSAEPLPELCYKADVQAFSRAFQPSVSLTVAALGLAGNGLVLATHLAARRAARSPTSAHLLQLALADLLLALTLPFAAAGALQGWSLGSATCRTISGLYSASFHAGFLFLACISADRYVAIARALPAGPRPSTPGRAHLVSVIVWLLSLLLALPALLFSQDGQREGQRRCRLIFPEGLTQTVKGASAVAQVALGFALPLGVMVACYALLGRTLLAARGPERRRALRVVVALVAAFVVLQLPYSLALLLDTADLLAARERSCPASKRKDVALLVTSGLALARCGLNPVLYAFLGLRFRQDLRRLLRGGSCPSGPQPRRGCPRRPRLSSCSAPTETHSLSWDN |
| UniProt | P46092 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P46092 |
| 3D structure model | This predicted structure model is from GPCR-EXP P46092. |
| BioLiP | N/A |
| Therapeutic Target Database | T71054 |
| ChEMBL | CHEMBL2321628 |
| IUPHAR | 67 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 521447 | 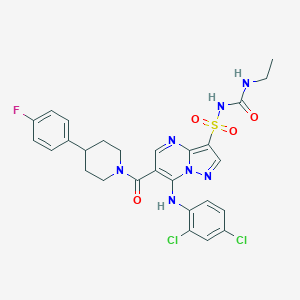 CHEMBL3731686 CHEMBL3731686 | C27H26Cl2FN7O4S | 634.508 | 8 / 3 | 4.7 | No |
| 535954 | 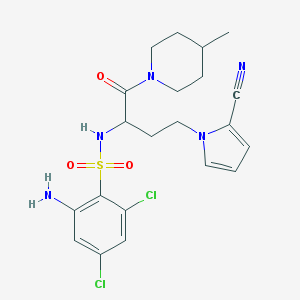 CHEMBL3959737 CHEMBL3959737 | C21H25Cl2N5O3S | 498.423 | 6 / 2 | 3.6 | Yes |
| 521518 | 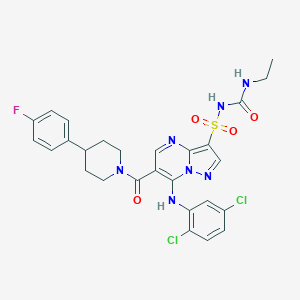 CHEMBL3727384 CHEMBL3727384 | C27H26Cl2FN7O4S | 634.508 | 8 / 3 | 4.7 | No |
| 536042 | 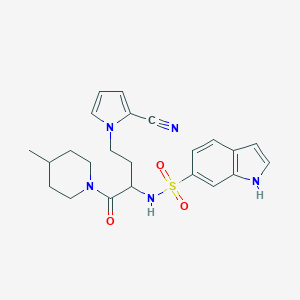 CHEMBL3923733 CHEMBL3923733 | C23H27N5O3S | 453.561 | 5 / 2 | 2.6 | Yes |
| 521603 | 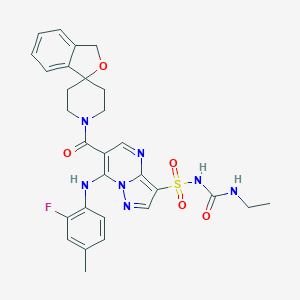 CHEMBL3733065 CHEMBL3733065 | C29H30FN7O5S | 607.661 | 9 / 3 | 2.9 | No |
| 536151 | 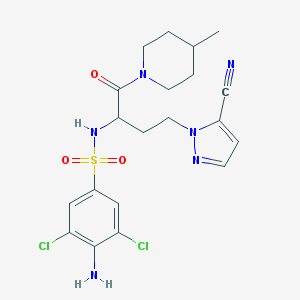 CHEMBL3919363 CHEMBL3919363 | C20H24Cl2N6O3S | 499.411 | 7 / 2 | 2.7 | Yes |
| 521742 | 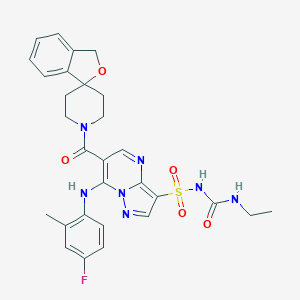 CHEMBL3729731 CHEMBL3729731 | C29H30FN7O5S | 607.661 | 9 / 3 | 2.9 | No |
| 521924 | 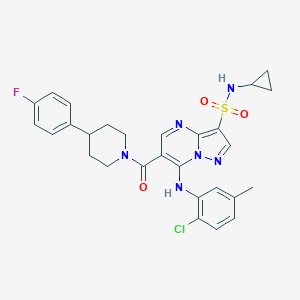 CHEMBL3732878 CHEMBL3732878 | C28H28ClFN6O3S | 583.079 | 8 / 2 | 5.1 | No |
| 522014 | 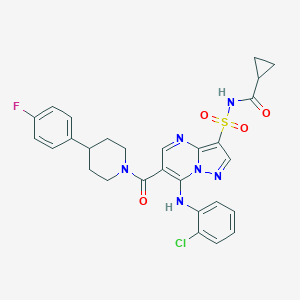 CHEMBL3730003 CHEMBL3730003 | C28H26ClFN6O4S | 597.062 | 8 / 2 | 4.4 | No |
| 536491 | 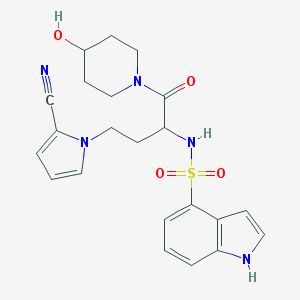 CHEMBL3960700 CHEMBL3960700 | C22H25N5O4S | 455.533 | 6 / 3 | 1.2 | Yes |
| 522314 | 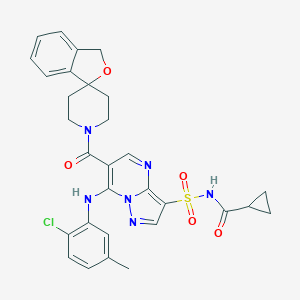 CHEMBL3731390 CHEMBL3731390 | C30H29ClN6O5S | 621.109 | 8 / 2 | 3.7 | No |
| 522403 | 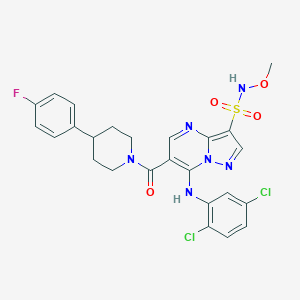 CHEMBL3731840 CHEMBL3731840 | C25H23Cl2FN6O4S | 593.455 | 9 / 2 | 4.8 | No |
| 553415 | 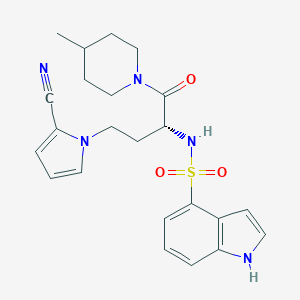 eut-22 eut-22 | C23H27N5O3S | 453.561 | 5 / 2 | 2.6 | Yes |
| 536781 | 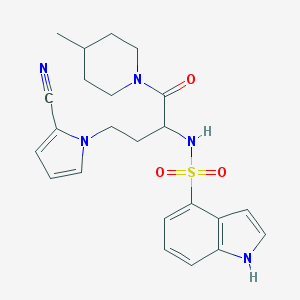 CHEMBL3889627 CHEMBL3889627 | C23H27N5O3S | 453.561 | 5 / 2 | 2.6 | Yes |
| 522594 | 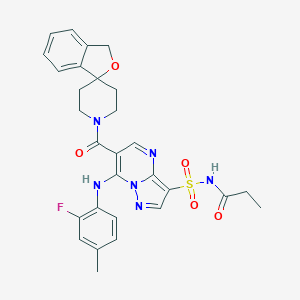 CHEMBL3728353 CHEMBL3728353 | C29H29FN6O5S | 592.646 | 9 / 2 | 3.2 | No |
| 536985 | 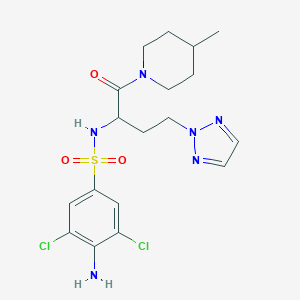 CHEMBL3943972 CHEMBL3943972 | C18H24Cl2N6O3S | 475.389 | 7 / 2 | 2.6 | Yes |
| 522726 | 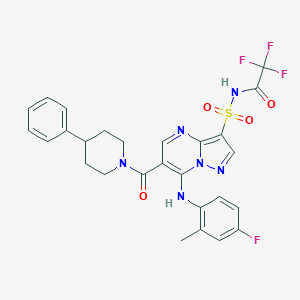 CHEMBL3728979 CHEMBL3728979 | C27H24F4N6O4S | 604.581 | 11 / 2 | 4.8 | No |
| 537039 | 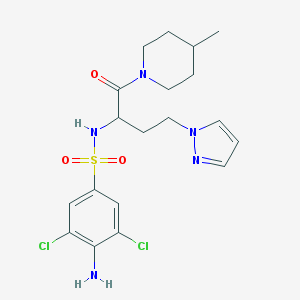 CHEMBL3948828 CHEMBL3948828 | C19H25Cl2N5O3S | 474.401 | 6 / 2 | 2.6 | Yes |
| 537045 | 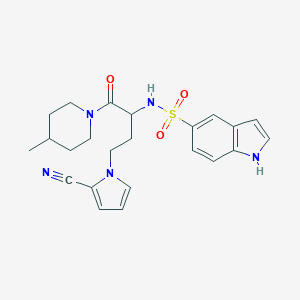 CHEMBL3971886 CHEMBL3971886 | C23H27N5O3S | 453.561 | 5 / 2 | 2.6 | Yes |
| 522939 | 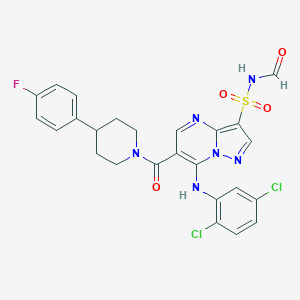 CHEMBL3728246 CHEMBL3728246 | C25H21Cl2FN6O4S | 591.439 | 8 / 2 | 4.6 | No |
| 522962 | 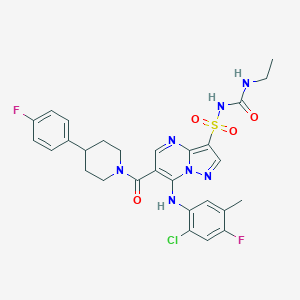 CHEMBL3729606 CHEMBL3729606 | C28H28ClF2N7O4S | 632.084 | 9 / 3 | 4.6 | No |
| 522978 | 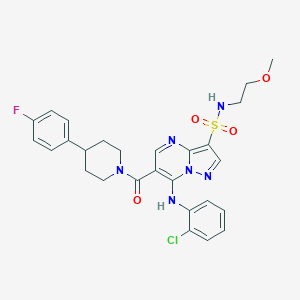 CHEMBL3728616 CHEMBL3728616 | C27H28ClFN6O4S | 587.067 | 9 / 2 | 4.0 | No |
| 523236 | 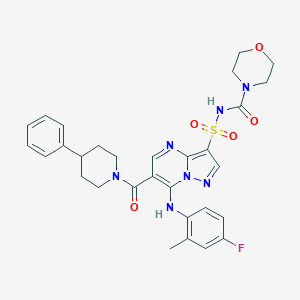 CHEMBL3731147 CHEMBL3731147 | C30H32FN7O5S | 621.688 | 9 / 2 | 3.3 | No |
| 523268 | 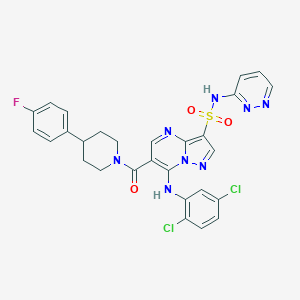 CHEMBL3728071 CHEMBL3728071 | C28H23Cl2FN8O3S | 641.503 | 10 / 2 | 4.6 | No |
| 523507 | 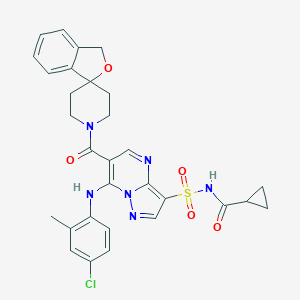 CHEMBL3731775 CHEMBL3731775 | C30H29ClN6O5S | 621.109 | 8 / 2 | 3.7 | No |
| 523700 | 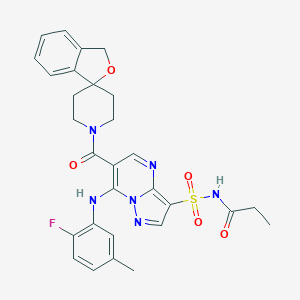 CHEMBL3731933 CHEMBL3731933 | C29H29FN6O5S | 592.646 | 9 / 2 | 3.2 | No |
| 523840 | 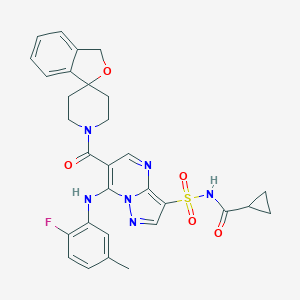 CHEMBL3732508 CHEMBL3732508 | C30H29FN6O5S | 604.657 | 9 / 2 | 3.1 | No |
| 523868 | 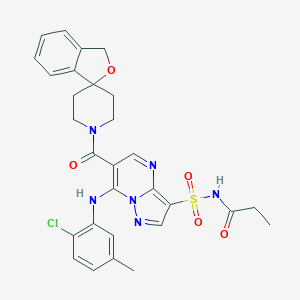 CHEMBL3728121 CHEMBL3728121 | C29H29ClN6O5S | 609.098 | 8 / 2 | 3.7 | No |
| 524014 | 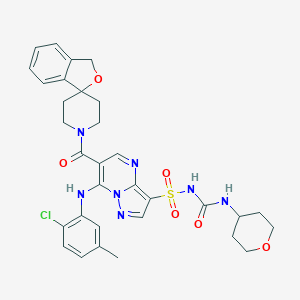 CHEMBL3731832 CHEMBL3731832 | C32H34ClN7O6S | 680.177 | 9 / 3 | 4.0 | No |
| 538137 | 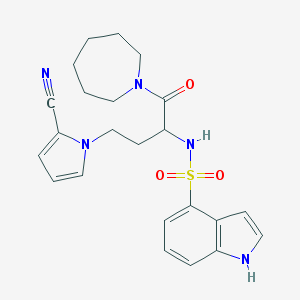 CHEMBL3909737 CHEMBL3909737 | C23H27N5O3S | 453.561 | 5 / 2 | 2.6 | Yes |
| 548932 | 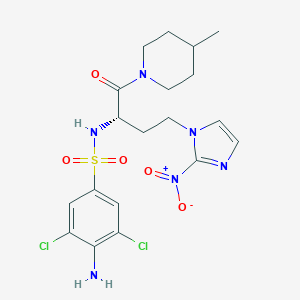 CHEMBL3932155 CHEMBL3932155 | C19H24Cl2N6O5S | 519.398 | 8 / 2 | 2.5 | No |
| 548933 | 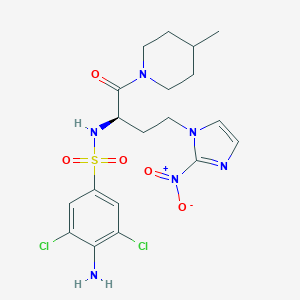 CHEMBL3908983 CHEMBL3908983 | C19H24Cl2N6O5S | 519.398 | 8 / 2 | 2.5 | No |
| 538193 | 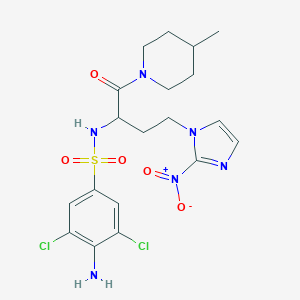 CHEMBL3908230 CHEMBL3908230 | C19H24Cl2N6O5S | 519.398 | 8 / 2 | 2.5 | No |
| 524127 | 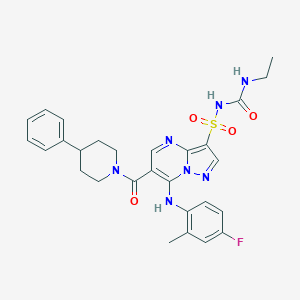 CHEMBL3731438 CHEMBL3731438 | C28H30FN7O4S | 579.651 | 8 / 3 | 3.9 | No |
| 524167 | 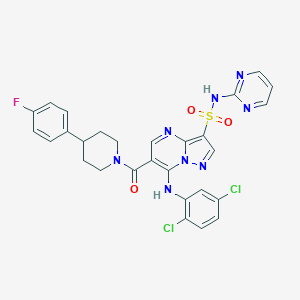 CHEMBL3733165 CHEMBL3733165 | C28H23Cl2FN8O3S | 641.503 | 10 / 2 | 4.9 | No |
| 538296 | 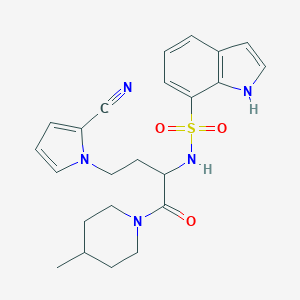 CHEMBL3942284 CHEMBL3942284 | C23H27N5O3S | 453.561 | 5 / 2 | 2.6 | Yes |
| 524254 | 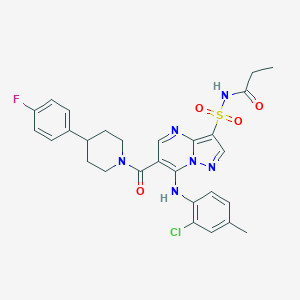 CHEMBL3728606 CHEMBL3728606 | C28H28ClFN6O4S | 599.078 | 8 / 2 | 4.8 | No |
| 538400 | 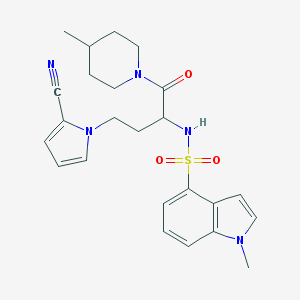 CHEMBL3945769 CHEMBL3945769 | C24H29N5O3S | 467.588 | 5 / 1 | 2.6 | Yes |
| 524987 | 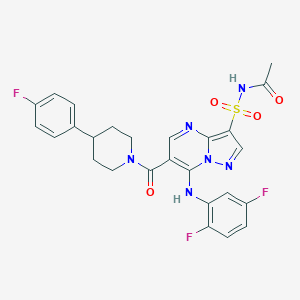 CHEMBL3730752 CHEMBL3730752 | C26H23F3N6O4S | 572.563 | 10 / 2 | 3.5 | No |
| 539149 | 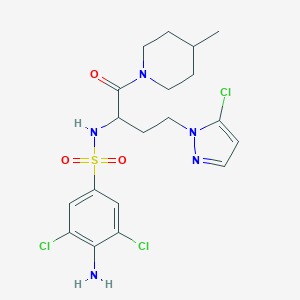 CHEMBL3979967 CHEMBL3979967 | C19H24Cl3N5O3S | 508.843 | 6 / 2 | 3.6 | No |
| 539191 | 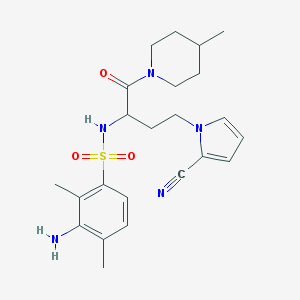 CHEMBL3933303 CHEMBL3933303 | C23H31N5O3S | 457.593 | 6 / 2 | 2.6 | Yes |
| 525211 | 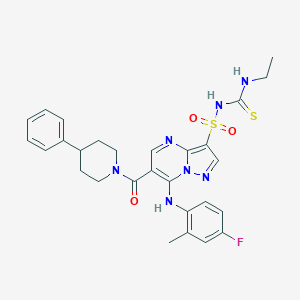 CHEMBL3732652 CHEMBL3732652 | C28H30FN7O3S2 | 595.712 | 8 / 3 | 4.5 | No |
| 525304 | 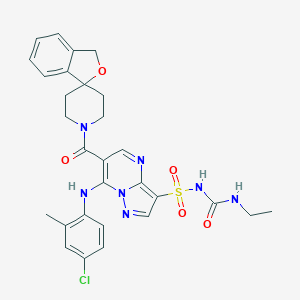 CHEMBL3732876 CHEMBL3732876 | C29H30ClN7O5S | 624.113 | 8 / 3 | 3.4 | No |
| 525327 | 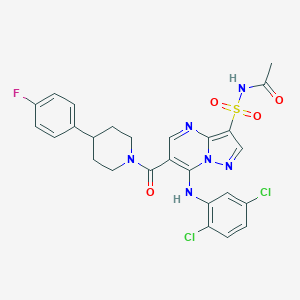 CHEMBL3732868 CHEMBL3732868 | C26H23Cl2FN6O4S | 605.466 | 8 / 2 | 4.6 | No |
| 525394 | 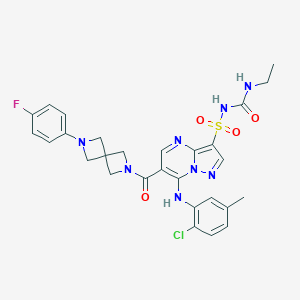 CHEMBL3729178 CHEMBL3729178 | C28H28ClFN8O4S | 627.092 | 9 / 3 | 3.7 | No |
| 525500 | 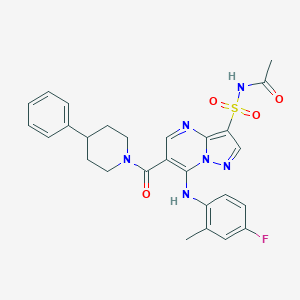 CHEMBL3729146 CHEMBL3729146 | C27H27FN6O4S | 550.609 | 8 / 2 | 3.7 | No |
| 525578 | 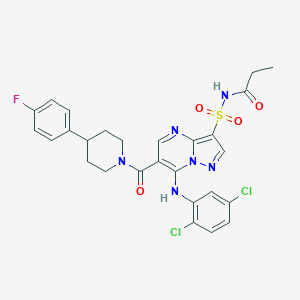 CHEMBL3731219 CHEMBL3731219 | C27H25Cl2FN6O4S | 619.493 | 8 / 2 | 5.0 | No |
| 525605 | 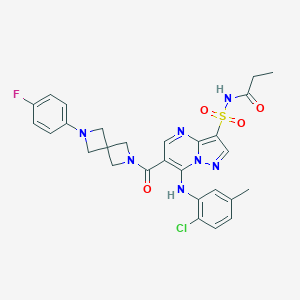 CHEMBL3727497 CHEMBL3727497 | C28H27ClFN7O4S | 612.077 | 9 / 2 | 4.0 | No |
| 525709 | 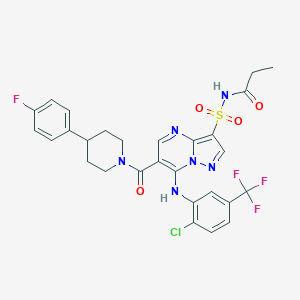 CHEMBL3732344 CHEMBL3732344 | C28H25ClF4N6O4S | 653.05 | 11 / 2 | 5.3 | No |
| 549797 | 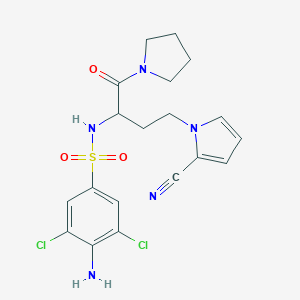 CHEMBL3928968 CHEMBL3928968 | C19H21Cl2N5O3S | 470.369 | 6 / 2 | 2.3 | Yes |
| 549812 | 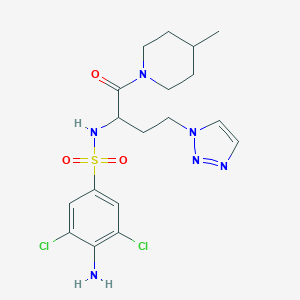 CHEMBL3950532 CHEMBL3950532 | C18H24Cl2N6O3S | 475.389 | 7 / 2 | 2.1 | Yes |
| 539879 | 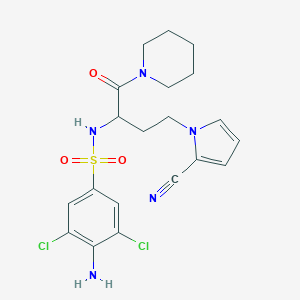 CHEMBL3969321 CHEMBL3969321 | C20H23Cl2N5O3S | 484.396 | 6 / 2 | 2.6 | Yes |
| 525940 | 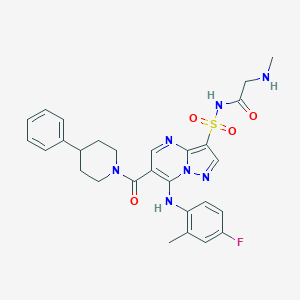 CHEMBL3728332 CHEMBL3728332 | C28H30FN7O4S | 579.651 | 9 / 3 | 3.3 | No |
| 526177 | 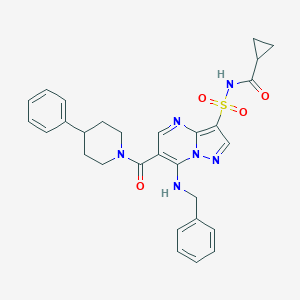 CHEMBL3731138 CHEMBL3731138 | C29H30N6O4S | 558.657 | 7 / 2 | 3.6 | No |
| 526298 | 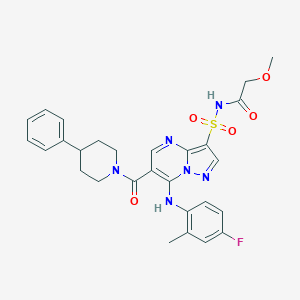 CHEMBL3730468 CHEMBL3730468 | C28H29FN6O5S | 580.635 | 9 / 2 | 3.5 | No |
| 526435 | 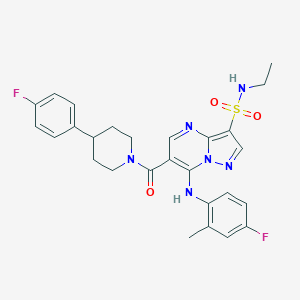 CHEMBL3729372 CHEMBL3729372 | C27H28F2N6O3S | 554.617 | 9 / 2 | 4.3 | No |
| 526523 | 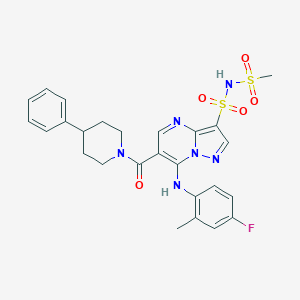 CHEMBL3727981 CHEMBL3727981 | C26H27FN6O5S2 | 586.657 | 10 / 2 | 3.4 | No |
| 526804 | 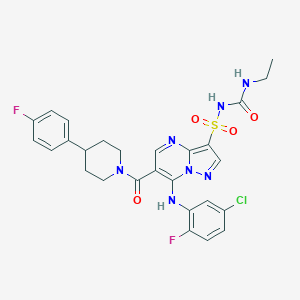 CHEMBL3732672 CHEMBL3732672 | C27H26ClF2N7O4S | 618.057 | 9 / 3 | 4.2 | No |
| 526823 | 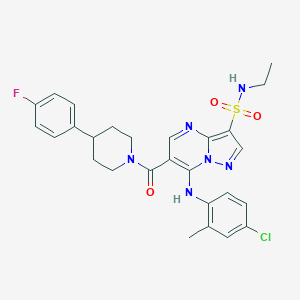 CHEMBL3732501 CHEMBL3732501 | C27H28ClFN6O3S | 571.068 | 8 / 2 | 4.9 | No |
| 526863 | 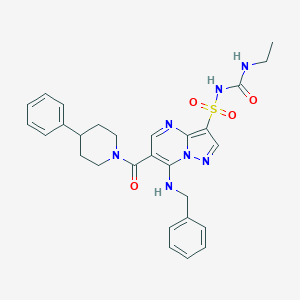 CHEMBL3730497 CHEMBL3730497 | C28H31N7O4S | 561.661 | 7 / 3 | 3.3 | No |
| 550245 | 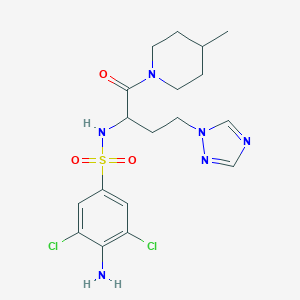 CHEMBL3973467 CHEMBL3973467 | C18H24Cl2N6O3S | 475.389 | 7 / 2 | 2.4 | Yes |
| 527155 | 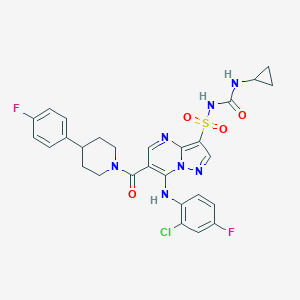 CHEMBL3731456 CHEMBL3731456 | C28H26ClF2N7O4S | 630.068 | 9 / 3 | 5.0 | No |
| 550311 | 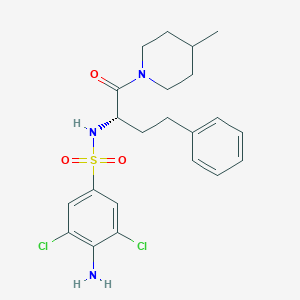 CHEMBL3964878 CHEMBL3964878 | C22H27Cl2N3O3S | 484.436 | 5 / 2 | 4.6 | Yes |
| 541068 | 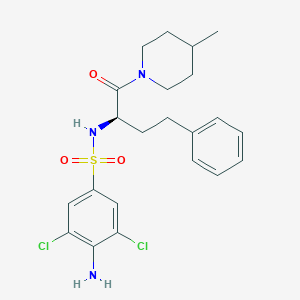 CHEMBL3926111 CHEMBL3926111 | C22H27Cl2N3O3S | 484.436 | 5 / 2 | 4.6 | Yes |
| 550329 | 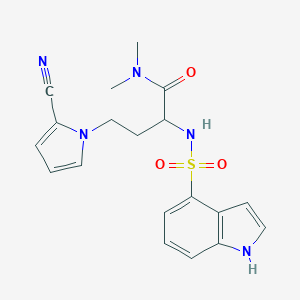 CHEMBL3979899 CHEMBL3979899 | C19H21N5O3S | 399.469 | 5 / 2 | 1.4 | Yes |
| 527288 | 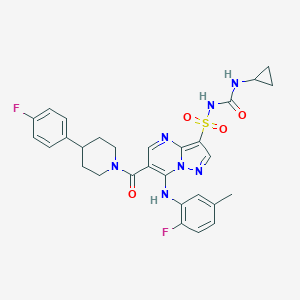 CHEMBL3728893 CHEMBL3728893 | C29H29F2N7O4S | 609.653 | 9 / 3 | 4.7 | No |
| 550419 | 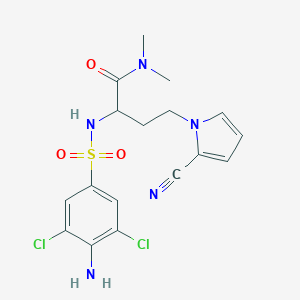 CHEMBL3900672 CHEMBL3900672 | C17H19Cl2N5O3S | 444.331 | 6 / 2 | 1.8 | Yes |
| 527463 | 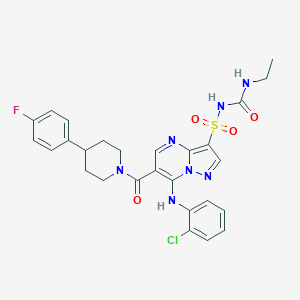 CHEMBL3728376 CHEMBL3728376 | C27H27ClFN7O4S | 600.066 | 8 / 3 | 4.1 | No |
| 541524 | 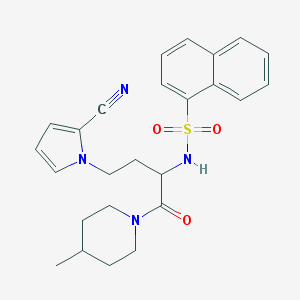 CHEMBL3975141 CHEMBL3975141 | C25H28N4O3S | 464.584 | 5 / 1 | 3.8 | Yes |
| 527762 | 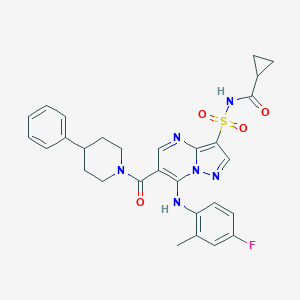 N-[7-(4-fluoro-2-methylphenylamino)-6-(4-phenylpiperidine-1-carbonyl)pyrazolo[1,5-a]pyrimidin-3-ylsulfonyl]cyclopropanecarboxamide N-[7-(4-fluoro-2-methylphenylamino)-6-(4-phenylpiperidine-1-carbonyl)pyrazolo[1,5-a]pyrimidin-3-ylsulfonyl]cyclopropanecarboxamide | C29H29FN6O4S | 576.647 | 8 / 2 | 4.1 | No |
| 528222 | 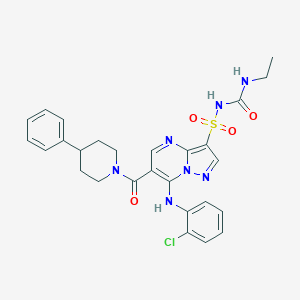 CHEMBL3728115 CHEMBL3728115 | C27H28ClN7O4S | 582.076 | 7 / 3 | 4.0 | No |
| 542337 | 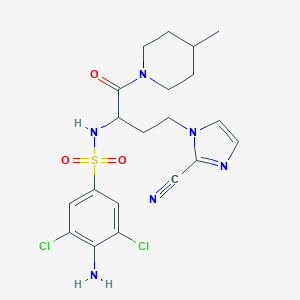 CHEMBL3951650 CHEMBL3951650 | C20H24Cl2N6O3S | 499.411 | 7 / 2 | 2.4 | Yes |
| 528670 | 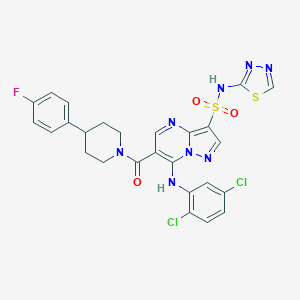 CHEMBL3733183 CHEMBL3733183 | C26H21Cl2FN8O3S2 | 647.525 | 11 / 2 | 5.1 | No |
| 528730 | 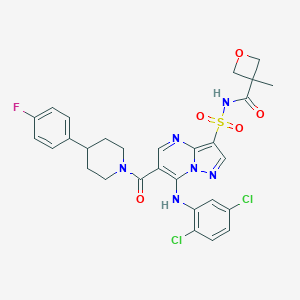 CHEMBL3732275 CHEMBL3732275 | C29H27Cl2FN6O5S | 661.53 | 9 / 2 | 4.5 | No |
| 542684 | 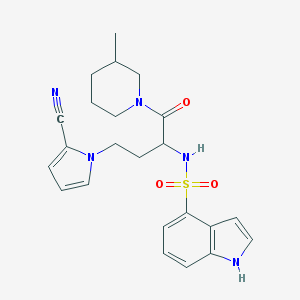 CHEMBL3890608 CHEMBL3890608 | C23H27N5O3S | 453.561 | 5 / 2 | 2.6 | Yes |
| 528971 | 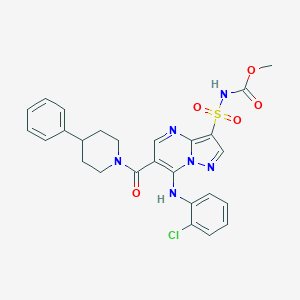 CHEMBL3733316 CHEMBL3733316 | C26H25ClN6O5S | 569.033 | 8 / 2 | 4.2 | No |
| 529049 | 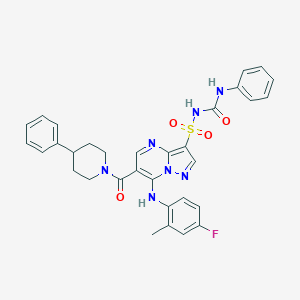 CHEMBL3731999 CHEMBL3731999 | C32H30FN7O4S | 627.695 | 8 / 3 | 5.6 | No |
| 542989 | 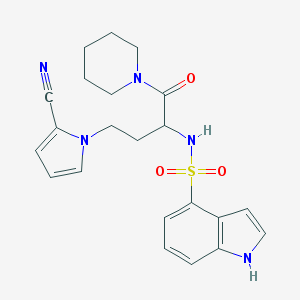 CHEMBL3971826 CHEMBL3971826 | C22H25N5O3S | 439.534 | 5 / 2 | 2.2 | Yes |
| 529096 | 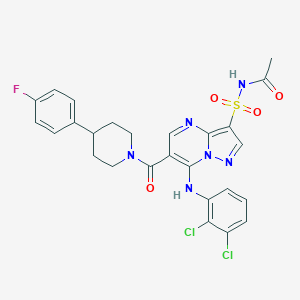 CHEMBL3729920 CHEMBL3729920 | C26H23Cl2FN6O4S | 605.466 | 8 / 2 | 4.6 | No |
| 529097 | 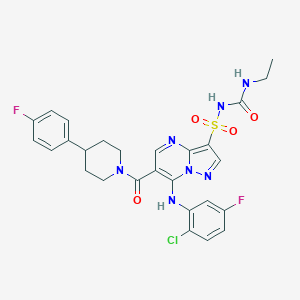 CHEMBL3728619 CHEMBL3728619 | C27H26ClF2N7O4S | 618.057 | 9 / 3 | 4.2 | No |
| 529272 | 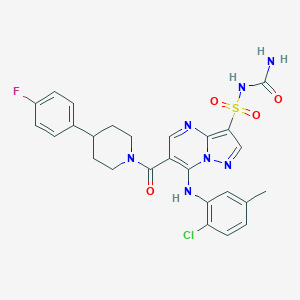 CHEMBL3728998 CHEMBL3728998 | C26H25ClFN7O4S | 586.039 | 8 / 3 | 3.7 | No |
| 529619 | 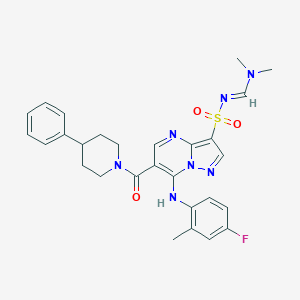 CHEMBL3731666 CHEMBL3731666 | C28H30FN7O3S | 563.652 | 8 / 1 | 4.1 | No |
| 529693 | 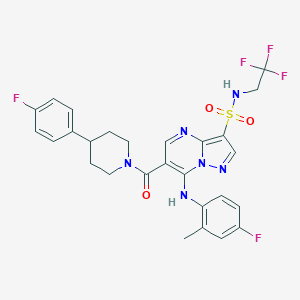 CHEMBL3727493 CHEMBL3727493 | C27H25F5N6O3S | 608.588 | 12 / 2 | 5.1 | No |
| 529838 | 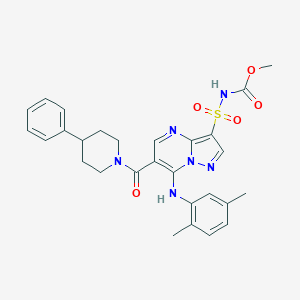 CHEMBL3728415 CHEMBL3728415 | C28H30N6O5S | 562.645 | 8 / 2 | 4.3 | No |
| 530021 | 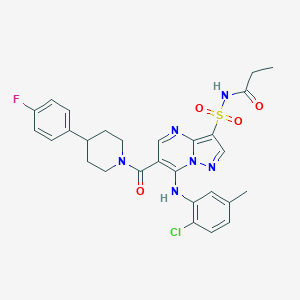 CHEMBL3730157 CHEMBL3730157 | C28H28ClFN6O4S | 599.078 | 8 / 2 | 4.8 | No |
| 544263 | 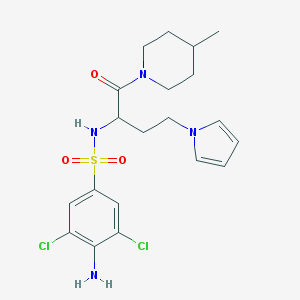 CHEMBL3940706 CHEMBL3940706 | C20H26Cl2N4O3S | 473.413 | 5 / 2 | 3.0 | Yes |
| 544327 | 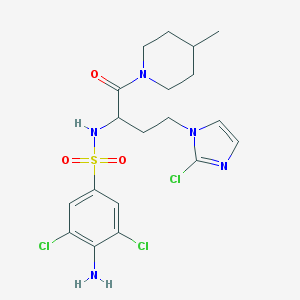 CHEMBL3964547 CHEMBL3964547 | C19H24Cl3N5O3S | 508.843 | 6 / 2 | 3.3 | No |
| 530407 | 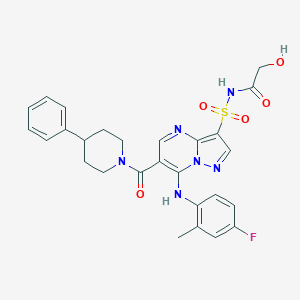 CHEMBL3733128 CHEMBL3733128 | C27H27FN6O5S | 566.608 | 9 / 3 | 3.0 | No |
| 530445 | 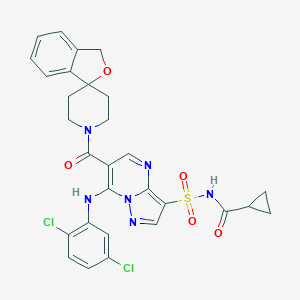 CHEMBL3732824 CHEMBL3732824 | C29H26Cl2N6O5S | 641.524 | 8 / 2 | 3.9 | No |
| 530456 | 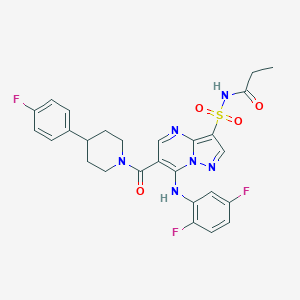 CHEMBL3729504 CHEMBL3729504 | C27H25F3N6O4S | 586.59 | 10 / 2 | 4.0 | No |
| 530457 | 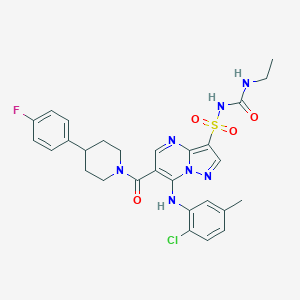 CHEMBL3727436 CHEMBL3727436 | C28H29ClFN7O4S | 614.093 | 8 / 3 | 4.5 | No |
| 544681 | 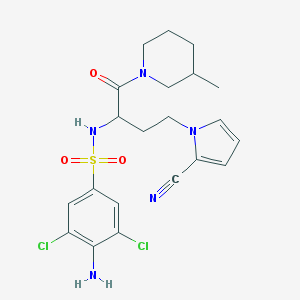 CHEMBL3955499 CHEMBL3955499 | C21H25Cl2N5O3S | 498.423 | 6 / 2 | 3.1 | Yes |
| 530703 | 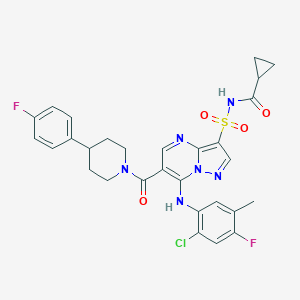 CHEMBL3731872 CHEMBL3731872 | C29H27ClF2N6O4S | 629.08 | 9 / 2 | 4.8 | No |
| 544839 | 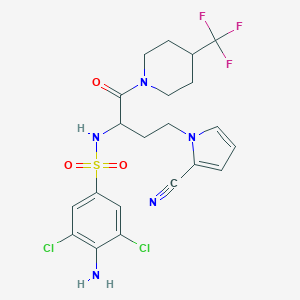 CHEMBL3946799 CHEMBL3946799 | C21H22Cl2F3N5O3S | 552.394 | 9 / 2 | 3.7 | No |
| 552025 | 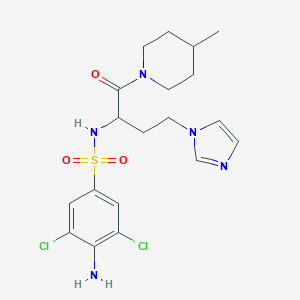 CHEMBL3944089 CHEMBL3944089 | C19H25Cl2N5O3S | 474.401 | 6 / 2 | 2.4 | Yes |
| 530861 | 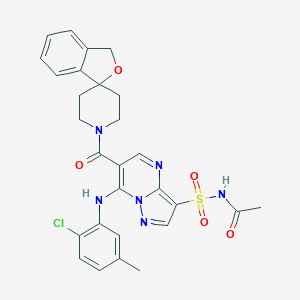 CHEMBL3732278 CHEMBL3732278 | C28H27ClN6O5S | 595.071 | 8 / 2 | 3.2 | No |
| 530943 | 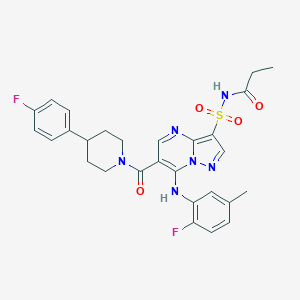 CHEMBL3731105 CHEMBL3731105 | C28H28F2N6O4S | 582.627 | 9 / 2 | 4.2 | No |
| 531009 | 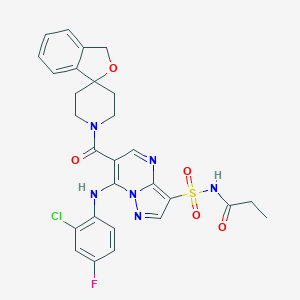 CHEMBL3732825 CHEMBL3732825 | C28H26ClFN6O5S | 613.061 | 9 / 2 | 3.4 | No |
| 545163 | 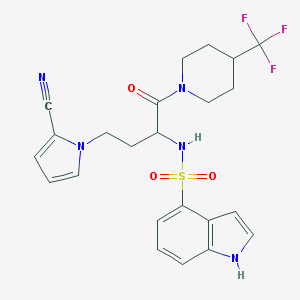 CHEMBL3891692 CHEMBL3891692 | C23H24F3N5O3S | 507.532 | 8 / 2 | 3.2 | No |
| 545262 | 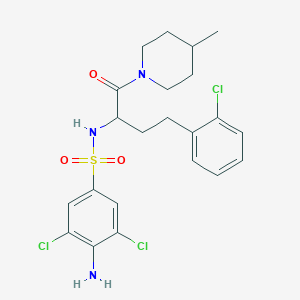 CHEMBL3891289 CHEMBL3891289 | C22H26Cl3N3O3S | 518.878 | 5 / 2 | 5.2 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218