

You can:
| Name | Glucagon receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | GCGR |
| Synonym | GL-R GGR GR glucagon receptor |
| Disease | Diabetes Type 2 diabetes Non-insulin dependent diabetes Obesity; Diabetes Type 1 diabetes [ Show all ] |
| Length | 477 |
| Amino acid sequence | MPPCQPQRPLLLLLLLLACQPQVPSAQVMDFLFEKWKLYGDQCHHNLSLLPPPTELVCNRTFDKYSCWPDTPANTTANISCPWYLPWHHKVQHRFVFKRCGPDGQWVRGPRGQPWRDASQCQMDGEEIEVQKEVAKMYSSFQVMYTVGYSLSLGALLLALAILGGLSKLHCTRNAIHANLFASFVLKASSVLVIDGLLRTRYSQKIGDDLSVSTWLSDGAVAGCRVAAVFMQYGIVANYCWLLVEGLYLHNLLGLATLPERSFFSLYLGIGWGAPMLFVVPWAVVKCLFENVQCWTSNDNMGFWWILRFPVFLAILINFFIFVRIVQLLVAKLRARQMHHTDYKFRLAKSTLTLIPLLGVHEVVFAFVTDEHAQGTLRSAKLFFDLFLSSFQGLLVAVLYCFLNKEVQSELRRRWHRWRLGKVLWEERNTSNHRASSSPGHGPPSKELQFGRGGGSQDSSAETPLAGGLPRLAESPF |
| UniProt | P47871 |
| Protein Data Bank | 5yqz, 5xez, 5ee7, 5xf1 |
| GPCR-HGmod model | P47871 |
| 3D structure model | This structure is from PDB ID 5yqz. |
| BioLiP | BL0379634,BL0379635, BL0402227, BL0379636,BL0379637, BL0343437 |
| Therapeutic Target Database | T60182, T87875 |
| ChEMBL | CHEMBL1985 |
| IUPHAR | 251 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 459237 | 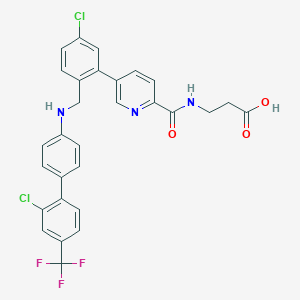 CHEMBL3680825 CHEMBL3680825 | C29H22Cl2F3N3O3 | 588.408 | 8 / 3 | 6.8 | No |
| 1704 | 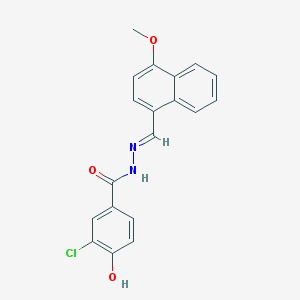 CHEMBL157064 CHEMBL157064 | C19H15ClN2O3 | 354.79 | 4 / 2 | 4.3 | Yes |
| 1959 | 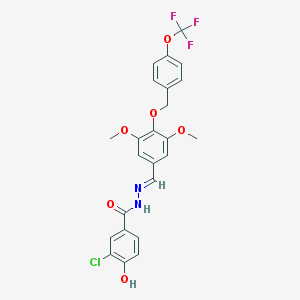 CHEMBL159093 CHEMBL159093 | C24H20ClF3N2O6 | 524.877 | 10 / 2 | 5.7 | No |
| 3421 | 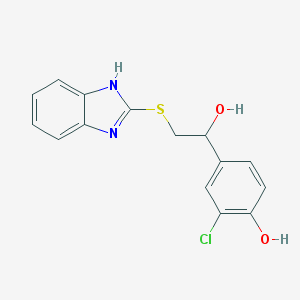 CHEMBL347544 CHEMBL347544 | C15H13ClN2O2S | 320.791 | 4 / 3 | 3.4 | Yes |
| 557383 | 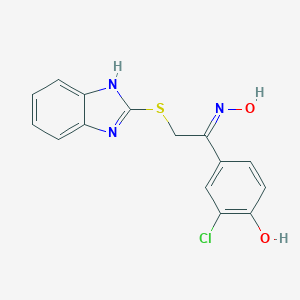 CID 135462249 CID 135462249 | C15H12ClN3O2S | 333.79 | 5 / 3 | 4.3 | Yes |
| 3825 | 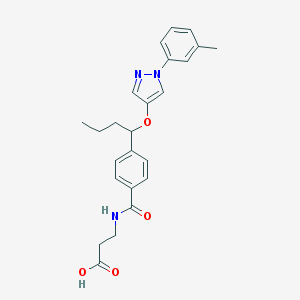 CHEMBL1933352 CHEMBL1933352 | C24H27N3O4 | 421.497 | 5 / 2 | 3.8 | Yes |
| 3954 | 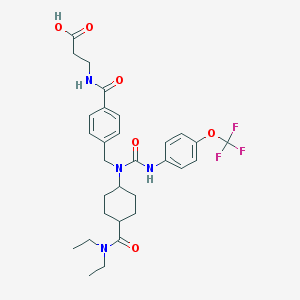 CHEMBL266715 CHEMBL266715 | C30H37F3N4O6 | 606.643 | 9 / 3 | 4.0 | No |
| 459269 | 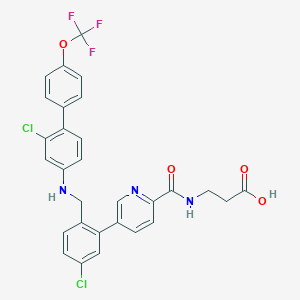 CHEMBL3675905 CHEMBL3675905 | C29H22Cl2F3N3O4 | 604.407 | 9 / 3 | 7.1 | No |
| 6115 | 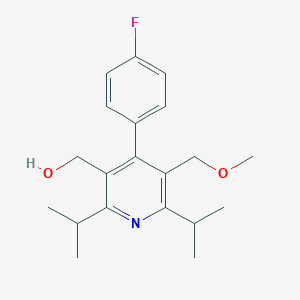 124864-27-3 124864-27-3 | C20H26FNO2 | 331.431 | 4 / 1 | 3.7 | Yes |
| 6809 | 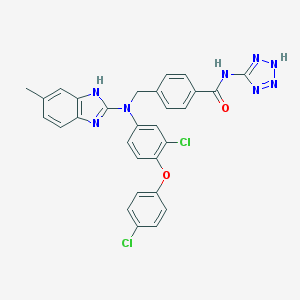 CHEMBL442855 CHEMBL442855 | C29H22Cl2N8O2 | 585.449 | 7 / 3 | 6.9 | No |
| 7496 | 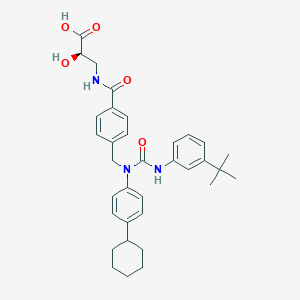 CHEMBL447742 CHEMBL447742 | C34H41N3O5 | 571.718 | 5 / 4 | 6.6 | No |
| 459292 | 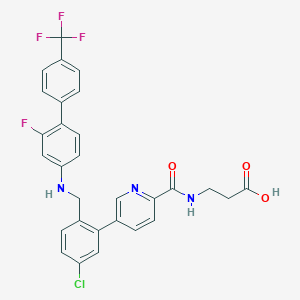 CHEMBL3675870 CHEMBL3675870 | C29H22ClF4N3O3 | 571.957 | 9 / 3 | 6.3 | No |
| 8543 | 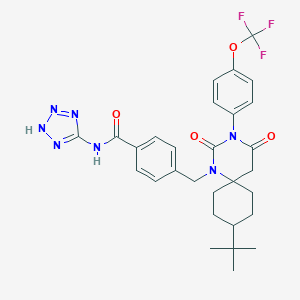 CHEMBL383350 CHEMBL383350 | C29H32F3N7O4 | 599.615 | 10 / 2 | 5.2 | No |
| 8952 | 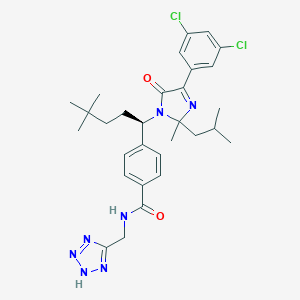 CHEMBL3656282 CHEMBL3656282 | C30H37Cl2N7O2 | 598.573 | 6 / 2 | 6.5 | No |
| 9694 | 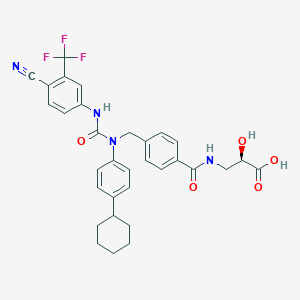 CHEMBL504245 CHEMBL504245 | C32H31F3N4O5 | 608.618 | 9 / 4 | 5.5 | No |
| 9725 | 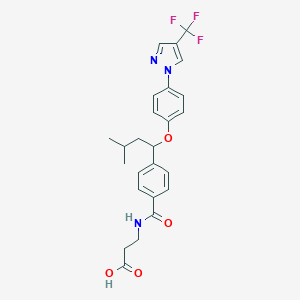 CHEMBL2381832 CHEMBL2381832 | C25H26F3N3O4 | 489.495 | 8 / 2 | 4.7 | Yes |
| 11086 | 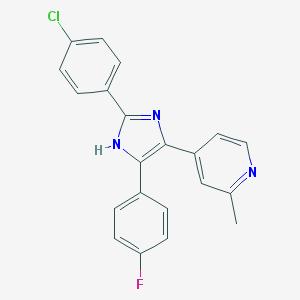 CHEMBL315836 CHEMBL315836 | C21H15ClFN3 | 363.82 | 3 / 1 | 5.1 | No |
| 11283 | 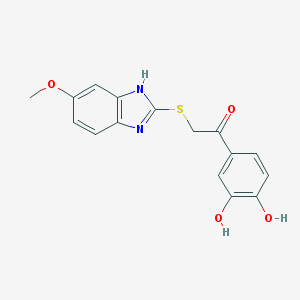 CHEMBL359176 CHEMBL359176 | C16H14N2O4S | 330.358 | 6 / 3 | 3.0 | Yes |
| 11363 | 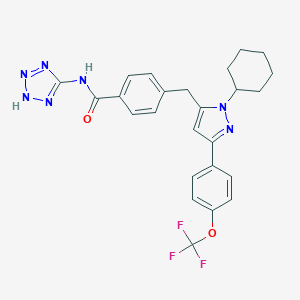 CHEMBL1643957 CHEMBL1643957 | C25H24F3N7O2 | 511.509 | 9 / 2 | 5.5 | No |
| 557633 | 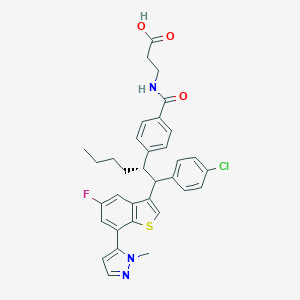 SCHEMBL1356029 SCHEMBL1356029 | C34H33ClFN3O3S | 618.164 | 6 / 2 | 8.0 | No |
| 548023 | 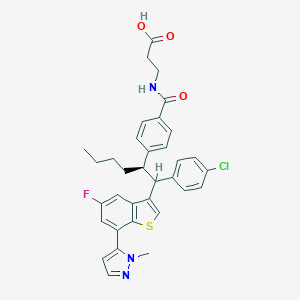 CHEMBL3944843 CHEMBL3944843 | C34H33ClFN3O3S | 618.164 | 6 / 2 | 8.0 | No |
| 13124 | 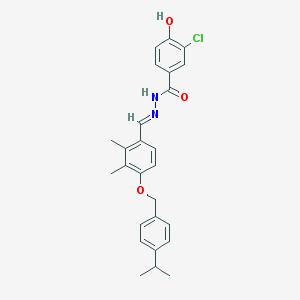 CHEMBL435445 CHEMBL435445 | C26H27ClN2O3 | 450.963 | 4 / 2 | 6.4 | No |
| 13342 | 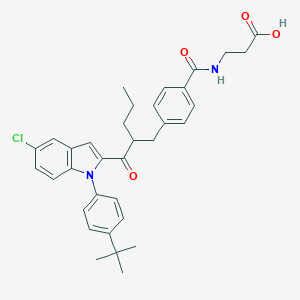 CHEMBL1922916 CHEMBL1922916 | C34H37ClN2O4 | 573.13 | 4 / 2 | 8.1 | No |
| 13738 | 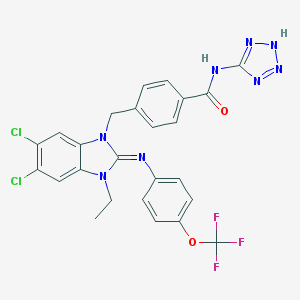 CHEMBL1922697 CHEMBL1922697 | C25H19Cl2F3N8O2 | 591.376 | 9 / 2 | 5.8 | No |
| 13789 | 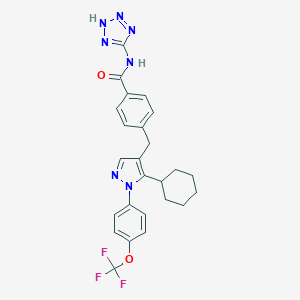 CHEMBL1644176 CHEMBL1644176 | C25H24F3N7O2 | 511.509 | 9 / 2 | 5.9 | No |
| 464563 | 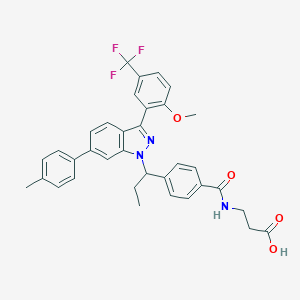 CHEMBL3616700 CHEMBL3616700 | C35H32F3N3O4 | 615.653 | 8 / 2 | 7.4 | No |
| 14336 | 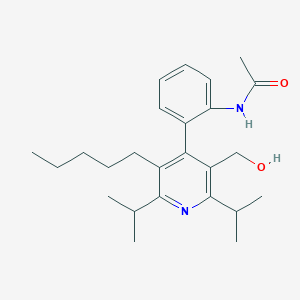 CHEMBL439915 CHEMBL439915 | C25H36N2O2 | 396.575 | 3 / 2 | 5.5 | No |
| 14572 | 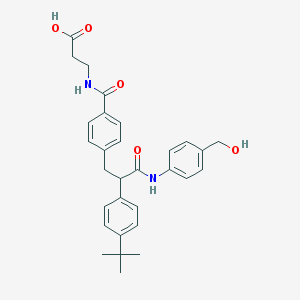 CHEMBL64955 CHEMBL64955 | C30H34N2O5 | 502.611 | 5 / 4 | 4.2 | No |
| 14818 | 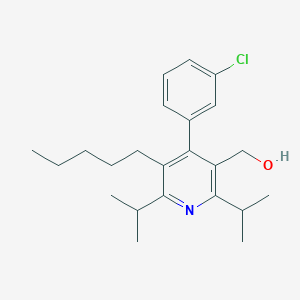 CHEMBL278361 CHEMBL278361 | C23H32ClNO | 373.965 | 2 / 1 | 7.0 | No |
| 15173 | 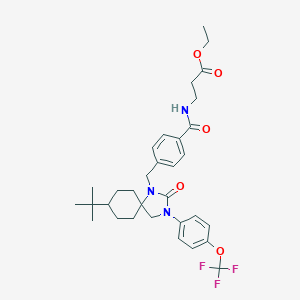 CHEMBL198231 CHEMBL198231 | C32H40F3N3O5 | 603.683 | 8 / 1 | 6.3 | No |
| 16041 | 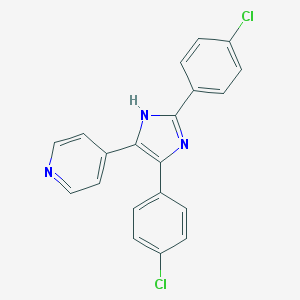 CHEMBL85009 CHEMBL85009 | C20H13Cl2N3 | 366.245 | 2 / 1 | 5.2 | No |
| 465048 | 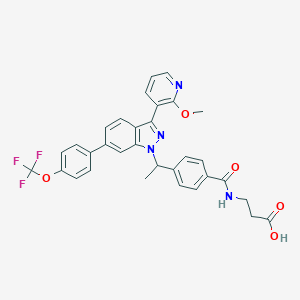 CHEMBL3616679 CHEMBL3616679 | C32H27F3N4O5 | 604.586 | 10 / 2 | 6.0 | No |
| 17842 | 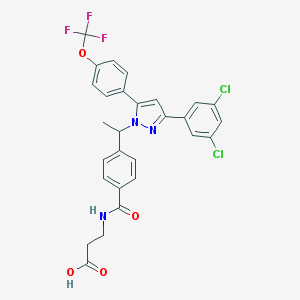 CHEMBL2159334 CHEMBL2159334 | C28H22Cl2F3N3O4 | 592.396 | 8 / 2 | 6.8 | No |
| 17959 | 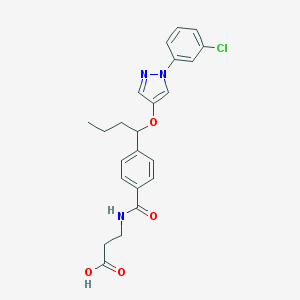 CHEMBL1933356 CHEMBL1933356 | C23H24ClN3O4 | 441.912 | 5 / 2 | 4.0 | Yes |
| 17991 | 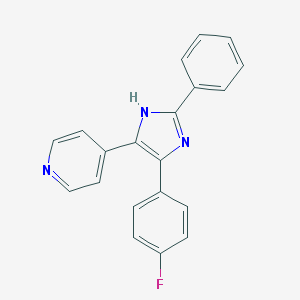 CHEMBL416169 CHEMBL416169 | C20H14FN3 | 315.351 | 3 / 1 | 4.1 | Yes |
| 522030 | 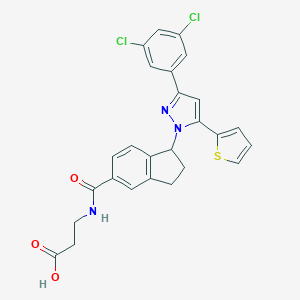 CHEMBL3800146 CHEMBL3800146 | C26H21Cl2N3O3S | 526.432 | 5 / 2 | 5.3 | No |
| 19436 | 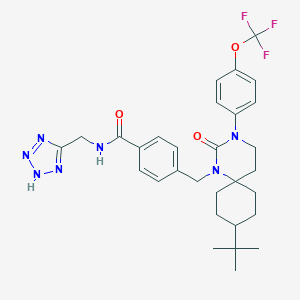 CHEMBL371235 CHEMBL371235 | C30H36F3N7O3 | 599.659 | 9 / 2 | 5.5 | No |
| 459390 | 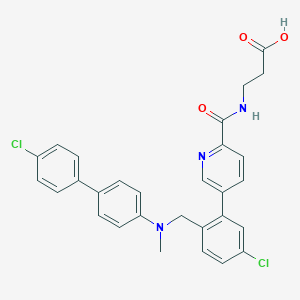 CHEMBL3675813 CHEMBL3675813 | C29H25Cl2N3O3 | 534.437 | 5 / 2 | 6.1 | No |
| 20021 | 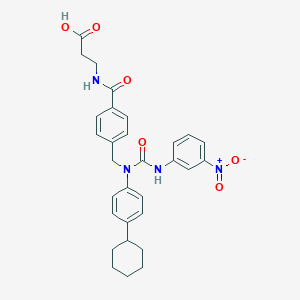 CHEMBL219968 CHEMBL219968 | C30H32N4O6 | 544.608 | 6 / 3 | 5.3 | No |
| 20097 | 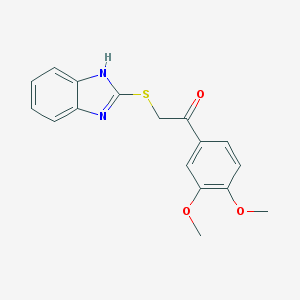 CHEMBL147177 CHEMBL147177 | C17H16N2O3S | 328.386 | 5 / 1 | 3.7 | Yes |
| 20613 | 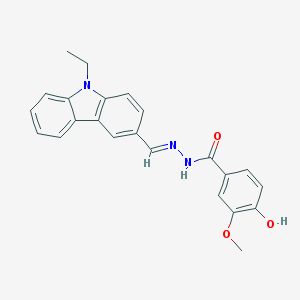 CHEMBL106577 CHEMBL106577 | C23H21N3O3 | 387.439 | 4 / 2 | 4.2 | Yes |
| 20968 | 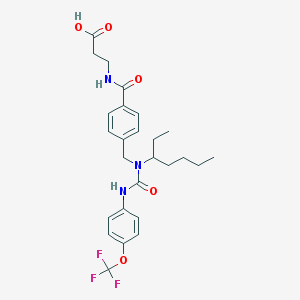 CHEMBL219308 CHEMBL219308 | C26H32F3N3O5 | 523.553 | 8 / 3 | 5.3 | No |
| 459406 | 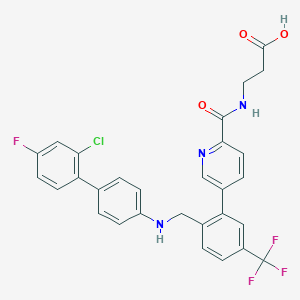 CHEMBL3675818 CHEMBL3675818 | C29H22ClF4N3O3 | 571.957 | 9 / 3 | 6.3 | No |
| 21921 | 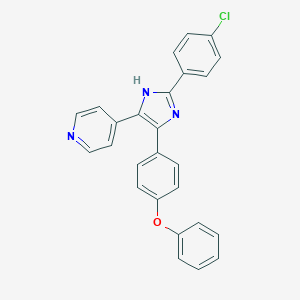 CHEMBL315613 CHEMBL315613 | C26H18ClN3O | 423.9 | 3 / 1 | 6.1 | No |
| 22261 | 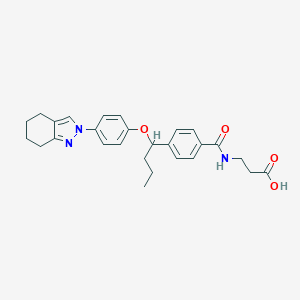 CHEMBL2381846 CHEMBL2381846 | C27H31N3O4 | 461.562 | 5 / 2 | 4.6 | Yes |
| 465735 | 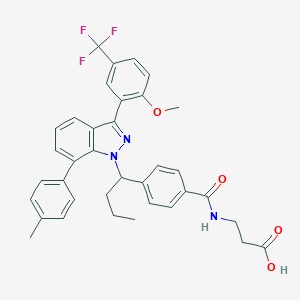 CHEMBL3616585 CHEMBL3616585 | C36H34F3N3O4 | 629.68 | 8 / 2 | 7.7 | No |
| 459413 | 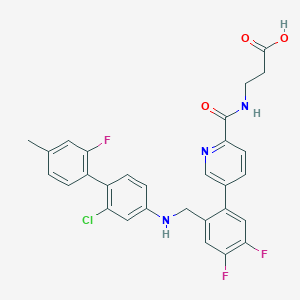 CHEMBL3675881 CHEMBL3675881 | C29H23ClF3N3O3 | 553.966 | 8 / 3 | 6.0 | No |
| 23523 | 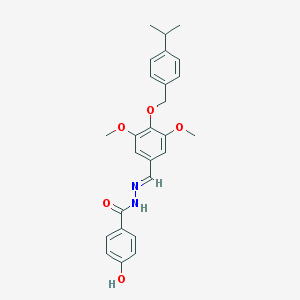 CHEMBL444343 CHEMBL444343 | C26H28N2O5 | 448.519 | 6 / 2 | 5.0 | Yes |
| 23885 | 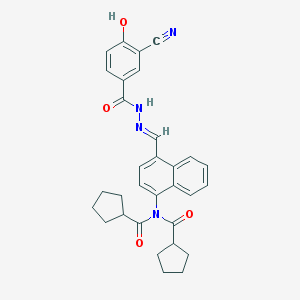 CHEMBL152477 CHEMBL152477 | C31H30N4O4 | 522.605 | 6 / 2 | 6.2 | No |
| 24745 | 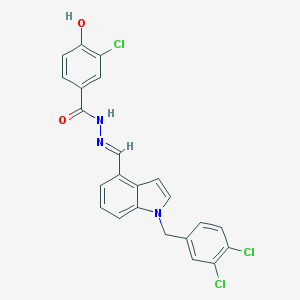 CHEMBL359126 CHEMBL359126 | C23H16Cl3N3O2 | 472.75 | 3 / 2 | 6.0 | No |
| 465981 | 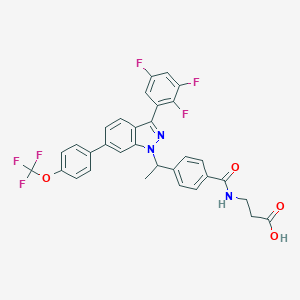 CHEMBL3616674 CHEMBL3616674 | C32H23F6N3O4 | 627.543 | 11 / 2 | 7.1 | No |
| 25536 | 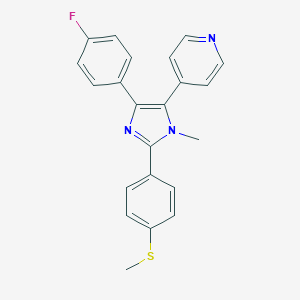 CHEMBL88045 CHEMBL88045 | C22H18FN3S | 375.465 | 4 / 0 | 4.5 | Yes |
| 25729 | 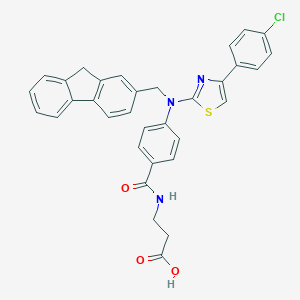 aminothiazole, 24 aminothiazole, 24 | C33H26ClN3O3S | 580.099 | 6 / 2 | 7.2 | No |
| 466116 | 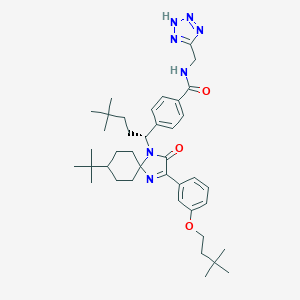 CHEMBL3617564 CHEMBL3617564 | C40H57N7O3 | 683.942 | 7 / 2 | 8.5 | No |
| 466194 | 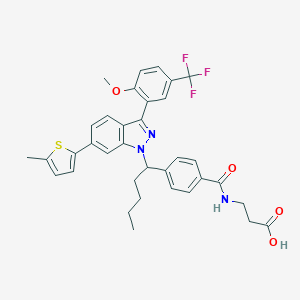 CHEMBL3616697 CHEMBL3616697 | C35H34F3N3O4S | 649.729 | 9 / 2 | 8.0 | No |
| 26961 | 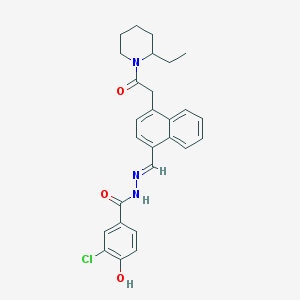 CHEMBL154400 CHEMBL154400 | C27H28ClN3O3 | 477.989 | 4 / 2 | 5.6 | No |
| 27788 | 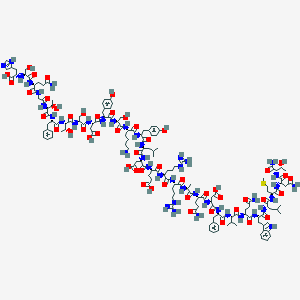 CID 49864554 CID 49864554 | C155H227N43O50S | 3524.83 | 56 / 55 | -16.3 | No |
| 27892 | 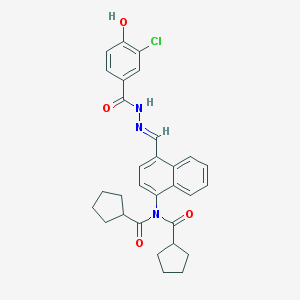 CHEMBL156724 CHEMBL156724 | C30H30ClN3O4 | 532.037 | 5 / 2 | 6.6 | No |
| 459462 | 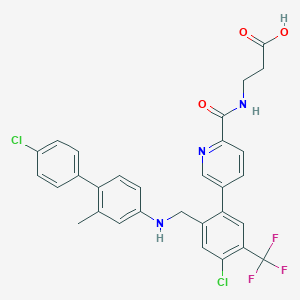 CHEMBL3675865 CHEMBL3675865 | C30H24Cl2F3N3O3 | 602.435 | 8 / 3 | 7.2 | No |
| 29017 | 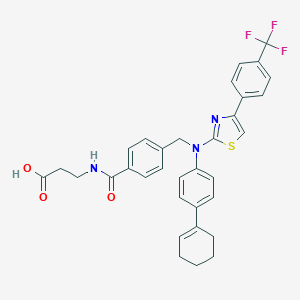 aminothiazole, 4 aminothiazole, 4 | C33H30F3N3O3S | 605.676 | 9 / 2 | 7.5 | No |
| 29162 | 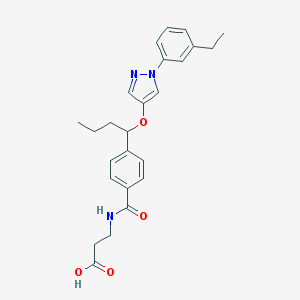 CHEMBL1933354 CHEMBL1933354 | C25H29N3O4 | 435.524 | 5 / 2 | 4.2 | Yes |
| 29230 | 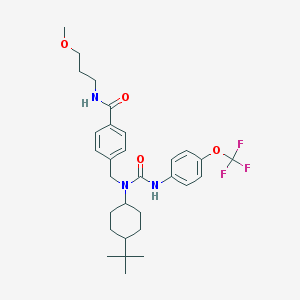 CHEMBL219450 CHEMBL219450 | C30H40F3N3O4 | 563.662 | 7 / 2 | 6.5 | No |
| 29358 | 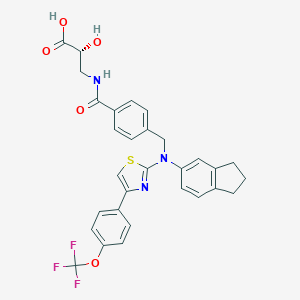 aminothiazole, 22 aminothiazole, 22 | C30H26F3N3O5S | 597.609 | 11 / 3 | 6.4 | No |
| 29794 | 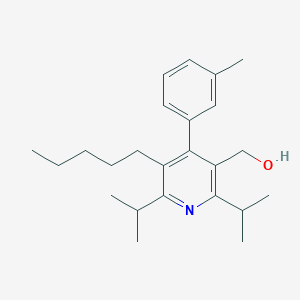 CHEMBL416559 CHEMBL416559 | C24H35NO | 353.55 | 2 / 1 | 6.7 | No |
| 459471 | 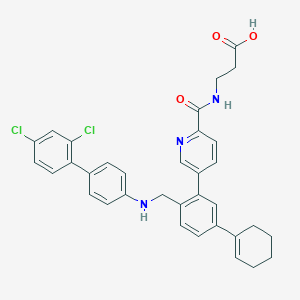 CHEMBL3680845 CHEMBL3680845 | C34H31Cl2N3O3 | 600.54 | 5 / 3 | 7.7 | No |
| 459479 | 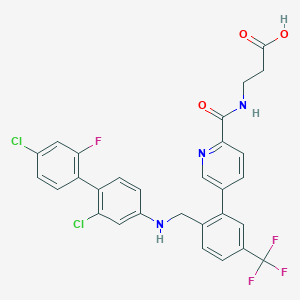 CHEMBL3675832 CHEMBL3675832 | C29H21Cl2F4N3O3 | 606.399 | 9 / 3 | 6.9 | No |
| 30607 | 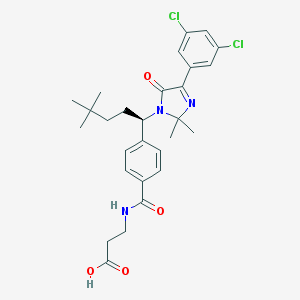 CHEMBL3656280 CHEMBL3656280 | C28H33Cl2N3O4 | 546.489 | 5 / 2 | 5.6 | No |
| 459484 | 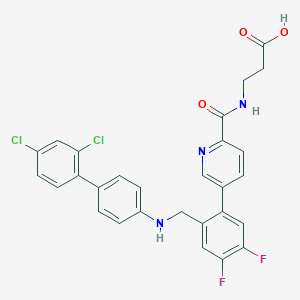 CHEMBL3675894 CHEMBL3675894 | C28H21Cl2F2N3O3 | 556.391 | 7 / 3 | 6.1 | No |
| 522477 | 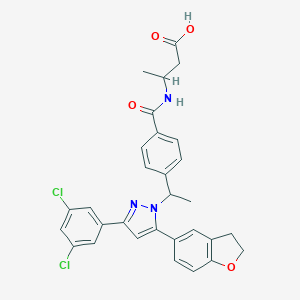 CHEMBL3797325 CHEMBL3797325 | C30H27Cl2N3O4 | 564.463 | 5 / 2 | 6.0 | No |
| 31166 | 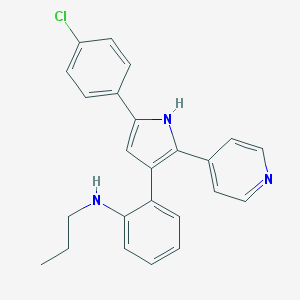 CHEMBL348555 CHEMBL348555 | C24H22ClN3 | 387.911 | 2 / 2 | 6.1 | No |
| 31349 | 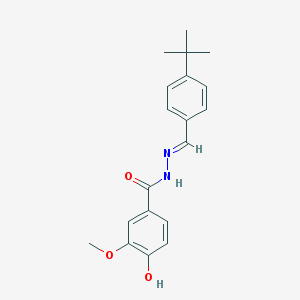 CHEMBL442389 CHEMBL442389 | C19H22N2O3 | 326.396 | 4 / 2 | 4.5 | Yes |
| 31689 | 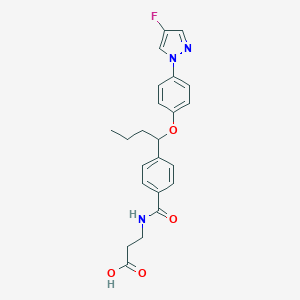 CHEMBL2381840 CHEMBL2381840 | C23H24FN3O4 | 425.46 | 6 / 2 | 3.5 | Yes |
| 459496 | 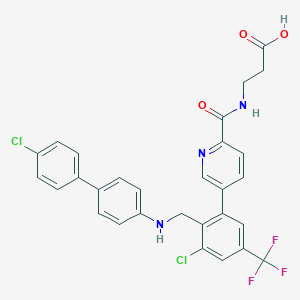 CHEMBL3675854 CHEMBL3675854 | C29H22Cl2F3N3O3 | 588.408 | 8 / 3 | 6.8 | No |
| 32341 | 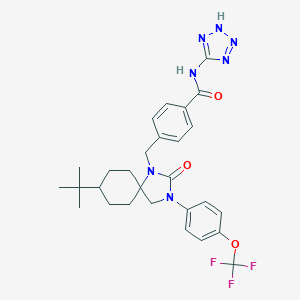 CHEMBL197851 CHEMBL197851 | C28H32F3N7O3 | 571.605 | 9 / 2 | 5.5 | No |
| 33184 | 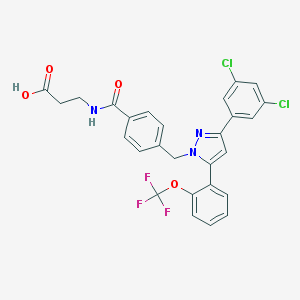 CHEMBL2159325 CHEMBL2159325 | C27H20Cl2F3N3O4 | 578.369 | 8 / 2 | 6.4 | No |
| 33456 | 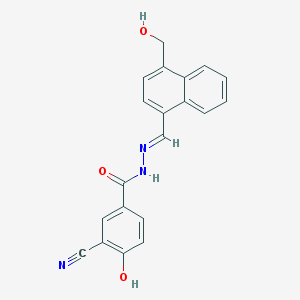 CHEMBL345589 CHEMBL345589 | C20H15N3O3 | 345.358 | 5 / 3 | 3.1 | Yes |
| 459515 | 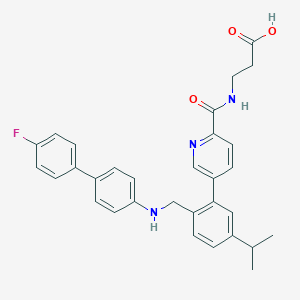 CHEMBL3680836 CHEMBL3680836 | C31H30FN3O3 | 511.597 | 6 / 3 | 5.9 | No |
| 467235 | 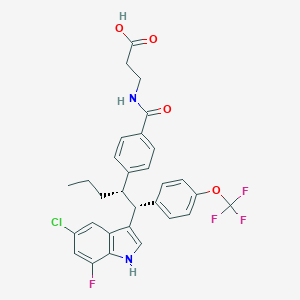 CHEMBL3617565 CHEMBL3617565 | C30H27ClF4N2O4 | 591.0 | 8 / 3 | 7.8 | No |
| 459529 | 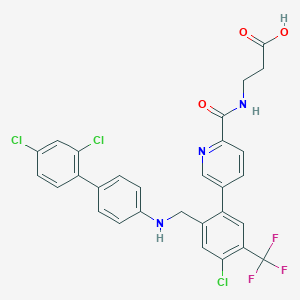 CHEMBL3675867 CHEMBL3675867 | C29H21Cl3F3N3O3 | 622.85 | 8 / 3 | 7.5 | No |
| 459530 | 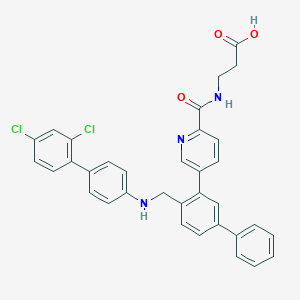 CHEMBL3680842 CHEMBL3680842 | C34H27Cl2N3O3 | 596.508 | 5 / 3 | 7.6 | No |
| 36023 | 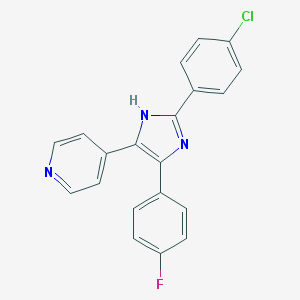 CHEMBL314238 CHEMBL314238 | C20H13ClFN3 | 349.793 | 3 / 1 | 4.7 | Yes |
| 467405 | 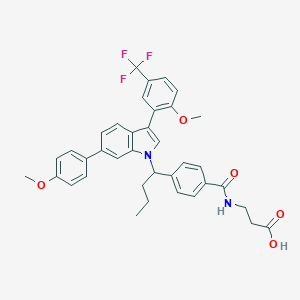 CHEMBL3616586 CHEMBL3616586 | C37H35F3N2O5 | 644.691 | 8 / 2 | 7.7 | No |
| 36640 | 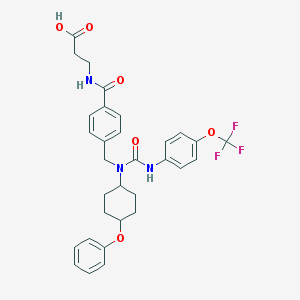 CHEMBL218833 CHEMBL218833 | C31H32F3N3O6 | 599.607 | 9 / 3 | 5.9 | No |
| 36674 | 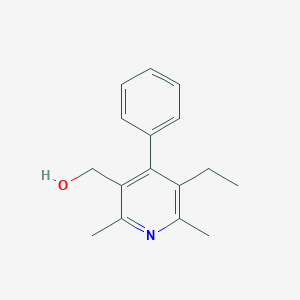 CHEMBL358249 CHEMBL358249 | C16H19NO | 241.334 | 2 / 1 | 3.2 | Yes |
| 36904 | 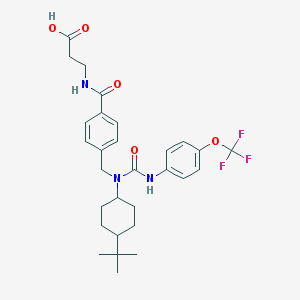 GRA EX-25 GRA EX-25 | C29H36F3N3O5 | 563.618 | 8 / 3 | 5.8 | No |
| 558404 | 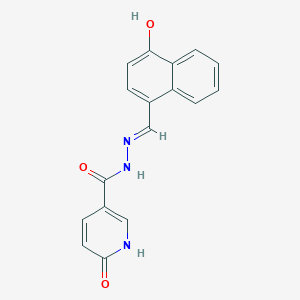 CHEMBL106950 CHEMBL106950 | C17H13N3O3 | 307.309 | 4 / 3 | 1.7 | Yes |
| 37599 | 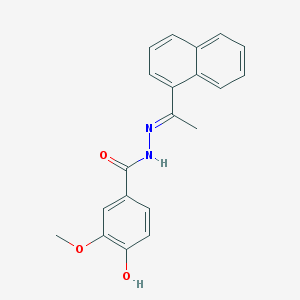 CHEMBL430509 CHEMBL430509 | C20H18N2O3 | 334.375 | 4 / 2 | 3.9 | Yes |
| 37716 | 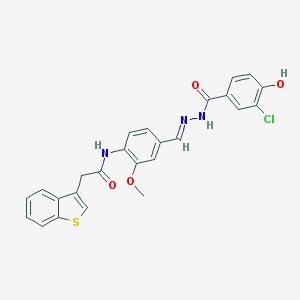 CHEMBL152348 CHEMBL152348 | C25H20ClN3O4S | 493.962 | 6 / 3 | 4.9 | Yes |
| 534077 | 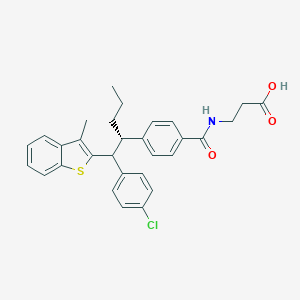 CHEMBL3943204 CHEMBL3943204 | C30H30ClNO3S | 520.084 | 4 / 2 | 7.8 | No |
| 522665 | 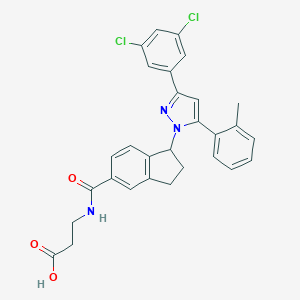 CHEMBL3799499 CHEMBL3799499 | C29H25Cl2N3O3 | 534.437 | 4 / 2 | 6.0 | No |
| 38844 | 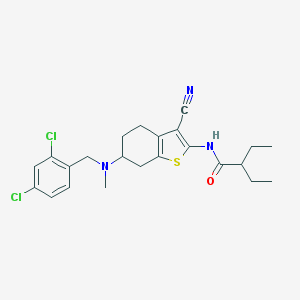 CHEMBL179181 CHEMBL179181 | C23H27Cl2N3OS | 464.449 | 4 / 1 | 6.7 | No |
| 39030 | 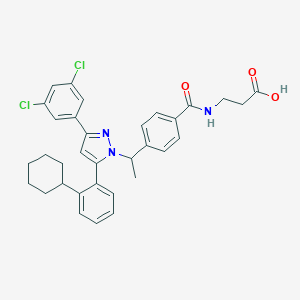 CHEMBL2159344 CHEMBL2159344 | C33H33Cl2N3O3 | 590.545 | 4 / 2 | 8.1 | No |
| 39428 | 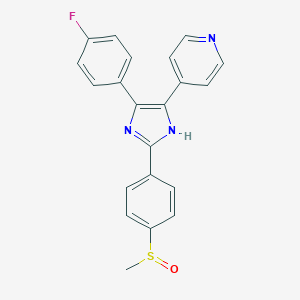 152121-47-6 152121-47-6 | C21H16FN3OS | 377.437 | 5 / 1 | 3.2 | Yes |
| 40060 | 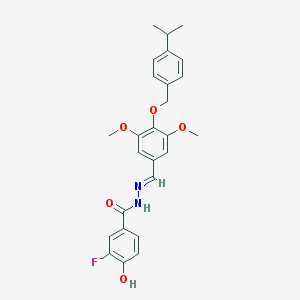 CHEMBL159071 CHEMBL159071 | C26H27FN2O5 | 466.509 | 7 / 2 | 5.1 | No |
| 459576 | 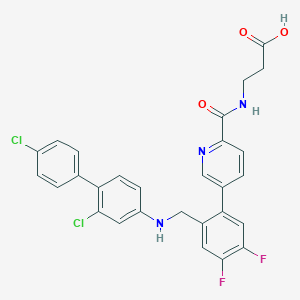 CHEMBL3675879 CHEMBL3675879 | C28H21Cl2F2N3O3 | 556.391 | 7 / 3 | 6.1 | No |
| 41087 | 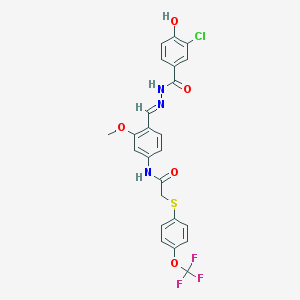 CHEMBL346694 CHEMBL346694 | C24H19ClF3N3O5S | 553.937 | 10 / 3 | 5.7 | No |
| 41258 | 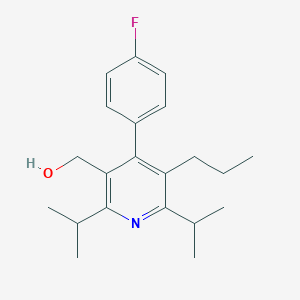 CHEMBL140514 CHEMBL140514 | C21H28FNO | 329.459 | 3 / 1 | 5.4 | No |
| 534090 | 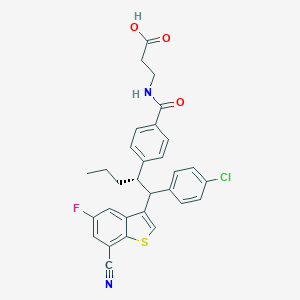 CHEMBL3939861 CHEMBL3939861 | C30H26ClFN2O3S | 549.057 | 6 / 2 | 7.2 | No |
| 42607 | 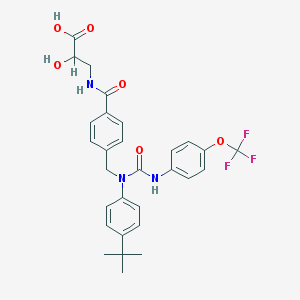 CHEMBL486650 CHEMBL486650 | C29H30F3N3O6 | 573.569 | 9 / 4 | 5.2 | No |
| 558593 | 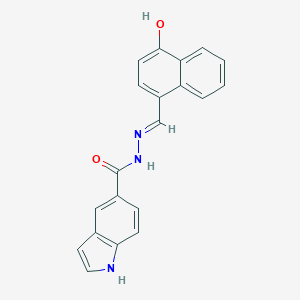 CHEMBL107480 CHEMBL107480 | C20H15N3O2 | 329.359 | 3 / 3 | 3.9 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218