

You can:
| Name | P2Y purinoceptor 1 |
|---|---|
| Species | Meleagris gallopavo (Wild turkey) |
| Gene | P2RY1 |
| Synonym | 6H1 orphan receptor ADP receptor P2Y1 Purinergic receptor |
| Disease | N/A for non-human GPCRs |
| Length | 362 |
| Amino acid sequence | MTEALISAALNGTQPELLAGGWAAGNASTKCSLTKTGFQFYYLPTVYILVFITGFLGNSVAIWMFVFHMRPWSGISVYMFNLALADFLYVLTLPALIFYYFNKTDWIFGDVMCKLQRFIFHVNLYGSILFLTCISVHRYTGVVHPLKSLGRLKKKNAVYVSSLVWALVVAVIAPILFYSGTGVRRNKTITCYDTTADEYLRSYFVYSMCTTVFMFCIPFIVILGCYGLIVKALIYKDLDNSPLRRKSIYLVIIVLTVFAVSYLPFHVMKTLNLRARLDFQTPQMCAFNDKVYATYQVTRGLASLNSCVDPILYFLAGDTFRRRLSRATRKSSRRSEPNVQSKSEEMTLNILTEYKQNGDTSL |
| UniProt | P49652 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL5720 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 792 | 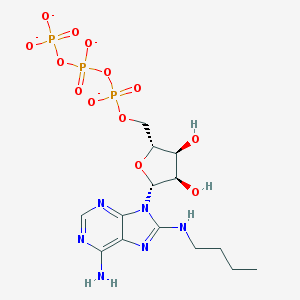 CHEMBL2364563 CHEMBL2364563 | C14H21N6O13P3-4 | 574.272 | 18 / 4 | -4.5 | No |
| 3668 | 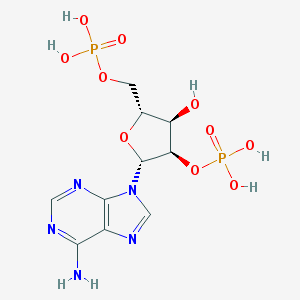 adenosine-2'-5'-diphosphate adenosine-2'-5'-diphosphate | C10H15N5O10P2 | 427.203 | 14 / 6 | -4.6 | No |
| 519714 | 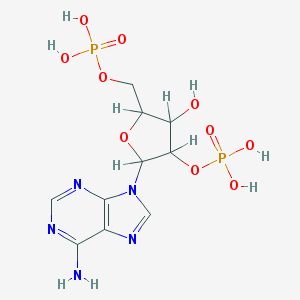 AC1L1CRI AC1L1CRI | C10H15N5O10P2 | 427.203 | 14 / 6 | -4.6 | No |
| 557505 | 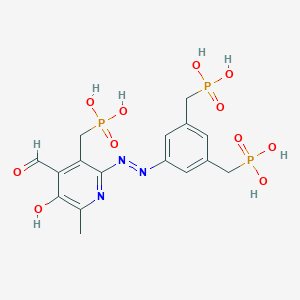 CHEMBL118007 CHEMBL118007 | C16H20N3O11P3 | 523.267 | 14 / 7 | -3.1 | No |
| 10011 | 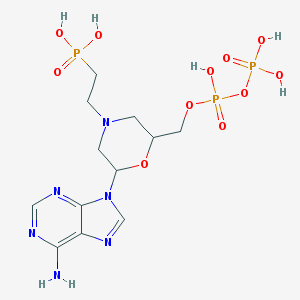 CHEMBL355325 CHEMBL355325 | C12H21N6O11P3 | 518.252 | 16 / 6 | -7.5 | No |
| 14333 | 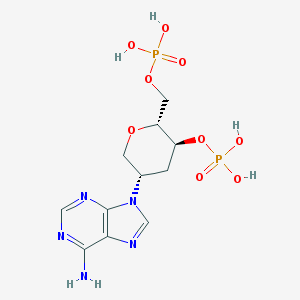 CHEMBL607338 CHEMBL607338 | C11H17N5O9P2 | 425.231 | 13 / 5 | -3.8 | No |
| 14334 | 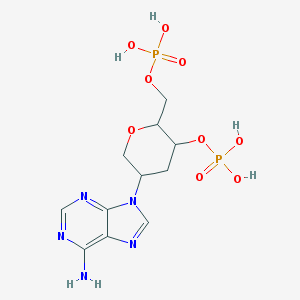 CHEMBL416252 CHEMBL416252 | C11H17N5O9P2 | 425.231 | 13 / 5 | -3.8 | No |
| 15080 | 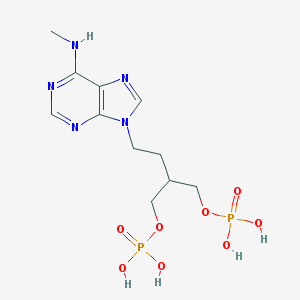 CHEMBL432028 CHEMBL432028 | C11H19N5O8P2 | 411.248 | 12 / 5 | -2.4 | No |
| 557736 | 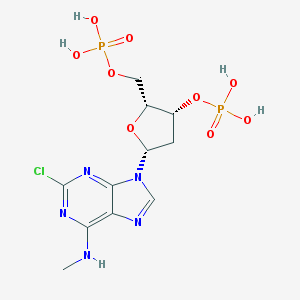 CHEMBL164811 CHEMBL164811 | C11H16ClN5O9P2 | 459.673 | 13 / 5 | -1.3 | No |
| 16322 | 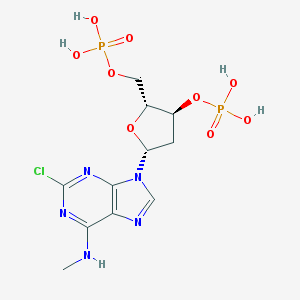 CHEMBL403456 CHEMBL403456 | C11H16ClN5O9P2 | 459.673 | 13 / 5 | -1.3 | No |
| 557737 | 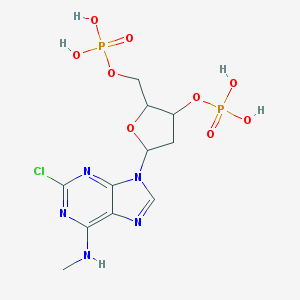 CHEMBL288798 CHEMBL288798 | C11H16ClN5O9P2 | 459.673 | 13 / 5 | -1.3 | No |
| 557958 | 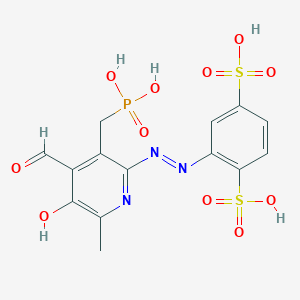 CHEMBL119416 CHEMBL119416 | C14H14N3O11PS2 | 495.37 | 14 / 5 | -1.5 | No |
| 558143 | 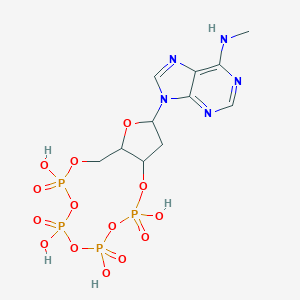 CHEMBL296960 CHEMBL296960 | C11H17N5O14P4 | 567.173 | 18 / 5 | -4.2 | No |
| 28869 | 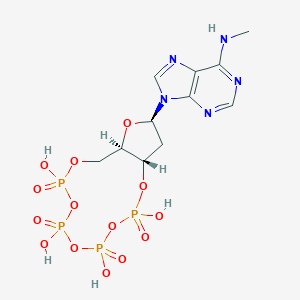 CHEMBL3144305 CHEMBL3144305 | C11H17N5O14P4 | 567.173 | 18 / 5 | -4.2 | No |
| 558213 | 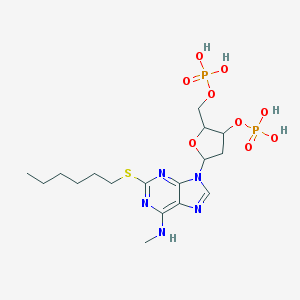 CHEMBL42627 CHEMBL42627 | C17H29N5O9P2S | 541.453 | 14 / 5 | 0.2 | No |
| 30523 | 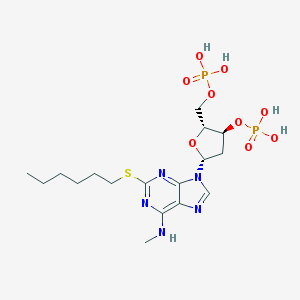 CHEMBL3144170 CHEMBL3144170 | C17H29N5O9P2S | 541.453 | 14 / 5 | 0.2 | No |
| 31737 | 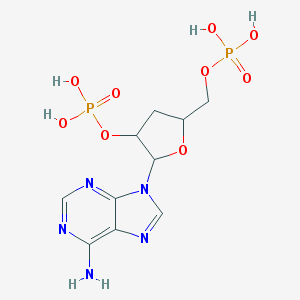 CHEMBL55804 CHEMBL55804 | C10H15N5O9P2 | 411.204 | 13 / 5 | -3.4 | No |
| 37597 | 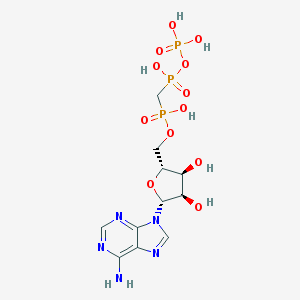 alpha,beta-Methylene ATP alpha,beta-Methylene ATP | C11H18N5O12P3 | 505.209 | 16 / 7 | -5.9 | No |
| 38812 | 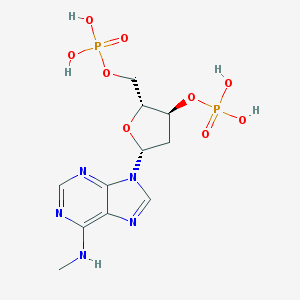 CHEMBL129841 CHEMBL129841 | C11H17N5O9P2 | 425.231 | 13 / 5 | -2.9 | No |
| 41708 | 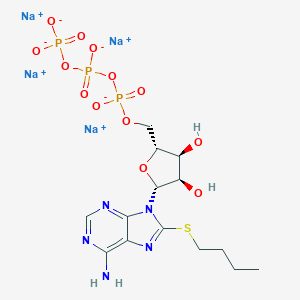 CHEMBL2364564 CHEMBL2364564 | C14H20N5Na4O13P3S | 683.276 | 18 / 3 | N/A | No |
| 558614 | 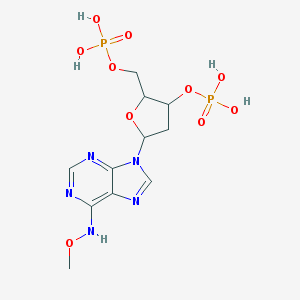 CHEMBL44317 CHEMBL44317 | C11H17N5O10P2 | 441.23 | 14 / 5 | -2.9 | No |
| 43416 | 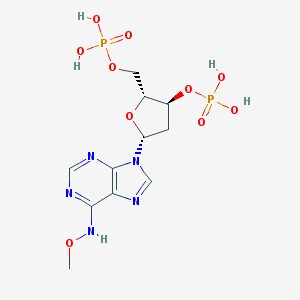 CHEMBL3144182 CHEMBL3144182 | C11H17N5O10P2 | 441.23 | 14 / 5 | -2.9 | No |
| 44919 | 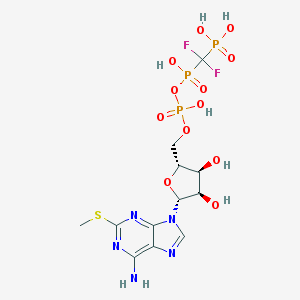 CHEMBL1089560 CHEMBL1089560 | C12H18F2N5O12P3S | 587.277 | 19 / 7 | -4.2 | No |
| 53090 | 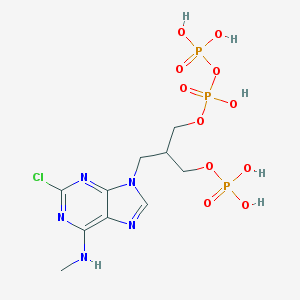 CHEMBL319906 CHEMBL319906 | C10H17ClN5O11P3 | 511.641 | 15 / 6 | -3.1 | No |
| 558967 | 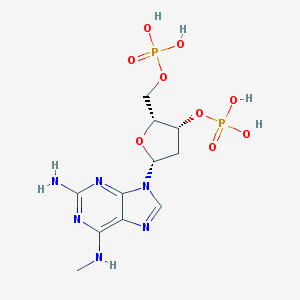 CHEMBL353089 CHEMBL353089 | C11H18N6O9P2 | 440.246 | 14 / 6 | -2.6 | No |
| 54632 | 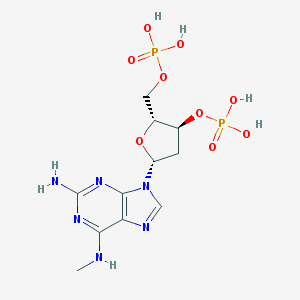 CHEMBL2368315 CHEMBL2368315 | C11H18N6O9P2 | 440.246 | 14 / 6 | -2.6 | No |
| 558968 | 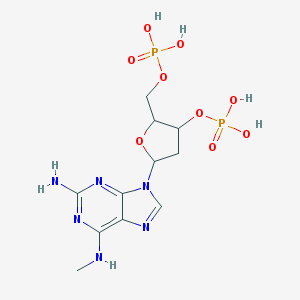 CHEMBL43507 CHEMBL43507 | C11H18N6O9P2 | 440.246 | 14 / 6 | -2.6 | No |
| 63613 | 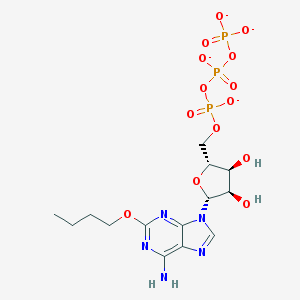 CHEMBL2364565 CHEMBL2364565 | C14H20N5O14P3-4 | 575.256 | 18 / 3 | -4.5 | No |
| 559328 | 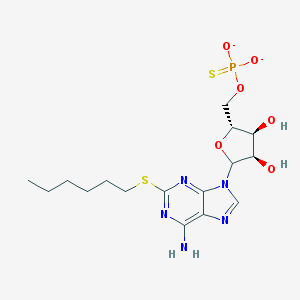 CHEMBL607771 CHEMBL607771 | C16H24N5O6PS2-2 | 477.491 | 12 / 3 | 1.2 | No |
| 68099 | 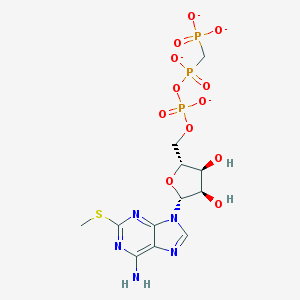 BDBM50268012 BDBM50268012 | C12H16N5O12P3S-4 | 547.264 | 17 / 3 | -5.3 | No |
| 68102 | 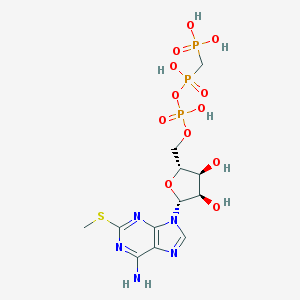 CHEMBL590527 CHEMBL590527 | C12H20N5O12P3S | 551.296 | 17 / 7 | -5.0 | No |
| 71159 | 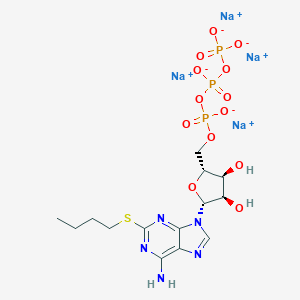 CHEMBL2364567 CHEMBL2364567 | C14H20N5Na4O13P3S | 683.276 | 18 / 3 | N/A | No |
| 72628 | 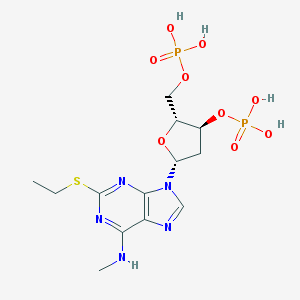 CHEMBL3144309 CHEMBL3144309 | C13H21N5O9P2S | 485.345 | 14 / 5 | -1.0 | No |
| 559550 | 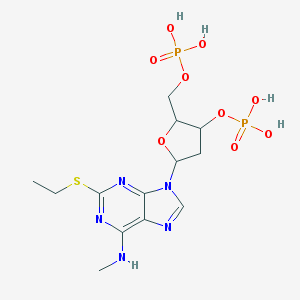 CHEMBL295819 CHEMBL295819 | C13H21N5O9P2S | 485.345 | 14 / 5 | -1.0 | No |
| 75225 | 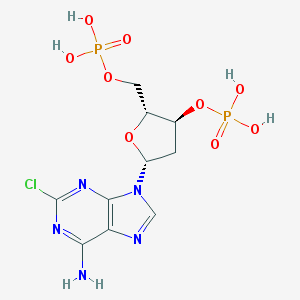 CHEMBL1094760 CHEMBL1094760 | C10H14ClN5O9P2 | 445.646 | 13 / 5 | -2.0 | No |
| 520033 | 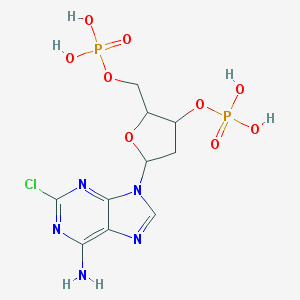 CHEMBL44408 CHEMBL44408 | C10H14ClN5O9P2 | 445.646 | 13 / 5 | -2.0 | No |
| 78876 | 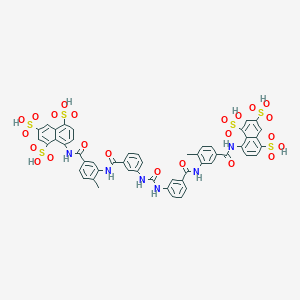 suramin suramin | C51H40N6O23S6 | 1297.26 | 23 / 12 | 1.5 | No |
| 82712 | 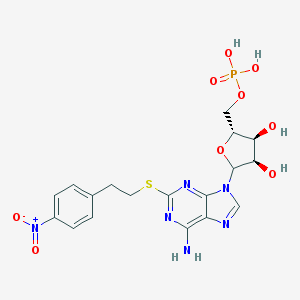 CHEMBL610985 CHEMBL610985 | C18H21N6O9PS | 528.433 | 14 / 5 | -0.9 | No |
| 559865 | 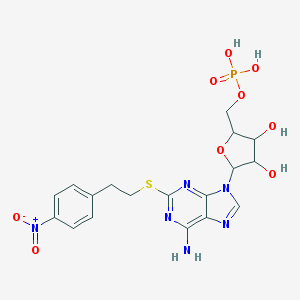 CHEMBL610985 CHEMBL610985 | C18H21N6O9PS | 528.433 | 14 / 5 | -0.9 | No |
| 84327 | 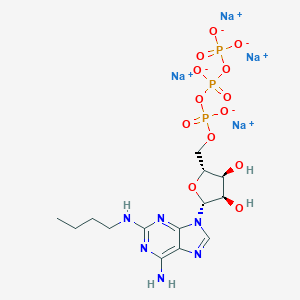 CHEMBL2364568 CHEMBL2364568 | C14H21N6Na4O13P3 | 666.231 | 18 / 4 | N/A | No |
| 84514 | 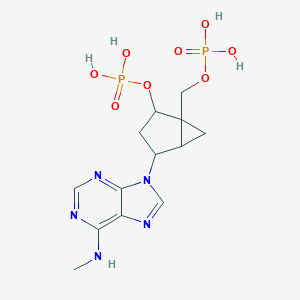 CHEMBL168427 CHEMBL168427 | C13H19N5O8P2 | 435.27 | 12 / 5 | -2.5 | No |
| 86028 | 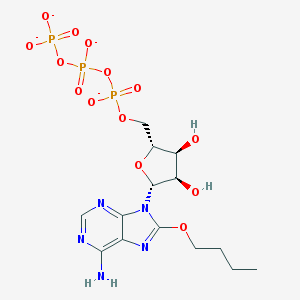 CHEMBL2364566 CHEMBL2364566 | C14H20N5O14P3-4 | 575.256 | 18 / 3 | -4.5 | No |
| 87026 | 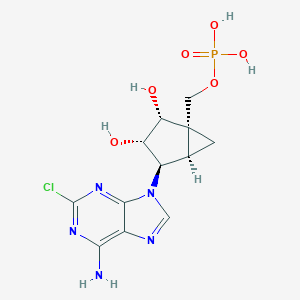 CHEMBL2373945 CHEMBL2373945 | C12H15ClN5O6P | 391.705 | 10 / 5 | -2.1 | Yes |
| 87027 | 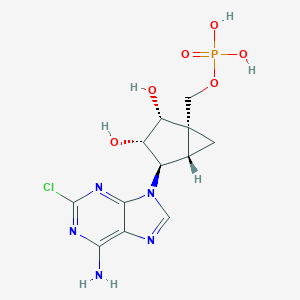 CHEMBL603128 CHEMBL603128 | C12H15ClN5O6P | 391.705 | 10 / 5 | -2.1 | Yes |
| 560190 | 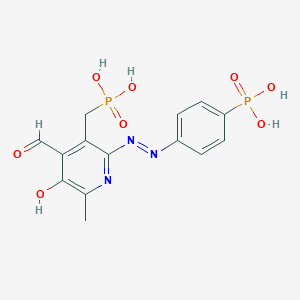 CHEMBL331250 CHEMBL331250 | C14H15N3O8P2 | 415.235 | 11 / 5 | -1.0 | No |
| 560325 | 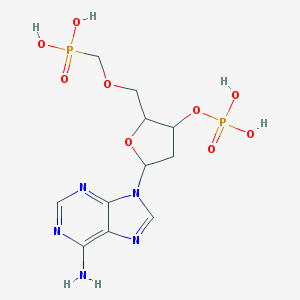 CHEMBL42554 CHEMBL42554 | C11H17N5O9P2 | 425.231 | 13 / 5 | -3.2 | No |
| 97200 | 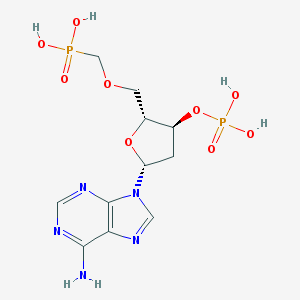 CHEMBL3144303 CHEMBL3144303 | C11H17N5O9P2 | 425.231 | 13 / 5 | -3.2 | No |
| 99787 | 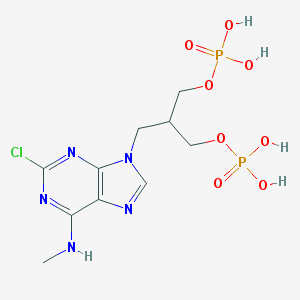 CHEMBL104784 CHEMBL104784 | C10H16ClN5O8P2 | 431.663 | 12 / 5 | -2.0 | No |
| 101694 | 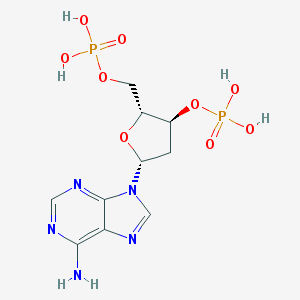 SCHEMBL375910 SCHEMBL375910 | C10H15N5O9P2 | 411.204 | 13 / 5 | -3.1 | No |
| 520133 | 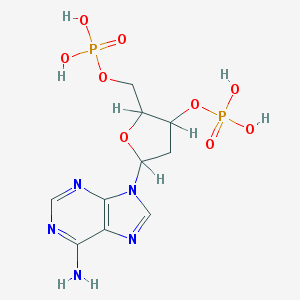 AC1NQBAN AC1NQBAN | C10H15N5O9P2 | 411.204 | 13 / 5 | -3.1 | No |
| 103550 | 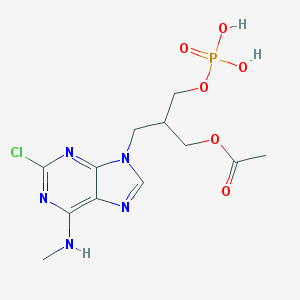 CHEMBL320073 CHEMBL320073 | C12H17ClN5O6P | 393.721 | 10 / 3 | -0.4 | Yes |
| 111330 | 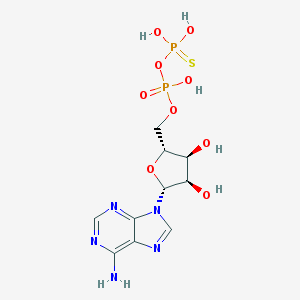 Adpbetas Adpbetas | C10H15N5O9P2S | 443.264 | 14 / 6 | -2.9 | No |
| 112892 | 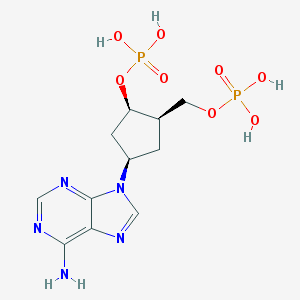 CHEMBL165225 CHEMBL165225 | C11H17N5O8P2 | 409.232 | 12 / 5 | -3.1 | No |
| 476814 | 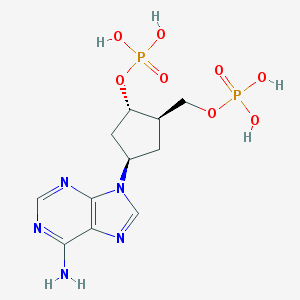 CHEMBL3706408 CHEMBL3706408 | C11H17N5O8P2 | 409.232 | 12 / 5 | -3.1 | No |
| 560807 | 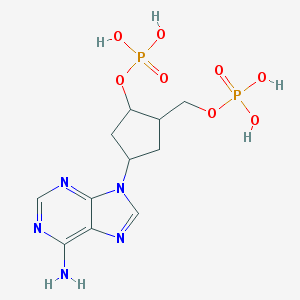 CHEMBL43903 CHEMBL43903 | C11H17N5O8P2 | 409.232 | 12 / 5 | -3.1 | No |
| 476848 | 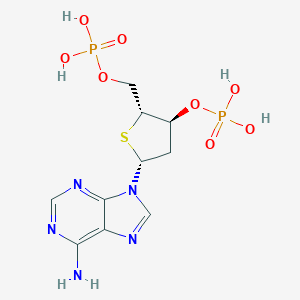 CHEMBL3706409 CHEMBL3706409 | C10H15N5O8P2S | 427.265 | 13 / 5 | -2.9 | No |
| 560816 | 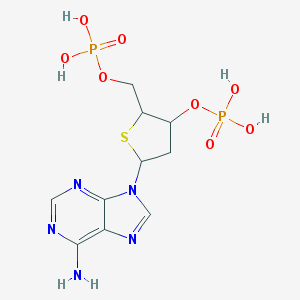 CHEMBL288993 CHEMBL288993 | C10H15N5O8P2S | 427.265 | 13 / 5 | -2.9 | No |
| 446178 | 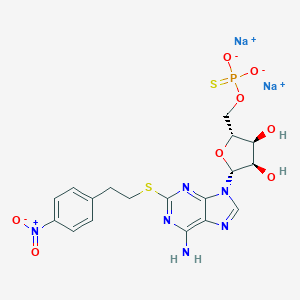 CHEMBL3351026 CHEMBL3351026 | C18H19N6Na2O8PS2 | 588.457 | 14 / 3 | N/A | No |
| 115007 | 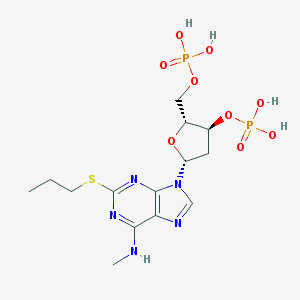 CHEMBL3144152 CHEMBL3144152 | C14H23N5O9P2S | 499.372 | 14 / 5 | -1.3 | No |
| 560846 | 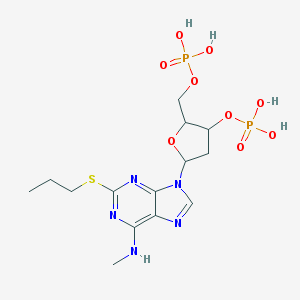 CHEMBL440662 CHEMBL440662 | C14H23N5O9P2S | 499.372 | 14 / 5 | -1.3 | No |
| 116532 | 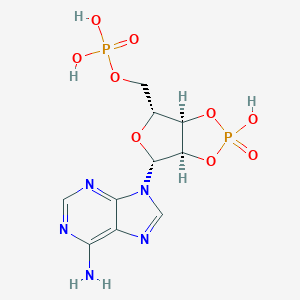 pA>p pA>p | C10H13N5O9P2 | 409.188 | 13 / 4 | -3.7 | No |
| 520186 | 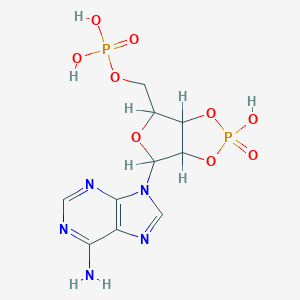 AC1NBNI5 AC1NBNI5 | C10H13N5O9P2 | 409.188 | 13 / 4 | -3.7 | No |
| 118084 | 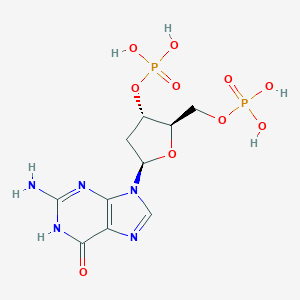 2'-Deoxyguanosine 3',5'-diphosphate 2'-Deoxyguanosine 3',5'-diphosphate | C10H15N5O10P2 | 427.203 | 12 / 6 | -4.1 | No |
| 560954 | 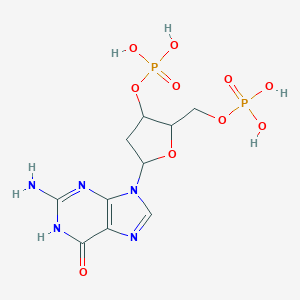 CHEMBL264849 CHEMBL264849 | C10H15N5O10P2 | 427.203 | 12 / 6 | -4.1 | No |
| 122706 | 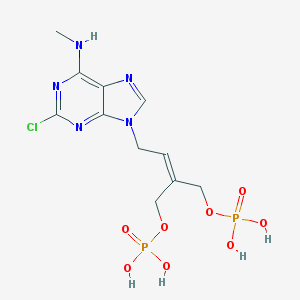 CHEMBL323457 CHEMBL323457 | C11H16ClN5O8P2 | 443.674 | 12 / 5 | -1.8 | No |
| 126010 | 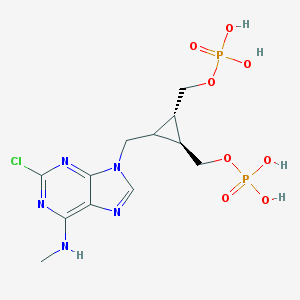 CHEMBL274496 CHEMBL274496 | C12H18ClN5O8P2 | 457.701 | 12 / 5 | -1.7 | No |
| 128330 | 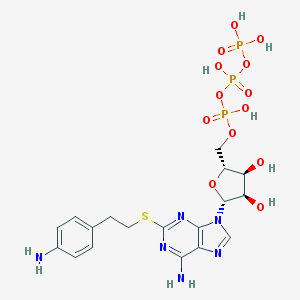 CHEMBL337062 CHEMBL337062 | C18H25N6O13P3S | 658.408 | 19 / 8 | -3.6 | No |
| 561281 | 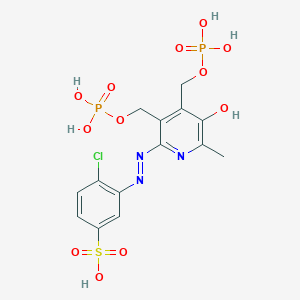 CHEMBL331238 CHEMBL331238 | C14H16ClN3O12P2S | 547.749 | 15 / 6 | -1.6 | No |
| 132208 | 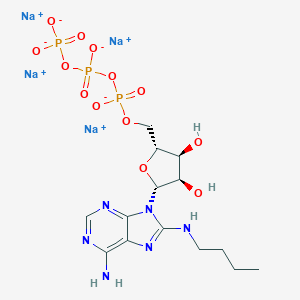 CHEMBL2364563 CHEMBL2364563 | C14H21N6Na4O13P3 | 666.231 | 18 / 4 | N/A | No |
| 133482 | 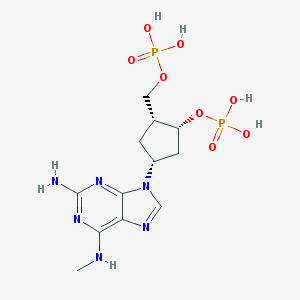 CHEMBL355406 CHEMBL355406 | C12H20N6O8P2 | 438.274 | 13 / 6 | -2.8 | No |
| 479343 | 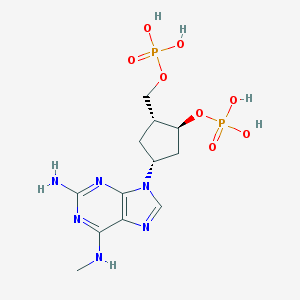 CHEMBL3706406 CHEMBL3706406 | C12H20N6O8P2 | 438.274 | 13 / 6 | -2.8 | No |
| 561444 | 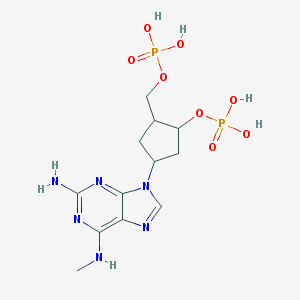 CHEMBL295051 CHEMBL295051 | C12H20N6O8P2 | 438.274 | 13 / 6 | -2.8 | No |
| 135293 | 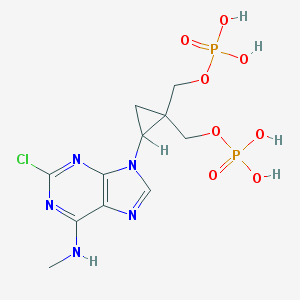 CHEMBL431649 CHEMBL431649 | C11H16ClN5O8P2 | 443.674 | 12 / 5 | -2.0 | No |
| 140621 | 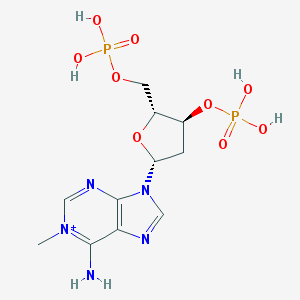 CHEMBL3144473 CHEMBL3144473 | C11H18N5O9P2+ | 426.239 | 12 / 5 | -3.5 | No |
| 145521 | 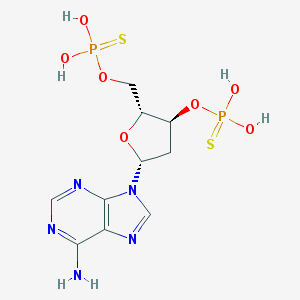 CHEMBL3144476 CHEMBL3144476 | C10H15N5O7P2S2 | 443.326 | 13 / 5 | 0.3 | No |
| 520313 | 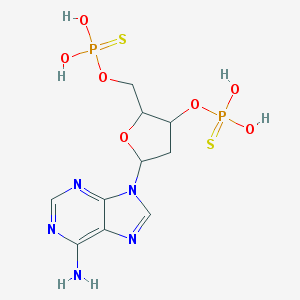 CHEMBL56787 CHEMBL56787 | C10H15N5O7P2S2 | 443.326 | 13 / 5 | 0.3 | No |
| 145547 | 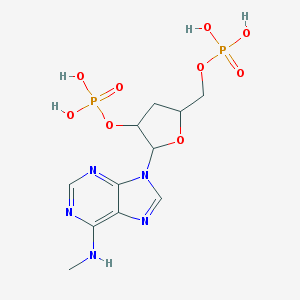 CHEMBL54116 CHEMBL54116 | C11H17N5O9P2 | 425.231 | 13 / 5 | -3.0 | No |
| 145860 | 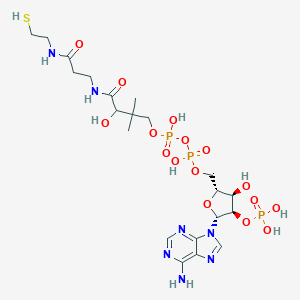 CHEMBL3144480 CHEMBL3144480 | C21H36N7O16P3S | 767.533 | 21 / 10 | -5.8 | No |
| 520315 | 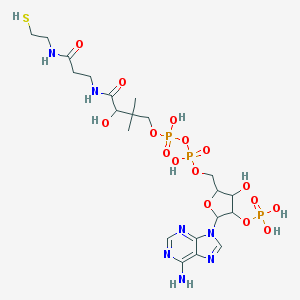 CHEMBL292013 CHEMBL292013 | C21H36N7O16P3S | 767.533 | 21 / 10 | -5.8 | No |
| 146120 | 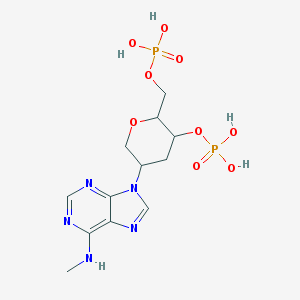 CHEMBL167191 CHEMBL167191 | C12H19N5O9P2 | 439.258 | 13 / 5 | -3.1 | No |
| 149835 | 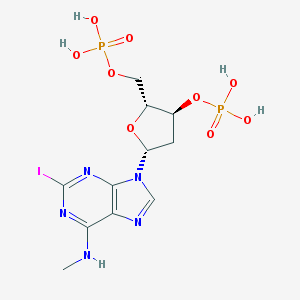 CHEMBL3144180 CHEMBL3144180 | C11H16IN5O9P2 | 551.127 | 13 / 5 | -2.3 | No |
| 561994 | 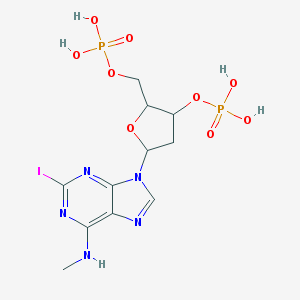 CHEMBL42028 CHEMBL42028 | C11H16IN5O9P2 | 551.127 | 13 / 5 | -2.3 | No |
| 562193 | 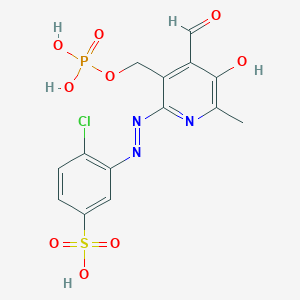 CHEMBL334823 CHEMBL334823 | C14H13ClN3O9PS | 465.754 | 12 / 4 | 0.4 | No |
| 156086 | 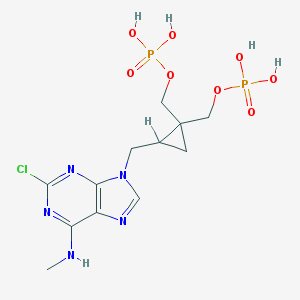 CHEMBL104752 CHEMBL104752 | C12H18ClN5O8P2 | 457.701 | 12 / 5 | -1.8 | No |
| 460644 | 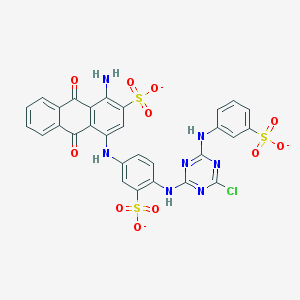 Basilen Blue Basilen Blue | C29H17ClN7O11S3-3 | 771.123 | 18 / 4 | 4.1 | No |
| 164699 | 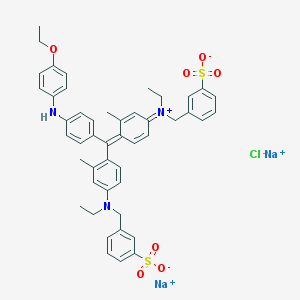 CHEMBL423337 CHEMBL423337 | C47H48ClN3Na2O7S2 | 912.465 | 10 / 1 | N/A | No |
| 172471 | 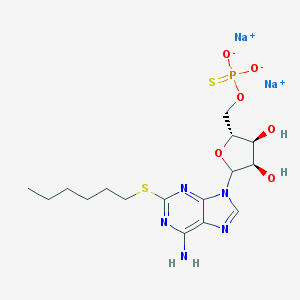 CHEMBL607771 CHEMBL607771 | C16H24N5Na2O6PS2 | 523.47 | 12 / 3 | N/A | No |
| 484842 | 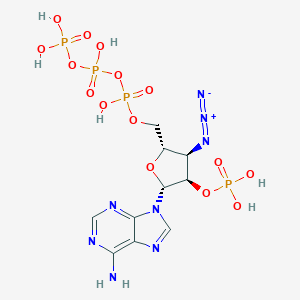 CHEMBL3706407 CHEMBL3706407 | C10H16N8O15P4 | 612.174 | 21 / 7 | -5.3 | No |
| 485717 | 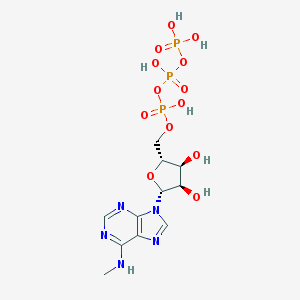 CHEMBL1094109 CHEMBL1094109 | C11H18N5O13P3 | 521.208 | 17 / 7 | -5.0 | No |
| 191793 | 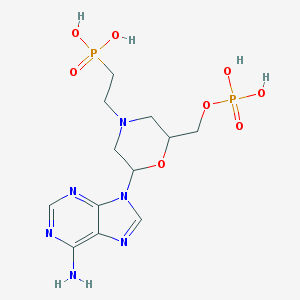 CHEMBL352431 CHEMBL352431 | C12H20N6O8P2 | 438.274 | 13 / 5 | -6.4 | No |
| 563341 | 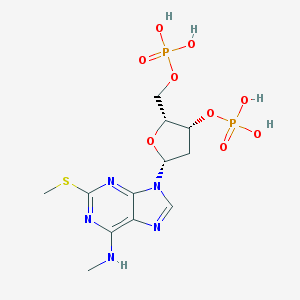 CHEMBL167061 CHEMBL167061 | C12H19N5O9P2S | 471.318 | 14 / 5 | -1.4 | No |
| 563342 | 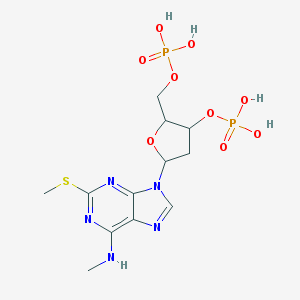 CHEMBL43924 CHEMBL43924 | C12H19N5O9P2S | 471.318 | 14 / 5 | -1.4 | No |
| 192045 | 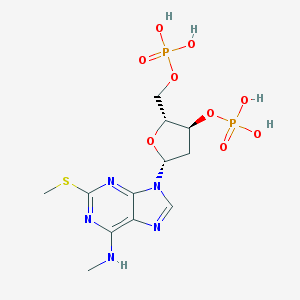 CHEMBL2368314 CHEMBL2368314 | C12H19N5O9P2S | 471.318 | 14 / 5 | -1.4 | No |
| 192296 | 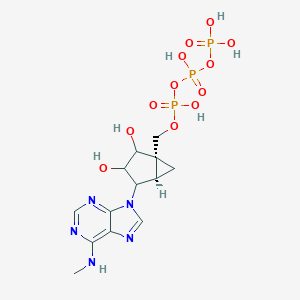 CHEMBL2373324 CHEMBL2373324 | C13H20N5O12P3 | 531.247 | 16 / 7 | -4.6 | No |
| 196164 | 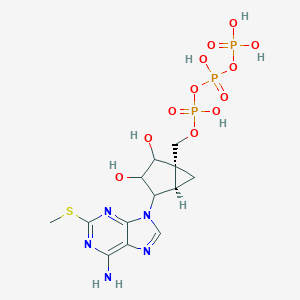 CHEMBL2373389 CHEMBL2373389 | C13H20N5O12P3S | 563.307 | 17 / 7 | -4.4 | No |
| 520508 | 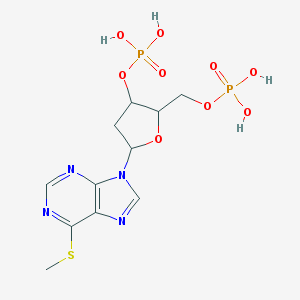 CHEMBL294601 CHEMBL294601 | C11H16N4O9P2S | 442.276 | 13 / 4 | -1.9 | No |
| 201491 | 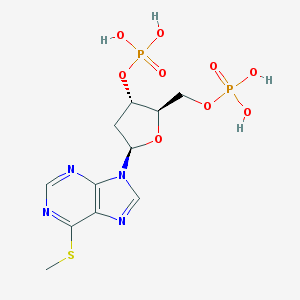 CHEMBL3144483 CHEMBL3144483 | C11H16N4O9P2S | 442.276 | 13 / 4 | -1.9 | No |
| 202916 | 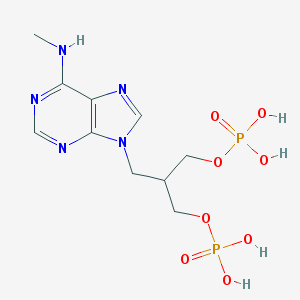 CHEMBL320924 CHEMBL320924 | C10H17N5O8P2 | 397.221 | 12 / 5 | -2.8 | No |
| 207978 | 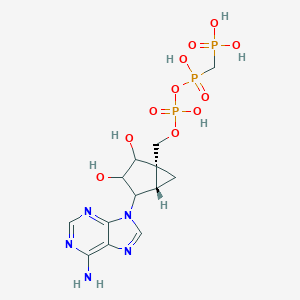 CHEMBL68067 CHEMBL68067 | C13H20N5O11P3 | 515.248 | 15 / 7 | -5.4 | No |
| 210414 | 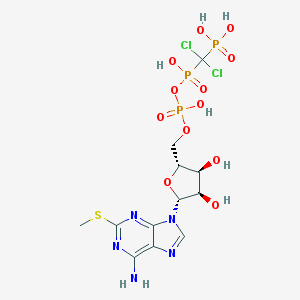 CHEMBL1089561 CHEMBL1089561 | C12H18Cl2N5O12P3S | 620.18 | 17 / 7 | -3.7 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218