

You can:
| Name | Prolactin-releasing peptide receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | PRLHR |
| Synonym | PrRP receptor hGR3 GPR10 G-protein coupled receptor 10 G protein-coupled receptor 10 [ Show all ] |
| Disease | N/A |
| Length | 370 |
| Amino acid sequence | MASSTTRGPRVSDLFSGLPPAVTTPANQSAEASAGNGSVAGADAPAVTPFQSLQLVHQLKGLIVLLYSVVVVVGLVGNCLLVLVIARVRRLHNVTNFLIGNLALSDVLMCTACVPLTLAYAFEPRGWVFGGGLCHLVFFLQPVTVYVSVFTLTTIAVDRYVVLVHPLRRRISLRLSAYAVLAIWALSAVLALPAAVHTYHVELKPHDVRLCEEFWGSQERQRQLYAWGLLLVTYLLPLLVILLSYVRVSVKLRNRVVPGCVTQSQADWDRARRRRTFCLLVVIVVVFAVCWLPLHVFNLLRDLDPHAIDPYAFGLVQLLCHWLAMSSACYNPFIYAWLHDSFREELRKLLVAWPRKIAPHGQNMTVSVVI |
| UniProt | P49683 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P49683 |
| 3D structure model | This predicted structure model is from GPCR-EXP P49683. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL1681611 |
| IUPHAR | 337 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 521508 | 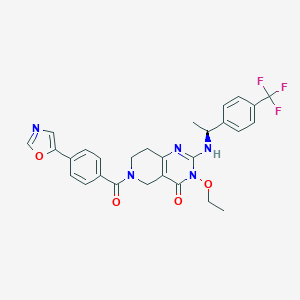 SCHEMBL434942 SCHEMBL434942 | C28H26F3N5O4 | 553.542 | 9 / 1 | 3.6 | No |
| 557358 | 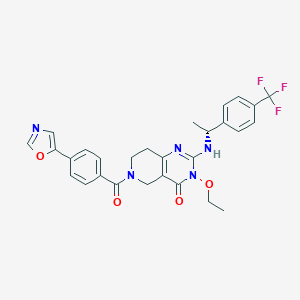 BDBM195750 BDBM195750 | C28H26F3N5O4 | 553.542 | 9 / 1 | 3.6 | No |
| 535979 | 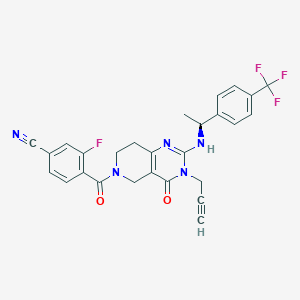 SCHEMBL433646 SCHEMBL433646 | C27H21F4N5O2 | 523.492 | 8 / 1 | 3.0 | No |
| 521923 | 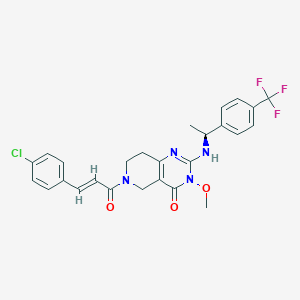 SCHEMBL448663 SCHEMBL448663 | C26H24ClF3N4O3 | 532.948 | 7 / 1 | 4.2 | No |
| 522006 | 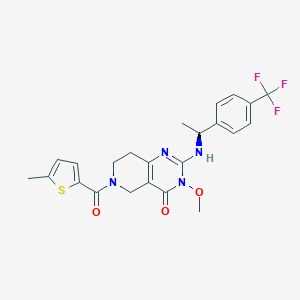 SCHEMBL448265 SCHEMBL448265 | C23H23F3N4O3S | 492.517 | 8 / 1 | 3.5 | Yes |
| 536443 | 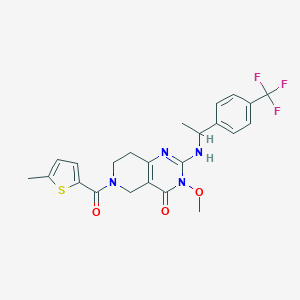 SCHEMBL437393 SCHEMBL437393 | C23H23F3N4O3S | 492.517 | 8 / 1 | 3.5 | Yes |
| 555586 | 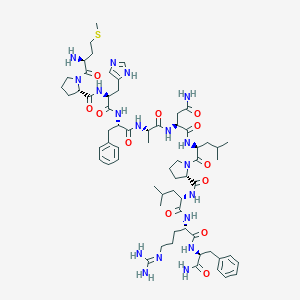 RFRP-1 RFRP-1 | C64H96N18O12S | 1341.65 | 16 / 14 | 0.2 | No |
| 536761 | 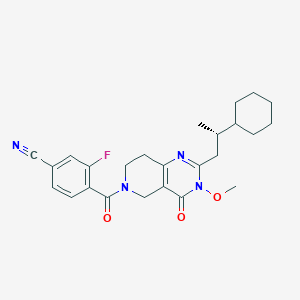 SCHEMBL437445 SCHEMBL437445 | C25H29FN4O3 | 452.53 | 6 / 0 | 3.8 | Yes |
| 522509 | 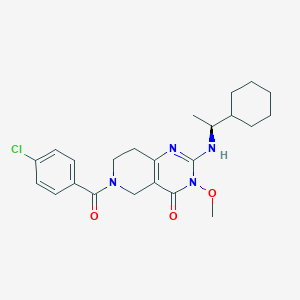 SCHEMBL450759 SCHEMBL450759 | C23H29ClN4O3 | 444.96 | 4 / 1 | 3.8 | Yes |
| 536816 | 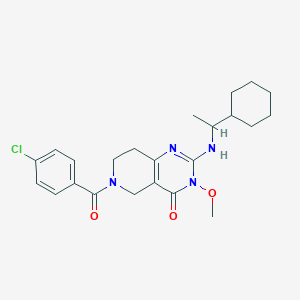 SCHEMBL437468 SCHEMBL437468 | C23H29ClN4O3 | 444.96 | 4 / 1 | 3.8 | Yes |
| 536862 | 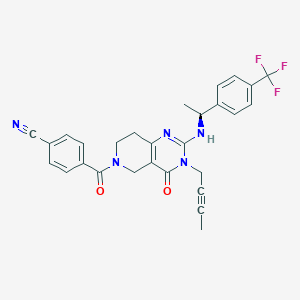 SCHEMBL435858 SCHEMBL435858 | C28H24F3N5O2 | 519.528 | 7 / 1 | 3.4 | No |
| 536907 | 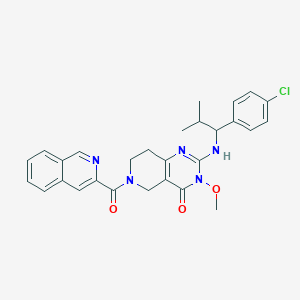 SCHEMBL435078 SCHEMBL435078 | C28H28ClN5O3 | 518.014 | 5 / 1 | 4.3 | No |
| 522610 | 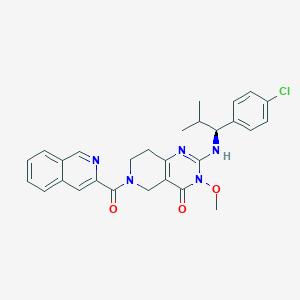 SCHEMBL448529 SCHEMBL448529 | C28H28ClN5O3 | 518.014 | 5 / 1 | 4.3 | No |
| 519858 | 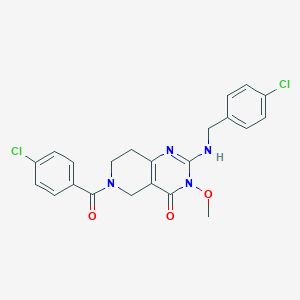 CHEMBL3734024 CHEMBL3734024 | C22H20Cl2N4O3 | 459.327 | 4 / 1 | 3.1 | Yes |
| 522636 | 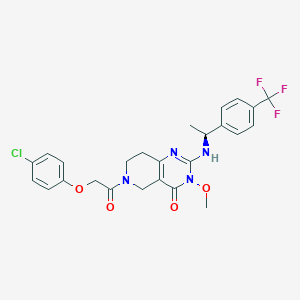 SCHEMBL435811 SCHEMBL435811 | C25H24ClF3N4O4 | 536.936 | 8 / 1 | 3.8 | No |
| 536933 | 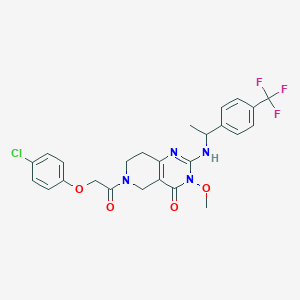 CHEMBL3914667 CHEMBL3914667 | C25H24ClF3N4O4 | 536.936 | 8 / 1 | 3.8 | No |
| 522900 | 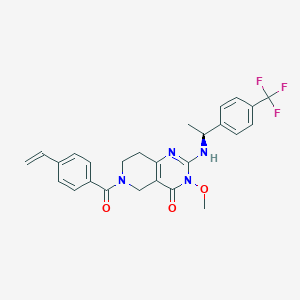 SCHEMBL437451 SCHEMBL437451 | C26H25F3N4O3 | 498.506 | 7 / 1 | 3.9 | Yes |
| 537205 | 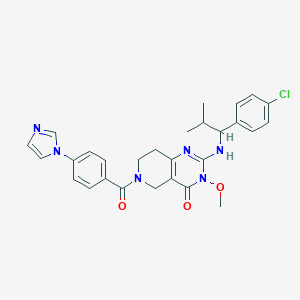 SCHEMBL435946 SCHEMBL435946 | C28H29ClN6O3 | 533.029 | 5 / 1 | 3.6 | No |
| 522937 | 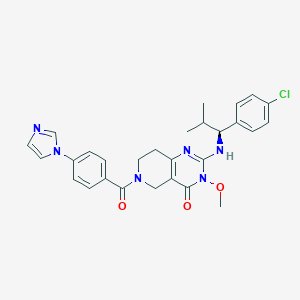 SCHEMBL448660 SCHEMBL448660 | C28H29ClN6O3 | 533.029 | 5 / 1 | 3.6 | No |
| 49374 | 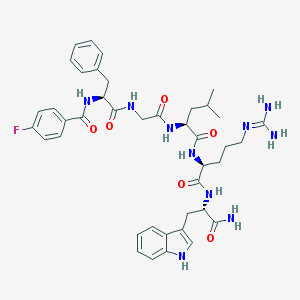 CHEMBL388586 CHEMBL388586 | C41H51FN10O6 | 798.921 | 8 / 9 | 2.4 | No |
| 519910 | 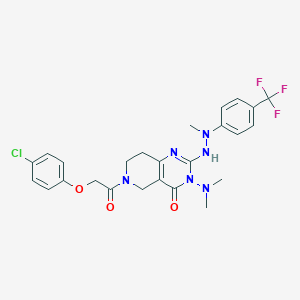 CHEMBL3986697 CHEMBL3986697 | C25H26ClF3N6O3 | 550.967 | 9 / 1 | 4.1 | No |
| 523171 | 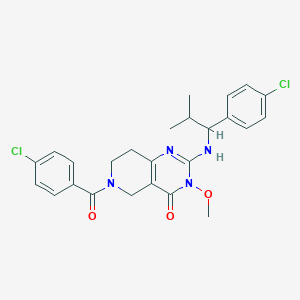 SCHEMBL437427 SCHEMBL437427 | C25H26Cl2N4O3 | 501.408 | 4 / 1 | 4.5 | No |
| 523265 | 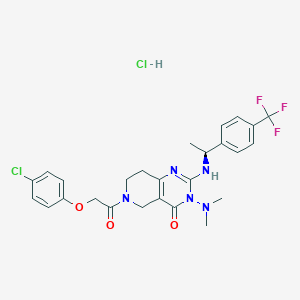 SCHEMBL451382 SCHEMBL451382 | C26H28Cl2F3N5O3 | 586.437 | 8 / 2 | N/A | No |
| 523473 | 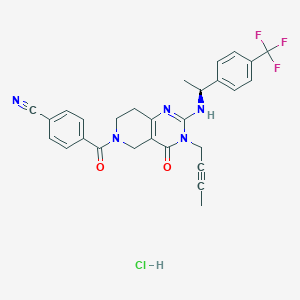 SCHEMBL447186 SCHEMBL447186 | C28H25ClF3N5O2 | 555.986 | 7 / 2 | N/A | No |
| 523804 | 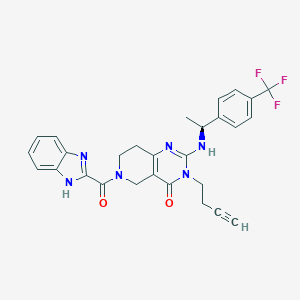 CHEMBL3729141 CHEMBL3729141 | C28H25F3N6O2 | 534.543 | 7 / 2 | 3.5 | No |
| 523836 | 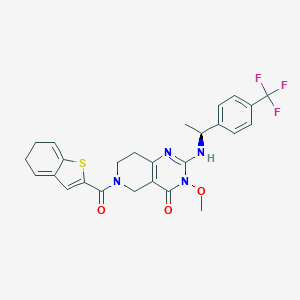 CHEMBL3728776 CHEMBL3728776 | C26H25F3N4O3S | 530.566 | 8 / 1 | 3.5 | No |
| 524057 | 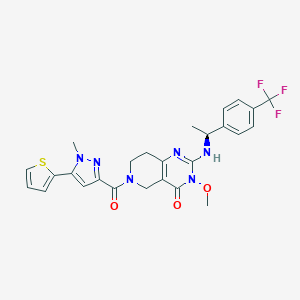 SCHEMBL448475 SCHEMBL448475 | C26H25F3N6O3S | 558.58 | 9 / 1 | 3.1 | No |
| 538168 | 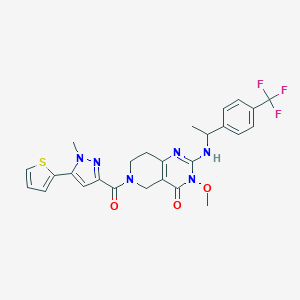 SCHEMBL437465 SCHEMBL437465 | C26H25F3N6O3S | 558.58 | 9 / 1 | 3.1 | No |
| 524058 | 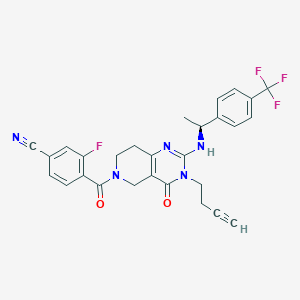 CHEMBL3728036 CHEMBL3728036 | C28H23F4N5O2 | 537.519 | 8 / 1 | 3.5 | No |
| 524424 | 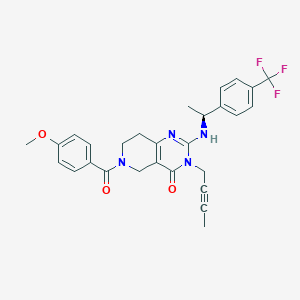 SCHEMBL435913 SCHEMBL435913 | C28H27F3N4O3 | 524.544 | 7 / 1 | 3.7 | No |
| 524516 | 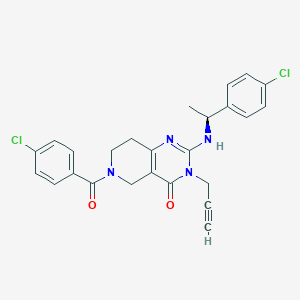 SCHEMBL436170 SCHEMBL436170 | C25H22Cl2N4O2 | 481.377 | 3 / 1 | 3.6 | Yes |
| 520152 | 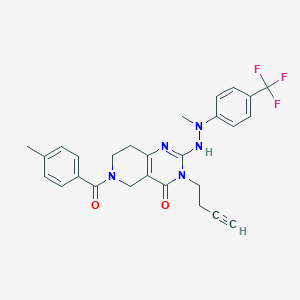 CHEMBL3916965 CHEMBL3916965 | C27H26F3N5O2 | 509.533 | 7 / 1 | 4.1 | No |
| 524712 | 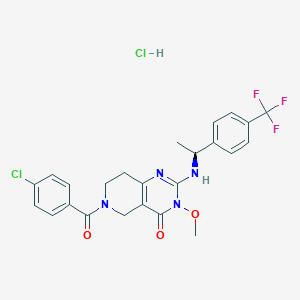 CHEMBL3727401 CHEMBL3727401 | C24H23Cl2F3N4O3 | 543.368 | 7 / 2 | N/A | No |
| 524838 | 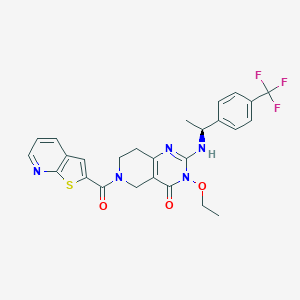 SCHEMBL435818 SCHEMBL435818 | C26H24F3N5O3S | 543.565 | 9 / 1 | 4.1 | No |
| 560842 | 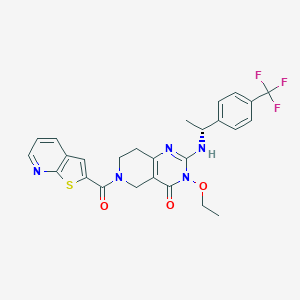 BDBM195751 BDBM195751 | C26H24F3N5O3S | 543.565 | 9 / 1 | 4.1 | No |
| 524895 | 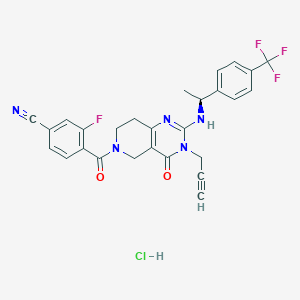 CHEMBL3730970 CHEMBL3730970 | C27H22ClF4N5O2 | 559.95 | 8 / 2 | N/A | No |
| 525055 | 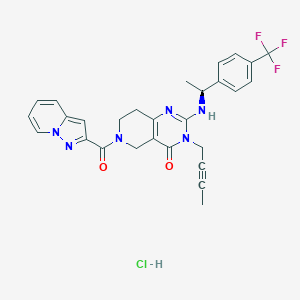 SCHEMBL448302 SCHEMBL448302 | C28H26ClF3N6O2 | 571.001 | 7 / 2 | N/A | No |
| 525153 | 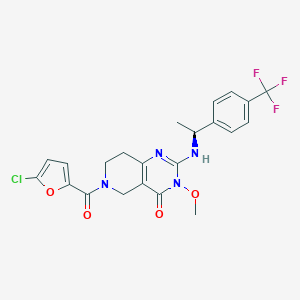 SCHEMBL452917 SCHEMBL452917 | C22H20ClF3N4O4 | 496.871 | 8 / 1 | 3.5 | Yes |
| 539102 | 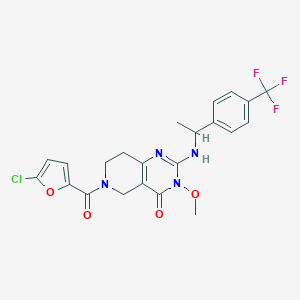 SCHEMBL437449 SCHEMBL437449 | C22H20ClF3N4O4 | 496.871 | 8 / 1 | 3.5 | Yes |
| 555978 | 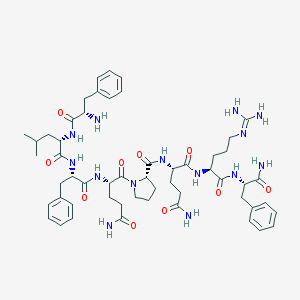 Neuropeptide FF Neuropeptide FF | C54H76N14O10 | 1081.29 | 12 / 12 | -0.3 | No |
| 555979 | 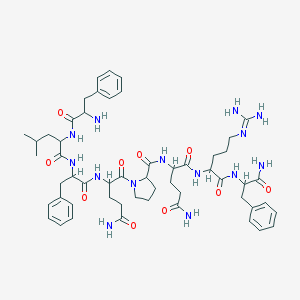 AC1NQ0SC AC1NQ0SC | C54H76N14O10 | 1081.29 | 12 / 12 | -0.3 | No |
| 525202 | 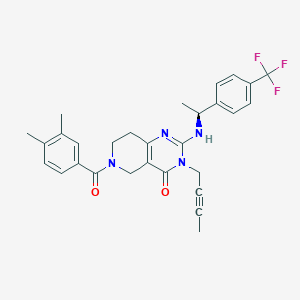 SCHEMBL435882 SCHEMBL435882 | C29H29F3N4O2 | 522.572 | 6 / 1 | 4.4 | No |
| 525254 | 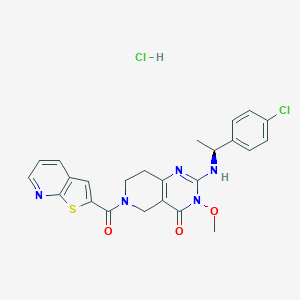 SCHEMBL452128 SCHEMBL452128 | C24H23Cl2N5O3S | 532.44 | 6 / 2 | N/A | No |
| 525272 | 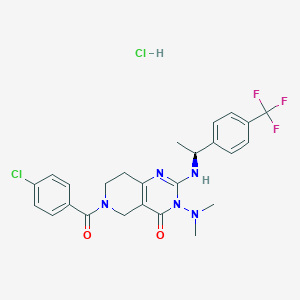 CHEMBL3728270 CHEMBL3728270 | C25H26Cl2F3N5O2 | 556.411 | 7 / 2 | N/A | No |
| 525297 | 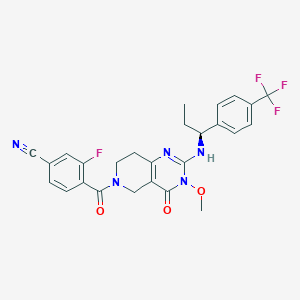 SCHEMBL447581 SCHEMBL447581 | C26H23F4N5O3 | 529.496 | 9 / 1 | 3.5 | No |
| 539289 | 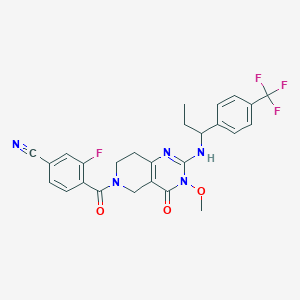 SCHEMBL437481 SCHEMBL437481 | C26H23F4N5O3 | 529.496 | 9 / 1 | 3.5 | No |
| 525586 | 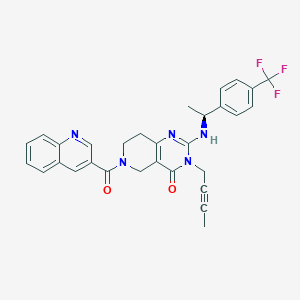 SCHEMBL437435 SCHEMBL437435 | C30H26F3N5O2 | 545.566 | 7 / 1 | 4.0 | No |
| 539614 | 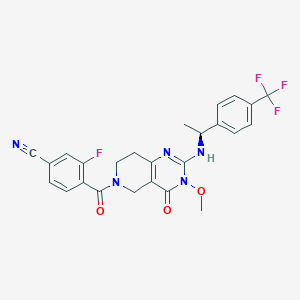 CHEMBL3734010 CHEMBL3734010 | C25H21F4N5O3 | 515.469 | 9 / 1 | 2.9 | No |
| 525649 | 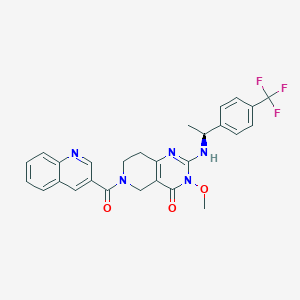 SCHEMBL452315 SCHEMBL452315 | C27H24F3N5O3 | 523.516 | 8 / 1 | 3.4 | No |
| 539631 | 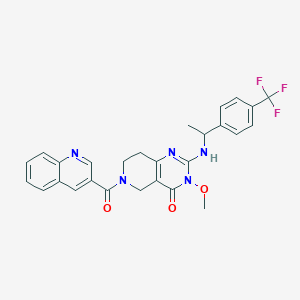 SCHEMBL437401 SCHEMBL437401 | C27H24F3N5O3 | 523.516 | 8 / 1 | 3.4 | No |
| 539674 | 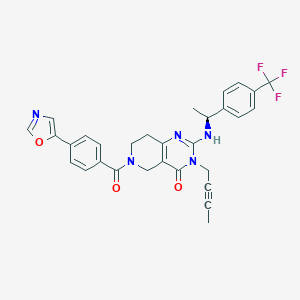 SCHEMBL434965 SCHEMBL434965 | C30H26F3N5O3 | 561.565 | 8 / 1 | 3.8 | No |
| 525739 | 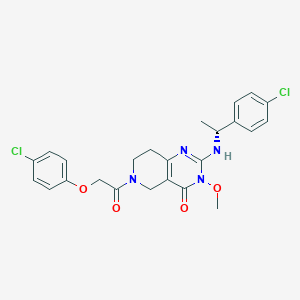 CHEMBL3729222 CHEMBL3729222 | C24H24Cl2N4O4 | 503.38 | 5 / 1 | 3.5 | No |
| 539722 | 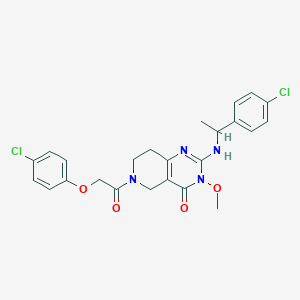 SCHEMBL434988 SCHEMBL434988 | C24H24Cl2N4O4 | 503.38 | 5 / 1 | 3.5 | No |
| 556049 | 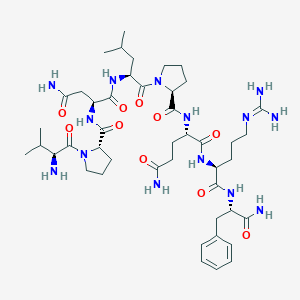 311309-27-0 311309-27-0 | C45H72N14O10 | 969.159 | 12 / 11 | -2.5 | No |
| 540051 | 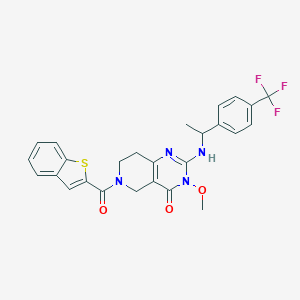 SCHEMBL435939 SCHEMBL435939 | C26H23F3N4O3S | 528.55 | 8 / 1 | 4.5 | No |
| 526033 | 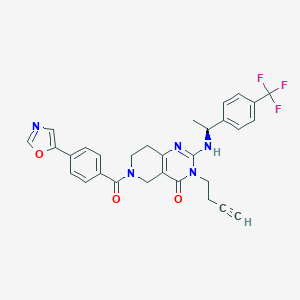 CHEMBL3728107 CHEMBL3728107 | C30H26F3N5O3 | 561.565 | 8 / 1 | 3.7 | No |
| 526316 | 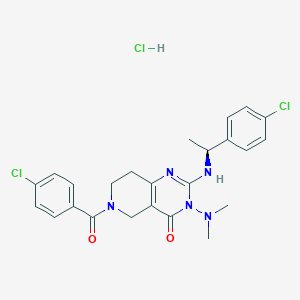 CHEMBL3731865 CHEMBL3731865 | C24H26Cl3N5O2 | 522.855 | 4 / 2 | N/A | No |
| 556174 | 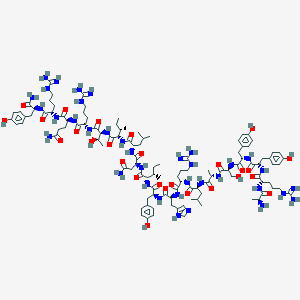 BDBM50368404 BDBM50368404 | C112H174N36O27 | 2456.85 | 33 / 41 | -3.6 | No |
| 526466 | 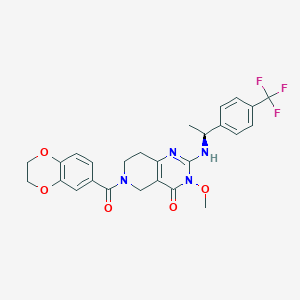 SCHEMBL448830 SCHEMBL448830 | C26H25F3N4O5 | 530.504 | 9 / 1 | 2.8 | No |
| 540485 | 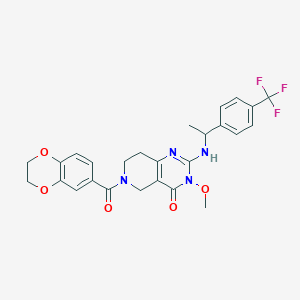 SCHEMBL435906 SCHEMBL435906 | C26H25F3N4O5 | 530.504 | 9 / 1 | 2.8 | No |
| 556189 | 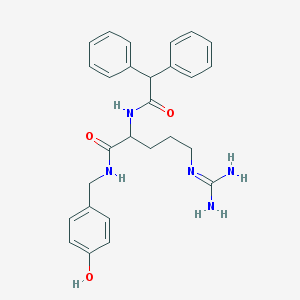 CHEMBL44246 CHEMBL44246 | C27H31N5O3 | 473.577 | 4 / 5 | 2.5 | Yes |
| 179485 | 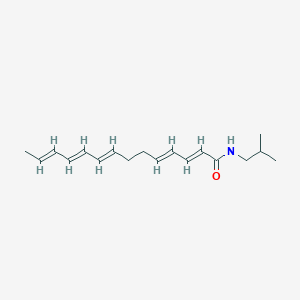 N-Isobutyl-2,4,8,10,12-tetradecapentaenamide N-Isobutyl-2,4,8,10,12-tetradecapentaenamide | C18H27NO | 273.42 | 1 / 1 | 4.9 | Yes |
| 526613 | 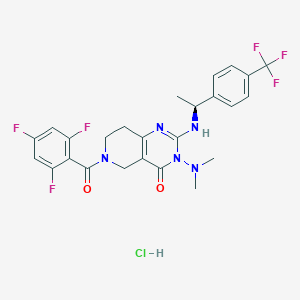 SCHEMBL447348 SCHEMBL447348 | C25H24ClF6N5O2 | 575.94 | 10 / 2 | N/A | No |
| 527047 | 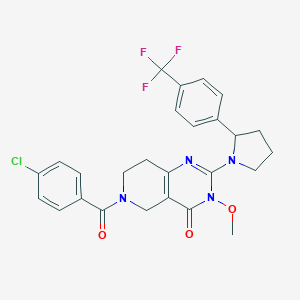 SCHEMBL437447 SCHEMBL437447 | C26H24ClF3N4O3 | 532.948 | 7 / 0 | 4.1 | No |
| 527222 | 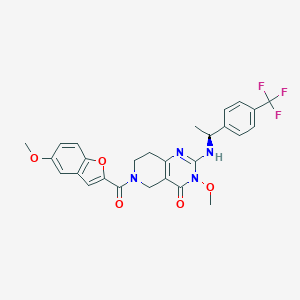 SCHEMBL448484 SCHEMBL448484 | C27H25F3N4O5 | 542.515 | 9 / 1 | 3.8 | No |
| 541126 | 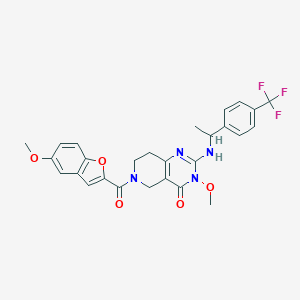 SCHEMBL435923 SCHEMBL435923 | C27H25F3N4O5 | 542.515 | 9 / 1 | 3.8 | No |
| 541131 | 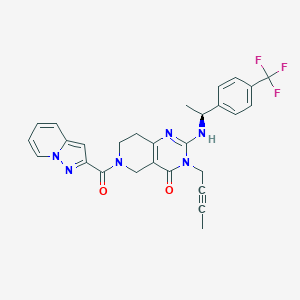 SCHEMBL437394 SCHEMBL437394 | C28H25F3N6O2 | 534.543 | 7 / 1 | 3.2 | No |
| 527236 | 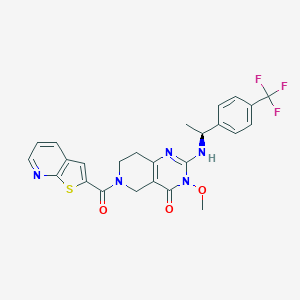 SCHEMBL452594 SCHEMBL452594 | C25H22F3N5O3S | 529.538 | 9 / 1 | 3.7 | No |
| 541134 | 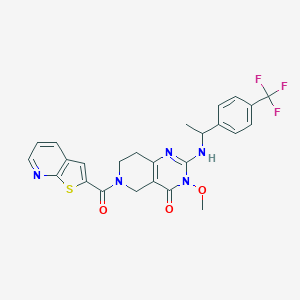 SCHEMBL436929 SCHEMBL436929 | C25H22F3N5O3S | 529.538 | 9 / 1 | 3.7 | No |
| 520496 | 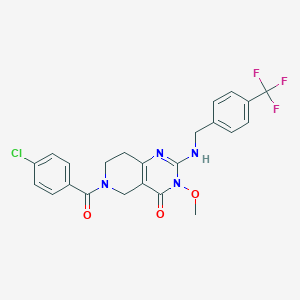 CHEMBL3734026 CHEMBL3734026 | C23H20ClF3N4O3 | 492.883 | 7 / 1 | 3.3 | Yes |
| 527441 | 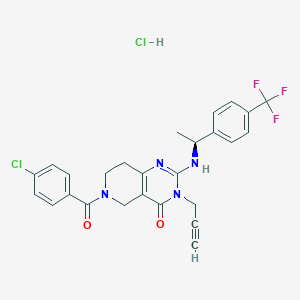 CHEMBL3732064 CHEMBL3732064 | C26H23Cl2F3N4O2 | 551.391 | 6 / 2 | N/A | No |
| 527644 | 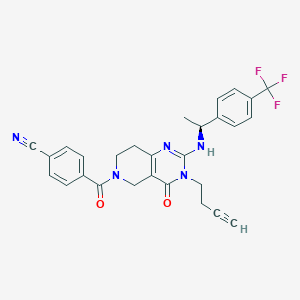 SCHEMBL452837 SCHEMBL452837 | C28H24F3N5O2 | 519.528 | 7 / 1 | 3.4 | No |
| 527702 | 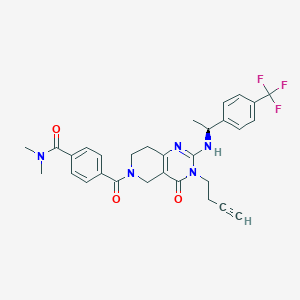 CHEMBL3731809 CHEMBL3731809 | C30H30F3N5O3 | 565.597 | 7 / 1 | 3.1 | No |
| 527740 | 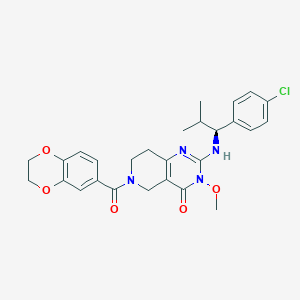 SCHEMBL448840 SCHEMBL448840 | C27H29ClN4O5 | 525.002 | 6 / 1 | 3.5 | No |
| 541665 | 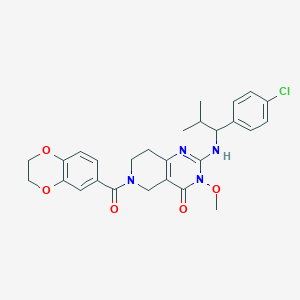 SCHEMBL437458 SCHEMBL437458 | C27H29ClN4O5 | 525.002 | 6 / 1 | 3.5 | No |
| 527800 | 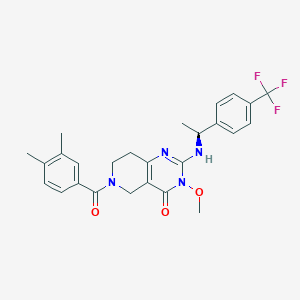 SCHEMBL450489 SCHEMBL450489 | C26H27F3N4O3 | 500.522 | 7 / 1 | 3.8 | No |
| 541751 | 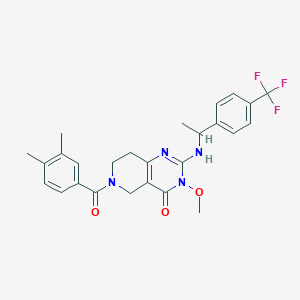 SCHEMBL437437 SCHEMBL437437 | C26H27F3N4O3 | 500.522 | 7 / 1 | 3.8 | No |
| 527841 | 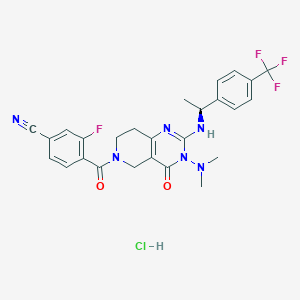 CHEMBL3728414 CHEMBL3728414 | C26H25ClF4N6O2 | 564.97 | 9 / 2 | N/A | No |
| 527874 | 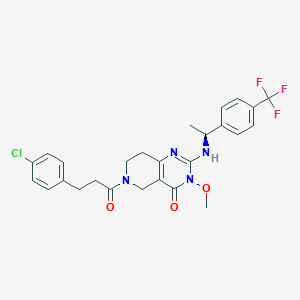 CHEMBL3727412 CHEMBL3727412 | C26H26ClF3N4O3 | 534.964 | 7 / 1 | 4.0 | No |
| 541803 | 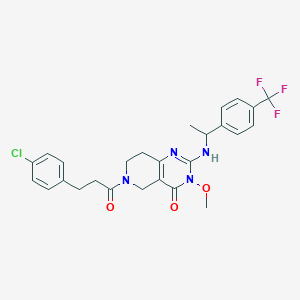 CHEMBL3901490 CHEMBL3901490 | C26H26ClF3N4O3 | 534.964 | 7 / 1 | 4.0 | No |
| 528134 | 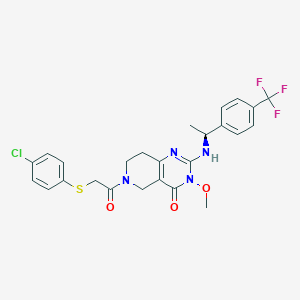 SCHEMBL437371 SCHEMBL437371 | C25H24ClF3N4O3S | 552.997 | 8 / 1 | 4.3 | No |
| 542029 | 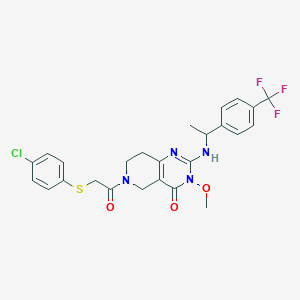 CHEMBL3969131 CHEMBL3969131 | C25H24ClF3N4O3S | 552.997 | 8 / 1 | 4.3 | No |
| 528197 | 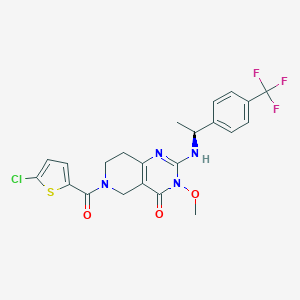 SCHEMBL448631 SCHEMBL448631 | C22H20ClF3N4O3S | 512.932 | 8 / 1 | 4.1 | No |
| 542132 | 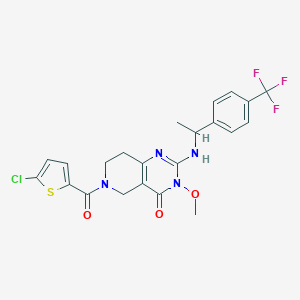 SCHEMBL436905 SCHEMBL436905 | C22H20ClF3N4O3S | 512.932 | 8 / 1 | 4.1 | No |
| 528298 | 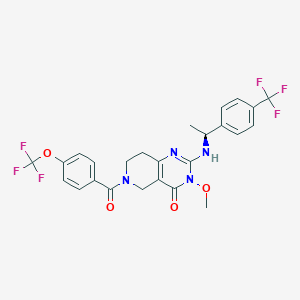 SCHEMBL435817 SCHEMBL435817 | C25H22F6N4O4 | 556.465 | 11 / 1 | 4.3 | No |
| 564710 | 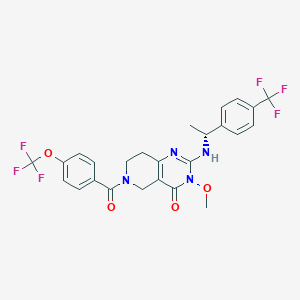 BDBM195746 BDBM195746 | C25H22F6N4O4 | 556.465 | 11 / 1 | 4.3 | No |
| 528452 | 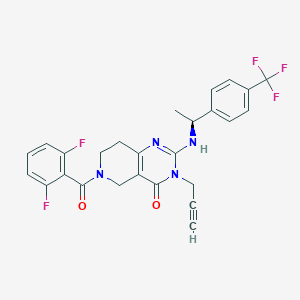 SCHEMBL450105 SCHEMBL450105 | C26H21F5N4O2 | 516.472 | 8 / 1 | 3.4 | No |
| 528469 | 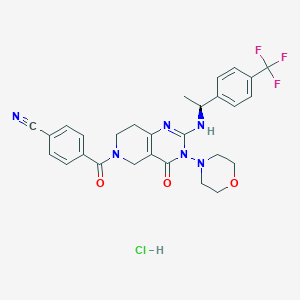 SCHEMBL447217 SCHEMBL447217 | C28H28ClF3N6O3 | 589.016 | 9 / 2 | N/A | No |
| 556417 | 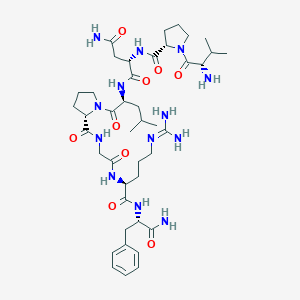 PrRP24-31 PrRP24-31 | C42H67N13O9 | 898.08 | 11 / 10 | -1.8 | No |
| 542435 | 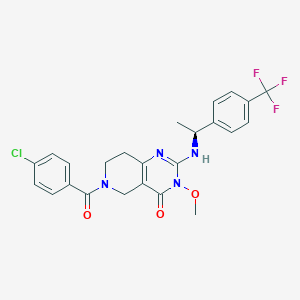 SCHEMBL435830 SCHEMBL435830 | C24H22ClF3N4O3 | 506.91 | 7 / 1 | 3.7 | No |
| 528645 | 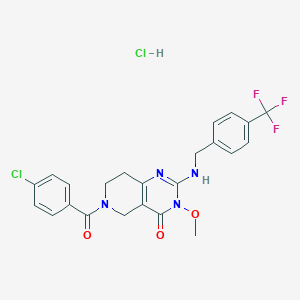 SCHEMBL452039 SCHEMBL452039 | C23H21Cl2F3N4O3 | 529.341 | 7 / 2 | N/A | No |
| 252535 | 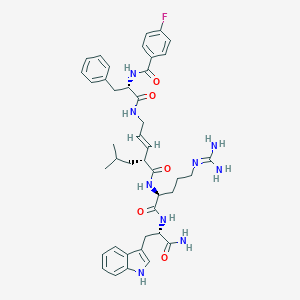 Peptide analogue, 19 Peptide analogue, 19 | C42H52FN9O5 | 781.934 | 7 / 8 | 3.4 | No |
| 528882 | 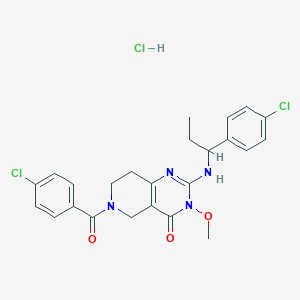 SCHEMBL449454 SCHEMBL449454 | C24H25Cl3N4O3 | 523.839 | 4 / 2 | N/A | No |
| 529047 | 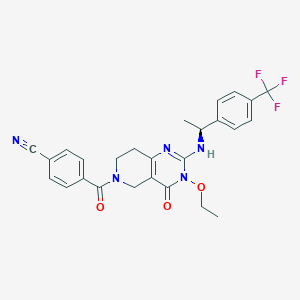 SCHEMBL434947 SCHEMBL434947 | C26H24F3N5O3 | 511.505 | 8 / 1 | 3.2 | No |
| 565432 | 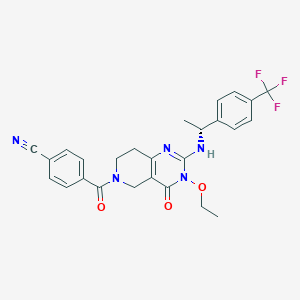 BDBM195748 BDBM195748 | C26H24F3N5O3 | 511.505 | 8 / 1 | 3.2 | No |
| 529257 | 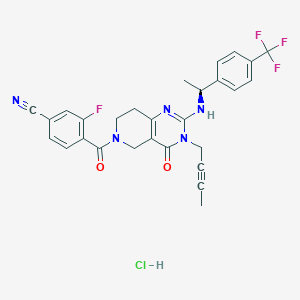 SCHEMBL448522 SCHEMBL448522 | C28H24ClF4N5O2 | 573.977 | 8 / 2 | N/A | No |
| 543361 | 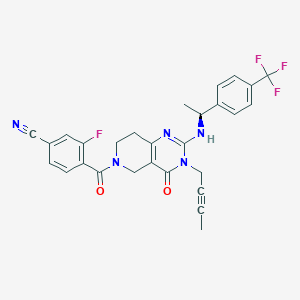 SCHEMBL434957 SCHEMBL434957 | C28H23F4N5O2 | 537.519 | 8 / 1 | 3.5 | No |
| 529447 | 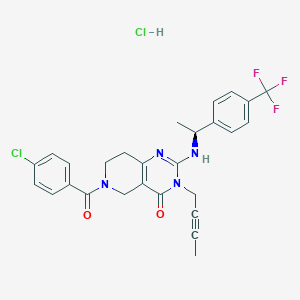 SCHEMBL450296 SCHEMBL450296 | C27H25Cl2F3N4O2 | 565.418 | 6 / 2 | N/A | No |
| 529474 | 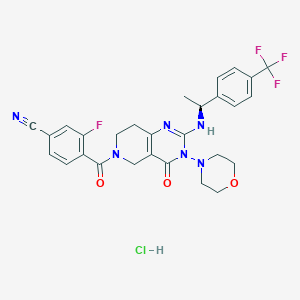 SCHEMBL450716 SCHEMBL450716 | C28H27ClF4N6O3 | 607.007 | 10 / 2 | N/A | No |
| 543585 | 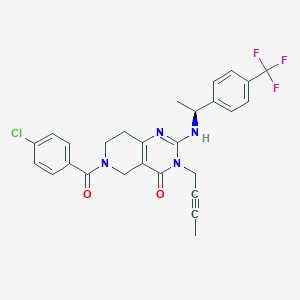 SCHEMBL435859 SCHEMBL435859 | C27H24ClF3N4O2 | 528.96 | 6 / 1 | 4.3 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218