

You can:
| Name | 5-hydroxytryptamine receptor 1B |
|---|---|
| Species | Cricetulus griseus (Chinese hamster) |
| Gene | HTR1B |
| Synonym | 5-HT-1B 5-HT1B Serotonin receptor 1B |
| Disease | N/A for non-human GPCRs |
| Length | 386 |
| Amino acid sequence | MEEQGIQCAPPPPAASQTGVPLVNLSHNCSAESHIYQDSIALPWKVLLVALLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPVSTMYTVTGRWTLGQVVCDFWLSSDITCCTASIMHLCVIALDRYWAITDAVEYAAKRTPKRAAIMIALVWVFSISISLPPFFWRQAKAEEEVLTCLVNTDHVLYTVYSTGGAFYLPTLLLIALYGRIYVEARSRILKQTPNKTGKRLTRAQLITDSPGSTTSVTSINSRAPDLPSESGSPVYVNQVKVRVSDALLEKKKLMAARERKATKTLGIILGAFIVCWLPFFIISLVMPICKDACWFHMATLDFFNWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFKCAG |
| UniProt | P46636 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3707466 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 2943 | 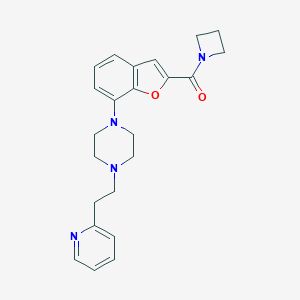 SCHEMBL931468 SCHEMBL931468 | C23H26N4O2 | 390.487 | 5 / 0 | 3.1 | Yes |
| 14729 | 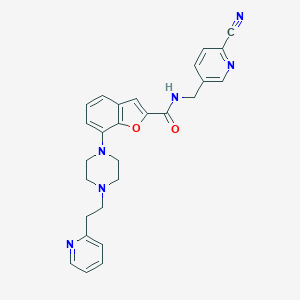 SCHEMBL931803 SCHEMBL931803 | C27H26N6O2 | 466.545 | 7 / 1 | 3.3 | Yes |
| 23268 | 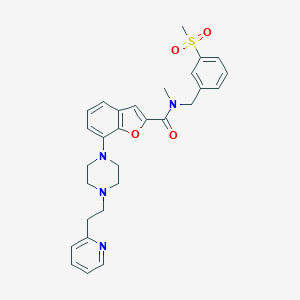 SCHEMBL932715 SCHEMBL932715 | C29H32N4O4S | 532.659 | 7 / 0 | 3.7 | No |
| 28848 | 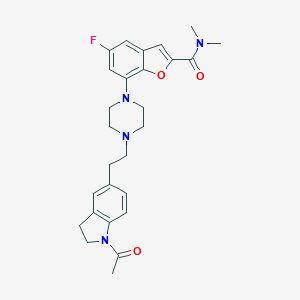 SCHEMBL932393 SCHEMBL932393 | C27H31FN4O3 | 478.568 | 6 / 0 | 3.5 | Yes |
| 32296 | 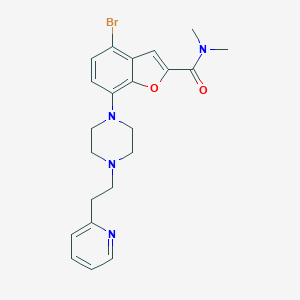 SCHEMBL931086 SCHEMBL931086 | C22H25BrN4O2 | 457.372 | 5 / 0 | 3.7 | Yes |
| 37618 | 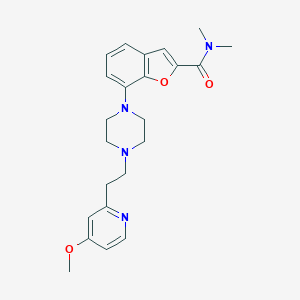 SCHEMBL930901 SCHEMBL930901 | C23H28N4O3 | 408.502 | 6 / 0 | 2.9 | Yes |
| 39618 | 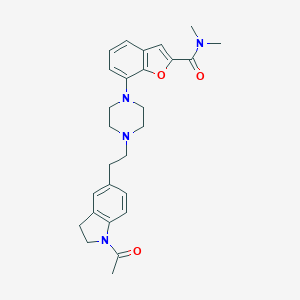 SCHEMBL943980 SCHEMBL943980 | C27H32N4O3 | 460.578 | 5 / 0 | 3.4 | Yes |
| 42688 | 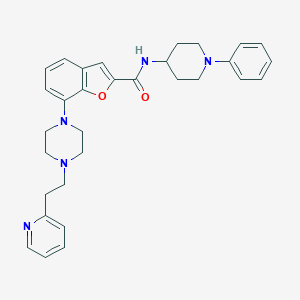 SCHEMBL3791467 SCHEMBL3791467 | C31H35N5O2 | 509.654 | 6 / 1 | 5.2 | No |
| 43034 | 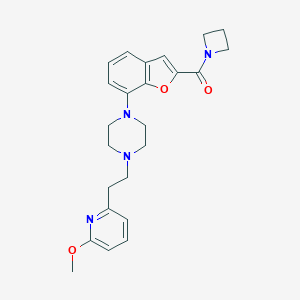 SCHEMBL931445 SCHEMBL931445 | C24H28N4O3 | 420.513 | 6 / 0 | 3.4 | Yes |
| 67959 | 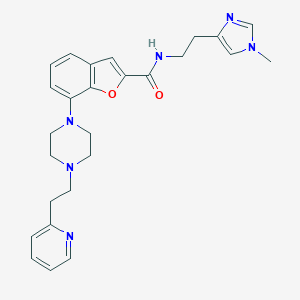 SCHEMBL932014 SCHEMBL932014 | C26H30N6O2 | 458.566 | 6 / 1 | 2.9 | Yes |
| 82021 | 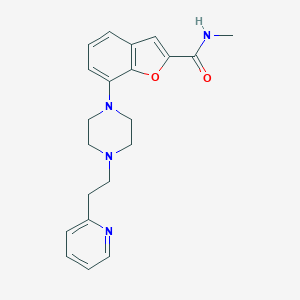 SCHEMBL932198 SCHEMBL932198 | C21H24N4O2 | 364.449 | 5 / 1 | 2.8 | Yes |
| 88229 | 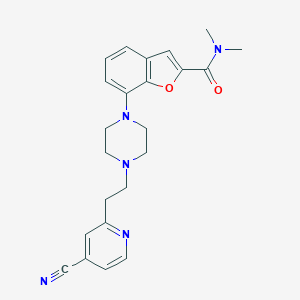 SCHEMBL931928 SCHEMBL931928 | C23H25N5O2 | 403.486 | 6 / 0 | 2.7 | Yes |
| 93345 | 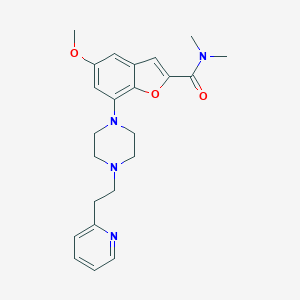 SCHEMBL932678 SCHEMBL932678 | C23H28N4O3 | 408.502 | 6 / 0 | 2.9 | Yes |
| 96162 | 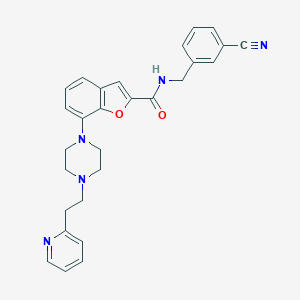 SCHEMBL932078 SCHEMBL932078 | C28H27N5O2 | 465.557 | 6 / 1 | 4.0 | Yes |
| 97599 | 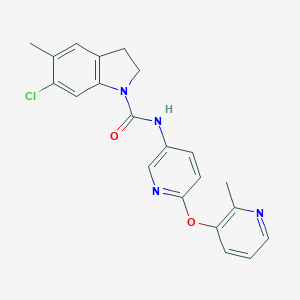 SB 242084 SB 242084 | C21H19ClN4O2 | 394.859 | 4 / 1 | 3.8 | Yes |
| 115785 | 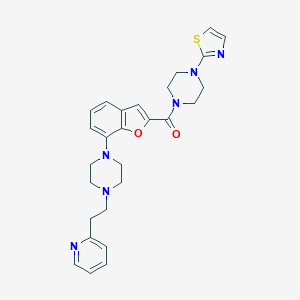 SCHEMBL931495 SCHEMBL931495 | C27H30N6O2S | 502.637 | 8 / 0 | 3.9 | No |
| 121021 | 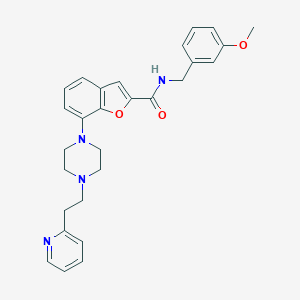 SCHEMBL931747 SCHEMBL931747 | C28H30N4O3 | 470.573 | 6 / 1 | 4.3 | Yes |
| 125778 | 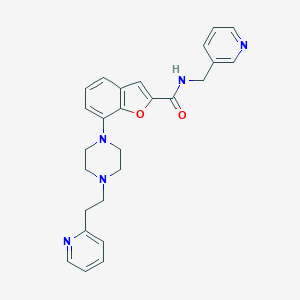 SCHEMBL3782291 SCHEMBL3782291 | C26H27N5O2 | 441.535 | 6 / 1 | 3.2 | Yes |
| 126241 | 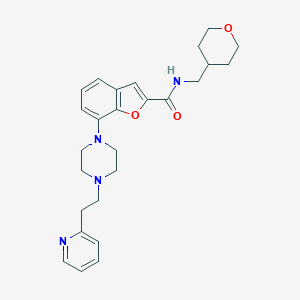 SCHEMBL932228 SCHEMBL932228 | C26H32N4O3 | 448.567 | 6 / 1 | 3.4 | Yes |
| 129625 | 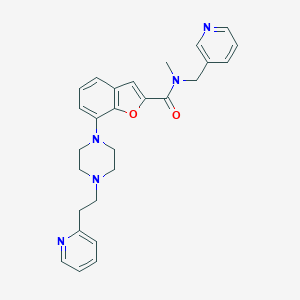 SCHEMBL931161 SCHEMBL931161 | C27H29N5O2 | 455.562 | 6 / 0 | 3.4 | Yes |
| 131853 | 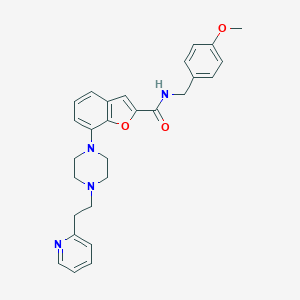 SCHEMBL931675 SCHEMBL931675 | C28H30N4O3 | 470.573 | 6 / 1 | 4.3 | Yes |
| 152295 | 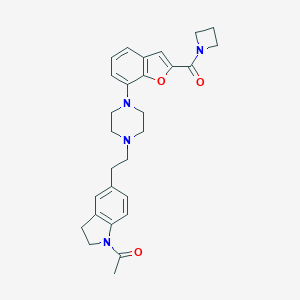 SCHEMBL932199 SCHEMBL932199 | C28H32N4O3 | 472.589 | 5 / 0 | 3.5 | Yes |
| 160142 | 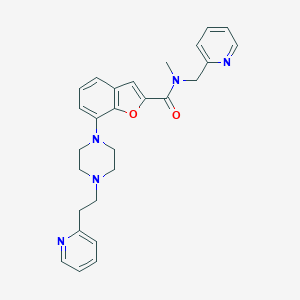 SCHEMBL931519 SCHEMBL931519 | C27H29N5O2 | 455.562 | 6 / 0 | 3.4 | Yes |
| 163456 | 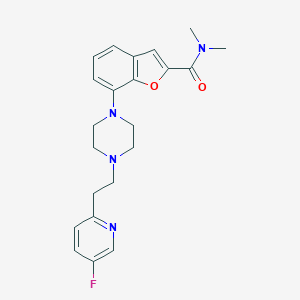 SCHEMBL932091 SCHEMBL932091 | C22H25FN4O2 | 396.466 | 6 / 0 | 3.1 | Yes |
| 169001 | 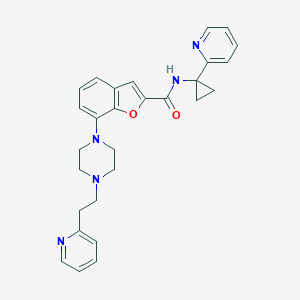 SCHEMBL3787910 SCHEMBL3787910 | C28H29N5O2 | 467.573 | 6 / 1 | 3.6 | Yes |
| 169662 | 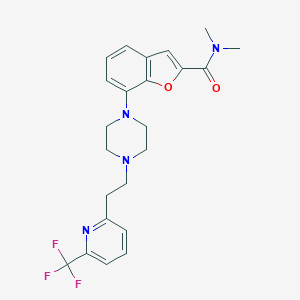 SCHEMBL932062 SCHEMBL932062 | C23H25F3N4O2 | 446.474 | 8 / 0 | 3.9 | Yes |
| 176751 | 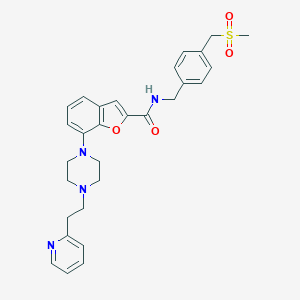 SCHEMBL931797 SCHEMBL931797 | C29H32N4O4S | 532.659 | 7 / 1 | 3.5 | No |
| 181389 | 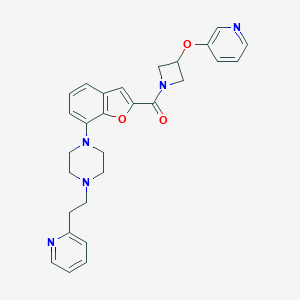 SCHEMBL931121 SCHEMBL931121 | C28H29N5O3 | 483.572 | 7 / 0 | 3.4 | Yes |
| 189908 | 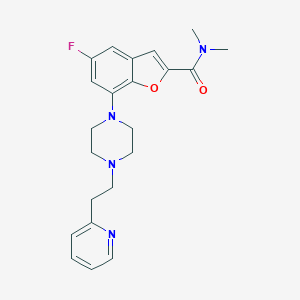 SCHEMBL931922 SCHEMBL931922 | C22H25FN4O2 | 396.466 | 6 / 0 | 3.1 | Yes |
| 207909 | 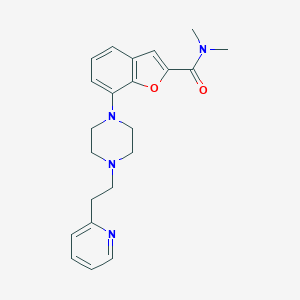 SCHEMBL931632 SCHEMBL931632 | C22H26N4O2 | 378.476 | 5 / 0 | 3.0 | Yes |
| 218273 | 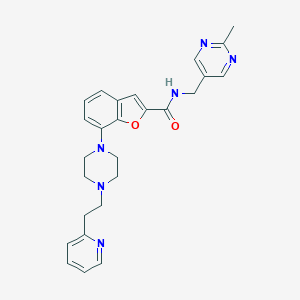 SCHEMBL932470 SCHEMBL932470 | C26H28N6O2 | 456.55 | 7 / 1 | 3.0 | Yes |
| 230685 | 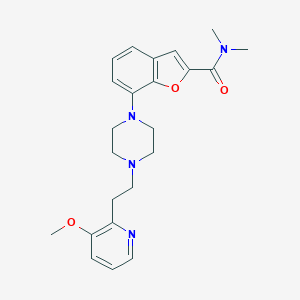 SCHEMBL931521 SCHEMBL931521 | C23H28N4O3 | 408.502 | 6 / 0 | 2.9 | Yes |
| 232352 | 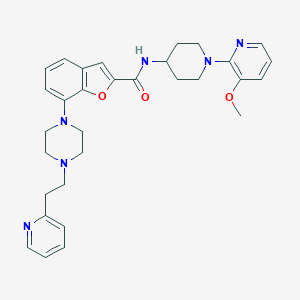 SCHEMBL3791937 SCHEMBL3791937 | C31H36N6O3 | 540.668 | 8 / 1 | 4.4 | No |
| 238040 | 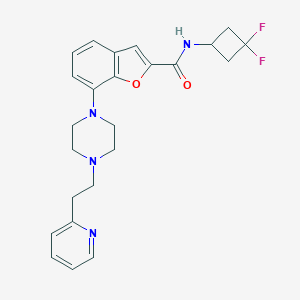 SCHEMBL931597 SCHEMBL931597 | C24H26F2N4O2 | 440.495 | 7 / 1 | 4.0 | Yes |
| 528601 | 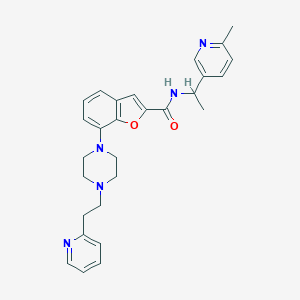 SCHEMBL931452 SCHEMBL931452 | C28H31N5O2 | 469.589 | 6 / 1 | 4.0 | Yes |
| 255025 | 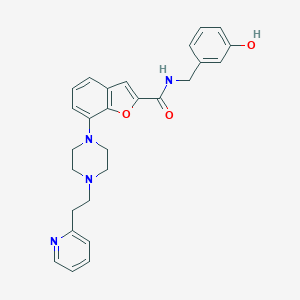 SCHEMBL931619 SCHEMBL931619 | C27H28N4O3 | 456.546 | 6 / 2 | 3.9 | Yes |
| 310526 | 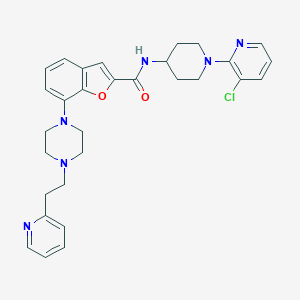 SCHEMBL3790615 SCHEMBL3790615 | C30H33ClN6O2 | 545.084 | 7 / 1 | 5.1 | No |
| 317342 | 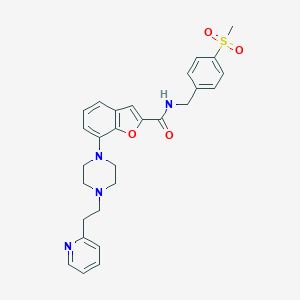 SCHEMBL3785805 SCHEMBL3785805 | C28H30N4O4S | 518.632 | 7 / 1 | 3.5 | No |
| 330713 | 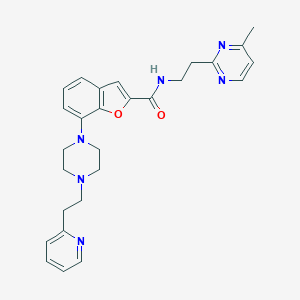 SCHEMBL932656 SCHEMBL932656 | C27H30N6O2 | 470.577 | 7 / 1 | 3.5 | Yes |
| 337548 | 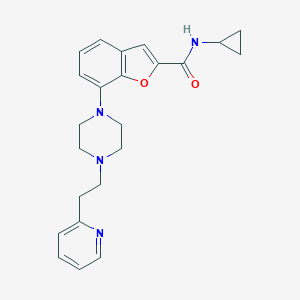 SCHEMBL931976 SCHEMBL931976 | C23H26N4O2 | 390.487 | 5 / 1 | 3.3 | Yes |
| 338531 | 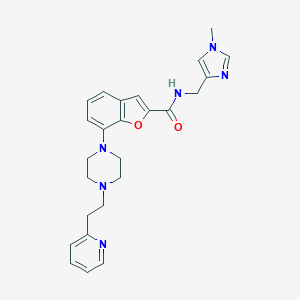 SCHEMBL932193 SCHEMBL932193 | C25H28N6O2 | 444.539 | 6 / 1 | 2.4 | Yes |
| 348931 | 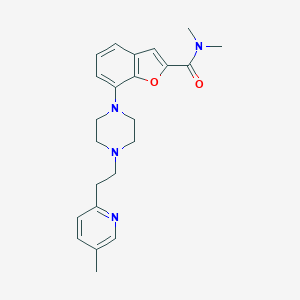 SCHEMBL932148 SCHEMBL932148 | C23H28N4O2 | 392.503 | 5 / 0 | 3.3 | Yes |
| 356002 | 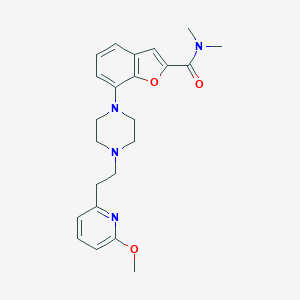 SCHEMBL931392 SCHEMBL931392 | C23H28N4O3 | 408.502 | 6 / 0 | 3.3 | Yes |
| 362378 | 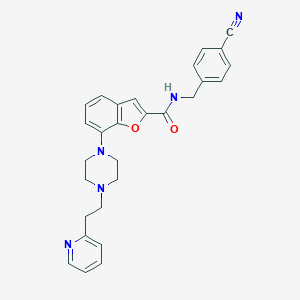 SCHEMBL931585 SCHEMBL931585 | C28H27N5O2 | 465.557 | 6 / 1 | 4.0 | Yes |
| 371266 | 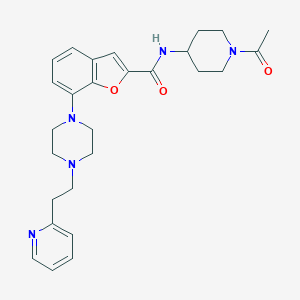 SCHEMBL3787659 SCHEMBL3787659 | C27H33N5O3 | 475.593 | 6 / 1 | 2.8 | Yes |
| 375167 | 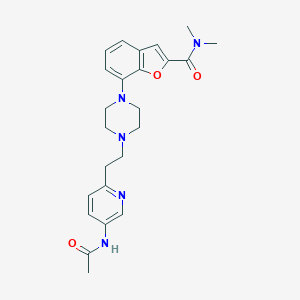 SCHEMBL931714 SCHEMBL931714 | C24H29N5O3 | 435.528 | 6 / 1 | 2.2 | Yes |
| 375436 | 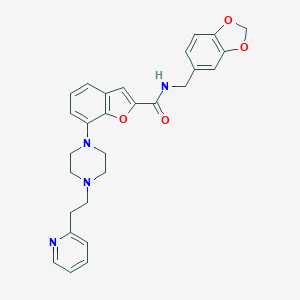 SCHEMBL931141 SCHEMBL931141 | C28H28N4O4 | 484.556 | 7 / 1 | 4.1 | Yes |
| 392664 | 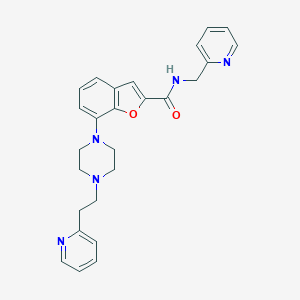 SCHEMBL931346 SCHEMBL931346 | C26H27N5O2 | 441.535 | 6 / 1 | 3.3 | Yes |
| 532726 | 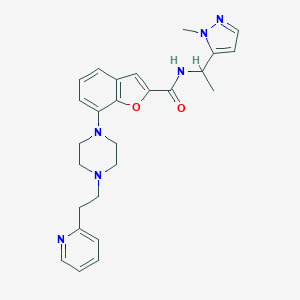 SCHEMBL943679 SCHEMBL943679 | C26H30N6O2 | 458.566 | 6 / 1 | 3.0 | Yes |
| 402844 | 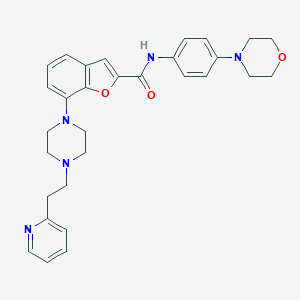 SCHEMBL3785037 SCHEMBL3785037 | C30H33N5O3 | 511.626 | 7 / 1 | 4.1 | No |
| 404301 | 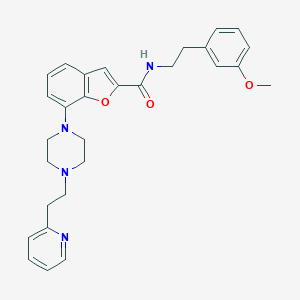 SCHEMBL931549 SCHEMBL931549 | C29H32N4O3 | 484.6 | 6 / 1 | 4.7 | Yes |
| 411894 | 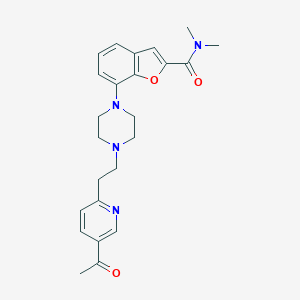 SCHEMBL931509 SCHEMBL931509 | C24H28N4O3 | 420.513 | 6 / 0 | 2.7 | Yes |
| 533711 | 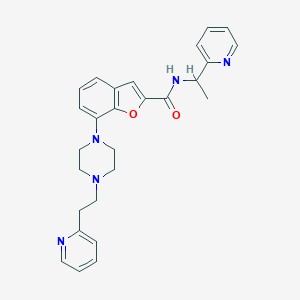 SCHEMBL3777493 SCHEMBL3777493 | C27H29N5O2 | 455.562 | 6 / 1 | 3.7 | Yes |
| 440068 | 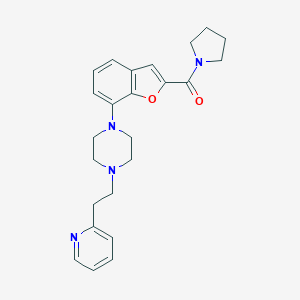 SCHEMBL931520 SCHEMBL931520 | C24H28N4O2 | 404.514 | 5 / 0 | 3.5 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218