

You can:
| Name | Melatonin receptor type 1C |
|---|---|
| Species | Xenopus laevis (African clawed frog) |
| Gene | mtnr1c |
| Synonym | Mel-1C-R Mel1c (alpha) receptor Mel1c receptor |
| Disease | N/A for non-human GPCRs |
| Length | 420 |
| Amino acid sequence | MMEVNSTCLDCRTPGTIRTEQDAQDSASQGLTSALAVVLIFTIVVDVLGNILVILSVLRNKKLQNAGNLFVVSLSIADLVVAVYPYPVILIAIFQNGWTLGNIHCQISGFLMGLSVIGSVFNITAIAINRYCYICHSLRYDKLYNQRSTWCYLGLTWILTIIAIVPNFFVGSLQYDPRIFSCTFAQTVSSSYTITVVVVHFIVPLSVVTFCYLRIWVLVIQVKHRVRQDFKQKLTQTDLRNFLTMFVVFVLFAVCWAPLNFIGLAVAINPFHVAPKIPEWLFVLSYFMAYFNSCLNAVIYGVLNQNFRKEYKRILMSLLTPRLLFLDTSRGGTEGLKSKPSPAVTNNNQADMLGEARSLWLSRRNGAKMVIIIRPRKAQIAIIHQIFWPQSSWATCRQDTKITGEEDGCRELCKDGISQR |
| UniProt | P49219 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL5495 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 16482 | 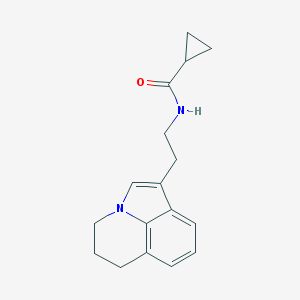 CHEMBL244471 CHEMBL244471 | C17H20N2O | 268.36 | 1 / 1 | 2.4 | Yes |
| 555577 | 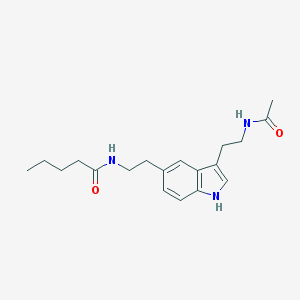 5-MeOT-N-pentanyl 5-MeOT-N-pentanyl | C19H27N3O2 | 329.444 | 2 / 3 | 2.6 | Yes |
| 40095 | 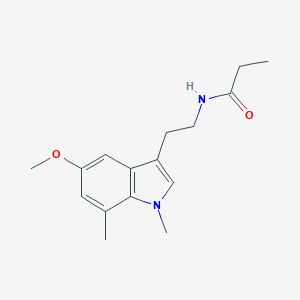 CHEMBL233516 CHEMBL233516 | C16H22N2O2 | 274.364 | 2 / 1 | 2.4 | Yes |
| 51776 | 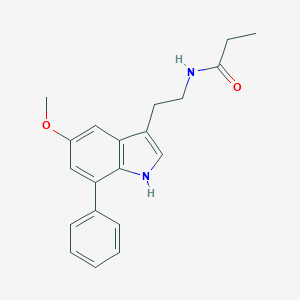 CHEMBL233517 CHEMBL233517 | C20H22N2O2 | 322.408 | 2 / 2 | 3.7 | Yes |
| 55349 | 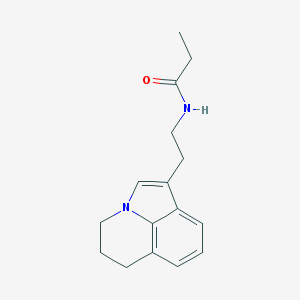 CHEMBL244957 CHEMBL244957 | C16H20N2O | 256.349 | 1 / 1 | 2.4 | Yes |
| 67089 | 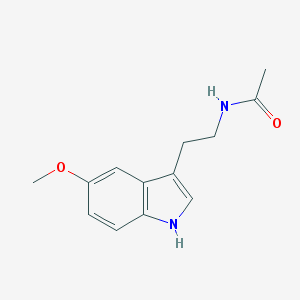 Melatonin Melatonin | C13H16N2O2 | 232.283 | 2 / 2 | 0.8 | Yes |
| 77929 | 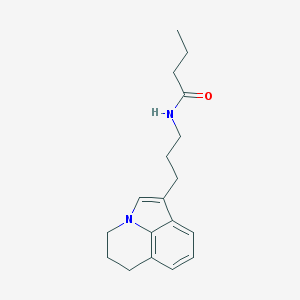 CHEMBL244472 CHEMBL244472 | C18H24N2O | 284.403 | 1 / 1 | 3.1 | Yes |
| 86095 | 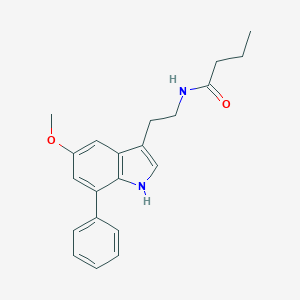 CHEMBL394408 CHEMBL394408 | C21H24N2O2 | 336.435 | 2 / 2 | 4.0 | Yes |
| 104555 | 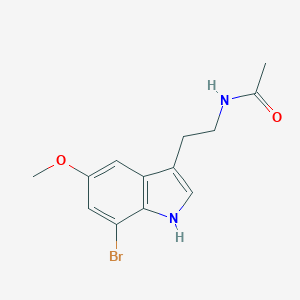 7-bromomelatonin 7-bromomelatonin | C13H15BrN2O2 | 311.179 | 2 / 2 | 2.3 | Yes |
| 555904 | 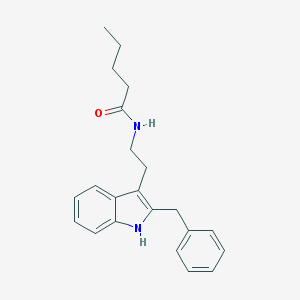 DH 97 DH 97 | C22H26N2O | 334.463 | 1 / 2 | 4.9 | Yes |
| 124988 | 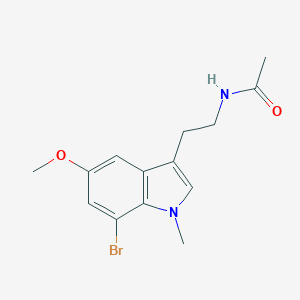 CHEMBL233309 CHEMBL233309 | C14H17BrN2O2 | 325.206 | 2 / 1 | 2.2 | Yes |
| 139664 | 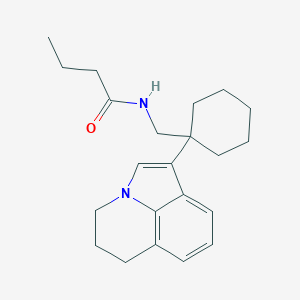 CHEMBL243003 CHEMBL243003 | C22H30N2O | 338.495 | 1 / 1 | 4.6 | Yes |
| 165860 | 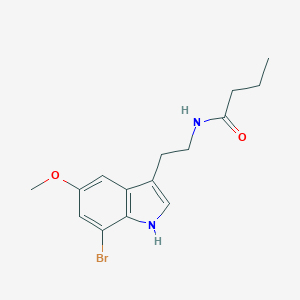 CHEMBL233308 CHEMBL233308 | C15H19BrN2O2 | 339.233 | 2 / 2 | 3.1 | Yes |
| 173230 | 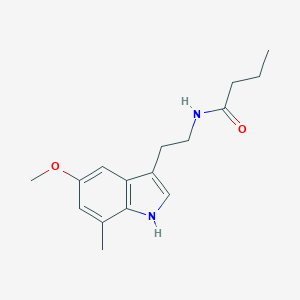 CHEMBL391350 CHEMBL391350 | C16H22N2O2 | 274.364 | 2 / 2 | 2.8 | Yes |
| 180487 | 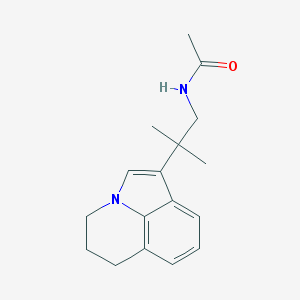 CHEMBL396991 CHEMBL396991 | C17H22N2O | 270.376 | 1 / 1 | 2.8 | Yes |
| 556208 | 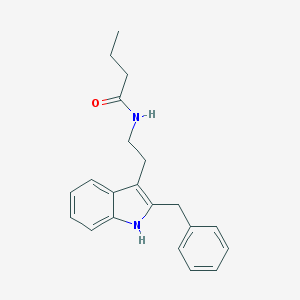 Luzindole,N-butanoyl Luzindole,N-butanoyl | C21H24N2O | 320.436 | 1 / 2 | 4.4 | Yes |
| 186836 | 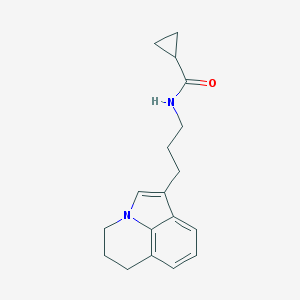 CHEMBL242573 CHEMBL242573 | C18H22N2O | 282.387 | 1 / 1 | 2.7 | Yes |
| 556304 | 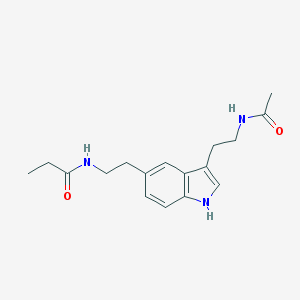 5-MeOT-N-propionyl 5-MeOT-N-propionyl | C17H23N3O2 | 301.39 | 2 / 3 | 1.7 | Yes |
| 227807 | 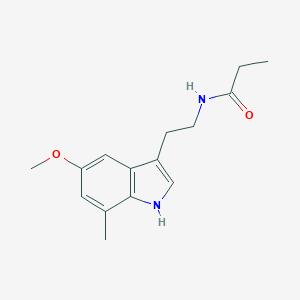 CHEMBL395221 CHEMBL395221 | C15H20N2O2 | 260.337 | 2 / 2 | 2.4 | Yes |
| 556423 | 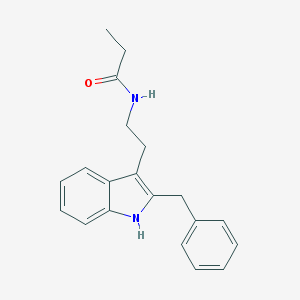 N-0889 N-0889 | C20H22N2O | 306.409 | 1 / 2 | 4.1 | Yes |
| 556442 | 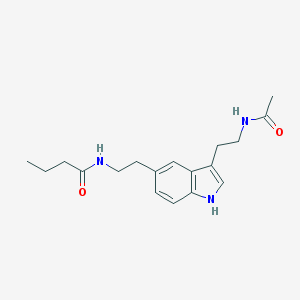 5-MeOT-N-butanyl 5-MeOT-N-butanyl | C18H25N3O2 | 315.417 | 2 / 3 | 2.0 | Yes |
| 279317 | 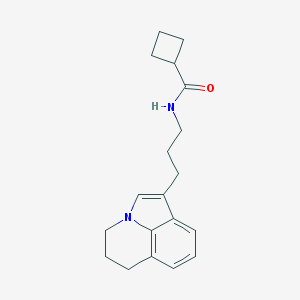 CHEMBL242786 CHEMBL242786 | C19H24N2O | 296.414 | 1 / 1 | 3.3 | Yes |
| 286060 | 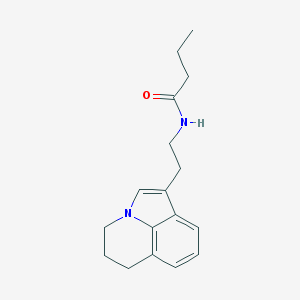 CHEMBL396501 CHEMBL396501 | C17H22N2O | 270.376 | 1 / 1 | 2.7 | Yes |
| 287214 | 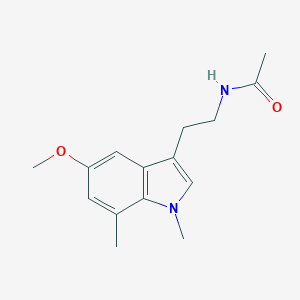 CHEMBL392421 CHEMBL392421 | C15H20N2O2 | 260.337 | 2 / 1 | 1.9 | Yes |
| 556606 | 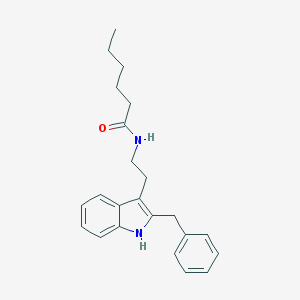 Luzindole,N-hexanoyl Luzindole,N-hexanoyl | C23H28N2O | 348.49 | 1 / 2 | 5.5 | No |
| 297450 | 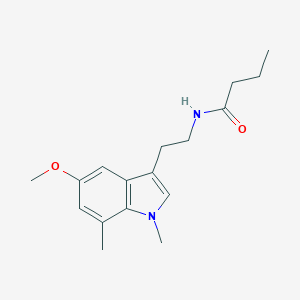 CHEMBL392422 CHEMBL392422 | C17H24N2O2 | 288.391 | 2 / 1 | 2.7 | Yes |
| 303513 | 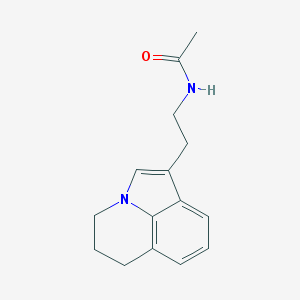 CHEMBL242806 CHEMBL242806 | C15H18N2O | 242.322 | 1 / 1 | 1.9 | Yes |
| 556693 | 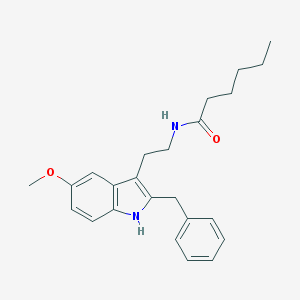 Luzindole,5-MeOT-N-hexanyl Luzindole,5-MeOT-N-hexanyl | C24H30N2O2 | 378.516 | 2 / 2 | 5.5 | No |
| 319020 | 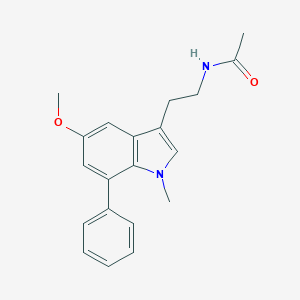 CHEMBL233740 CHEMBL233740 | C20H22N2O2 | 322.408 | 2 / 1 | 3.2 | Yes |
| 328296 | 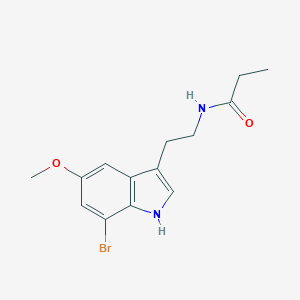 CHEMBL233729 CHEMBL233729 | C14H17BrN2O2 | 325.206 | 2 / 2 | 2.8 | Yes |
| 337422 | 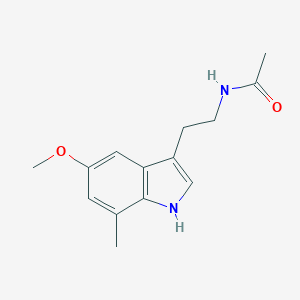 CHEMBL231872 CHEMBL231872 | C14H18N2O2 | 246.31 | 2 / 2 | 2.0 | Yes |
| 374541 | 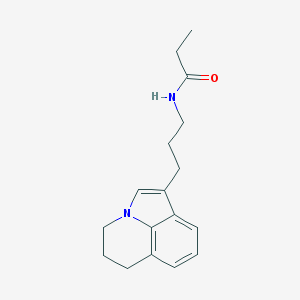 CHEMBL244263 CHEMBL244263 | C17H22N2O | 270.376 | 1 / 1 | 2.7 | Yes |
| 557051 | 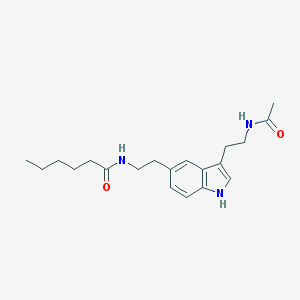 5-MeOT-N-hexanyl 5-MeOT-N-hexanyl | C20H29N3O2 | 343.471 | 2 / 3 | 3.1 | Yes |
| 383593 | 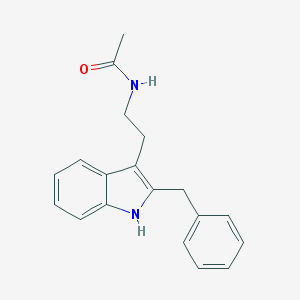 luzindole luzindole | C19H20N2O | 292.382 | 1 / 2 | 3.6 | Yes |
| 405861 | 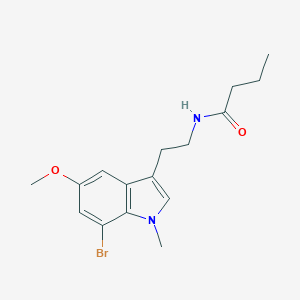 CHEMBL394406 CHEMBL394406 | C16H21BrN2O2 | 353.26 | 2 / 1 | 3.1 | Yes |
| 408815 | 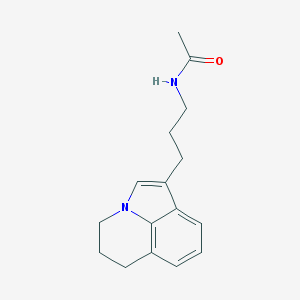 CHEMBL241928 CHEMBL241928 | C16H20N2O | 256.349 | 1 / 1 | 2.3 | Yes |
| 410780 | 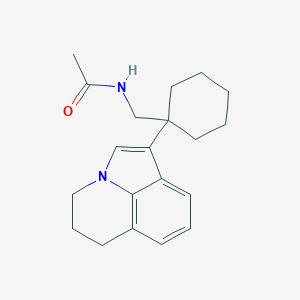 CHEMBL243002 CHEMBL243002 | C20H26N2O | 310.441 | 1 / 1 | 3.8 | Yes |
| 437295 | 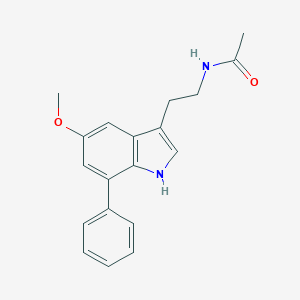 CHEMBL233311 CHEMBL233311 | C19H20N2O2 | 308.381 | 2 / 2 | 3.2 | Yes |
| 439158 | 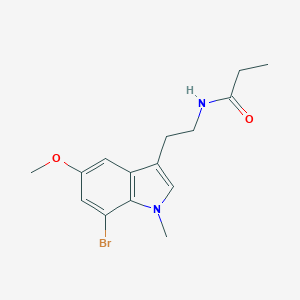 CHEMBL233310 CHEMBL233310 | C15H19BrN2O2 | 339.233 | 2 / 1 | 2.7 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218