

You can:
| Name | C-C chemokine receptor type 1 |
|---|---|
| Species | Mus musculus (Mouse) |
| Gene | Ccr1 |
| Synonym | MIP1aR MIP-1alphaR MIP-1alpha/RANTES MIP-1alpha-R macrophage inflammatory protein-1 alpha receptor [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 355 |
| Amino acid sequence | MEISDFTEAYPTTTEFDYGDSTPCQKTAVRAFGAGLLPPLYSLVFIIGVVGNVLVILVLMQHRRLQSMTSIYLFNLAVSDLVFLFTLPFWIDYKLKDDWIFGDAMCKLLSGFYYLGLYSEIFFIILLTIDRYLAIVHAVFALRARTVTFGIITSIITWALAILASMPALYFFKAQWEFTHRTCSPHFPYKSLKQWKRFQALKLNLLGLILPLLVMIICYAGIIRILLRRPSEKKVKAVRLIFAITLLFFLLWTPYNLSVFVSAFQDVLFTNQCEQSKQLDLAMQVTEVIAYTHCCVNPIIYVFVGERFWKYLRQLFQRHVAIPLAKWLPFLSVDQLERTSSISPSTGEHELSAGF |
| UniProt | P51675 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3872 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 6001 | 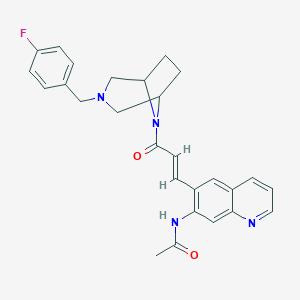 CHEMBL200680 CHEMBL200680 | C27H27FN4O2 | 458.537 | 5 / 1 | 3.3 | Yes |
| 19136 | 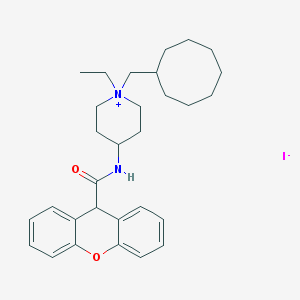 CHEMBL280751 CHEMBL280751 | C30H41IN2O2 | 588.574 | 3 / 1 | N/A | No |
| 33398 | 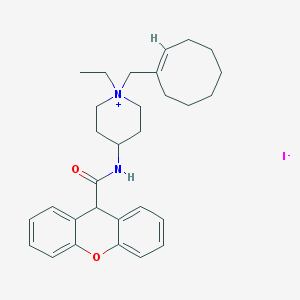 CHEMBL34732 CHEMBL34732 | C30H39IN2O2 | 586.558 | 3 / 1 | N/A | No |
| 36133 | 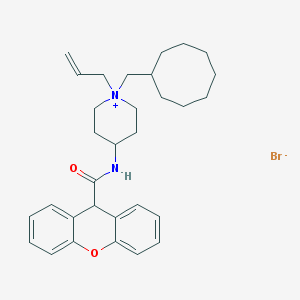 CHEMBL33104 CHEMBL33104 | C31H41BrN2O2 | 553.585 | 3 / 1 | N/A | No |
| 459541 | 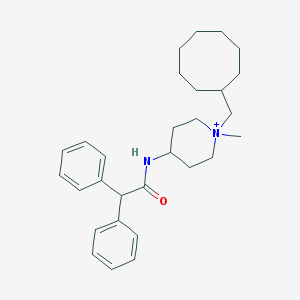 CHEMBL33761 CHEMBL33761 | C29H41N2O+ | 433.66 | 1 / 1 | 6.9 | No |
| 37962 | 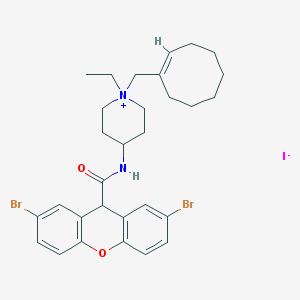 CHEMBL289902 CHEMBL289902 | C30H37Br2IN2O2 | 744.35 | 3 / 1 | N/A | No |
| 45506 | 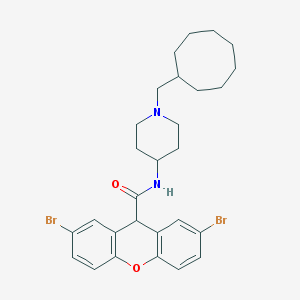 CHEMBL32305 CHEMBL32305 | C28H34Br2N2O2 | 590.4 | 3 / 1 | 7.9 | No |
| 53458 | 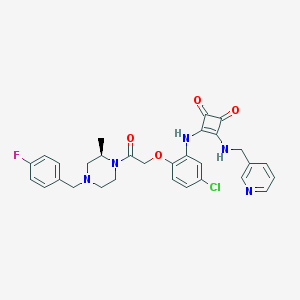 CHEMBL397910 CHEMBL397910 | C30H29ClFN5O4 | 578.041 | 9 / 2 | 3.9 | No |
| 62037 | 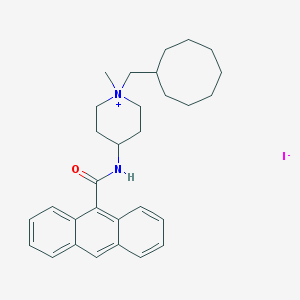 CHEMBL289682 CHEMBL289682 | C30H39IN2O | 570.559 | 2 / 1 | N/A | No |
| 64358 | 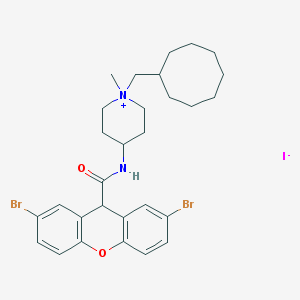 CHEMBL33223 CHEMBL33223 | C29H37Br2IN2O2 | 732.339 | 3 / 1 | N/A | No |
| 68992 | 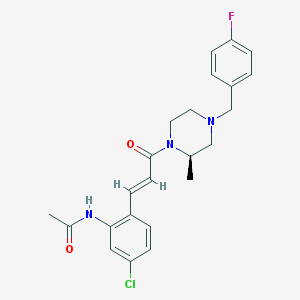 CHEMBL198949 CHEMBL198949 | C23H25ClFN3O2 | 429.92 | 4 / 1 | 3.5 | Yes |
| 459897 | 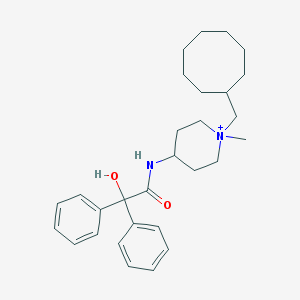 CHEMBL433232 CHEMBL433232 | C29H41N2O2+ | 449.659 | 2 / 2 | 6.1 | No |
| 74392 | 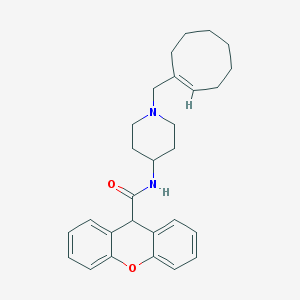 CHEMBL33277 CHEMBL33277 | C28H34N2O2 | 430.592 | 3 / 1 | 5.7 | No |
| 83079 | 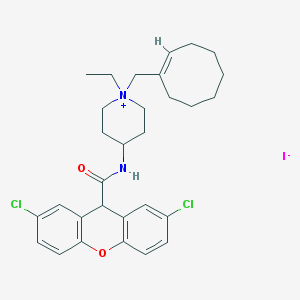 UNII-HD099HOT7H UNII-HD099HOT7H | C30H37Cl2IN2O2 | 655.442 | 3 / 1 | N/A | No |
| 83533 | 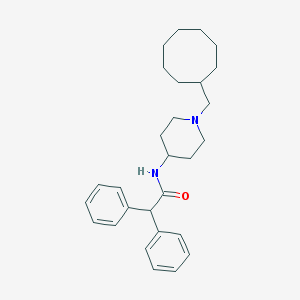 CHEMBL286355 CHEMBL286355 | C28H38N2O | 418.625 | 2 / 1 | 6.9 | No |
| 84495 | 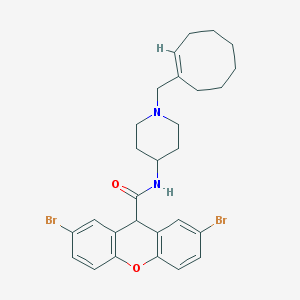 CHEMBL33580 CHEMBL33580 | C28H32Br2N2O2 | 588.384 | 3 / 1 | 7.1 | No |
| 85737 | 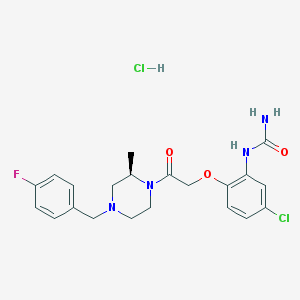 BX471 hydrochloride BX471 hydrochloride | C21H25Cl2FN4O3 | 471.354 | 5 / 3 | N/A | N/A |
| 460037 | 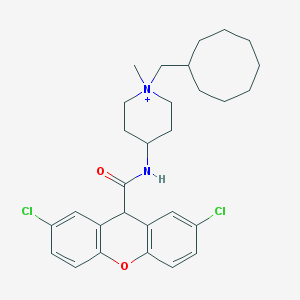 CHEMBL33929 CHEMBL33929 | C29H37Cl2N2O2+ | 516.527 | 2 / 1 | 7.8 | No |
| 460108 | 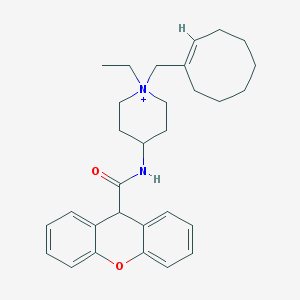 CHEMBL34732 CHEMBL34732 | C30H39N2O2+ | 459.654 | 2 / 1 | 6.1 | No |
| 103858 | 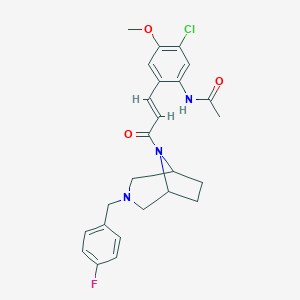 CHEMBL372807 CHEMBL372807 | C25H27ClFN3O3 | 471.957 | 5 / 1 | 3.6 | Yes |
| 445832 | 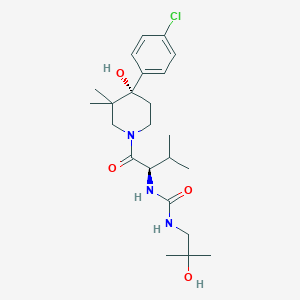 UNII-AKQ3X6FEH0 UNII-AKQ3X6FEH0 | C23H36ClN3O4 | 454.008 | 4 / 4 | 2.8 | Yes |
| 112887 | 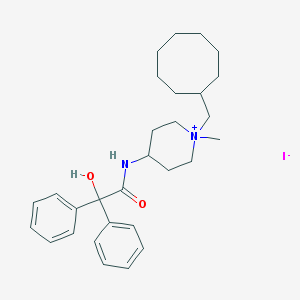 CHEMBL433232 CHEMBL433232 | C29H41IN2O2 | 576.563 | 3 / 2 | N/A | No |
| 113263 | 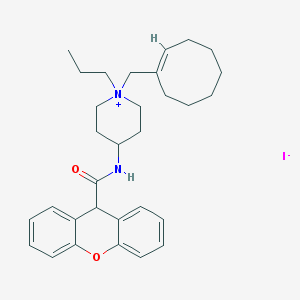 CHEMBL33050 CHEMBL33050 | C31H41IN2O2 | 600.585 | 3 / 1 | N/A | No |
| 144576 | 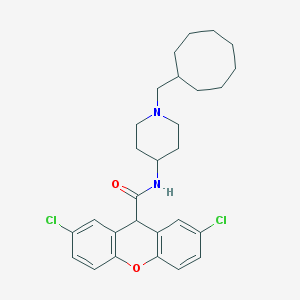 CHEMBL33278 CHEMBL33278 | C28H34Cl2N2O2 | 501.492 | 3 / 1 | 7.7 | No |
| 152804 | 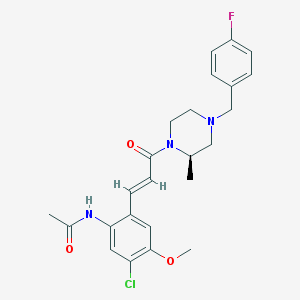 CHEMBL200629 CHEMBL200629 | C24H27ClFN3O3 | 459.946 | 5 / 1 | 3.4 | Yes |
| 158668 | 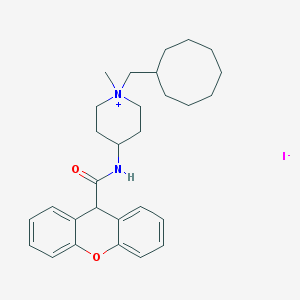 CHEMBL34165 CHEMBL34165 | C29H39IN2O2 | 574.547 | 3 / 1 | N/A | No |
| 165441 | 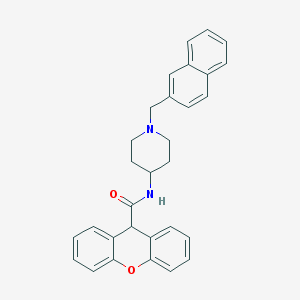 CHEMBL32766 CHEMBL32766 | C30H28N2O2 | 448.566 | 3 / 1 | 5.8 | No |
| 167191 | 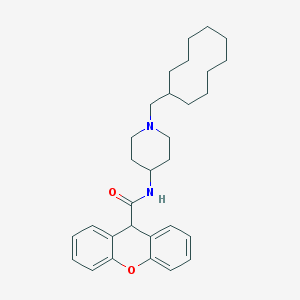 CHEMBL285703 CHEMBL285703 | C30H40N2O2 | 460.662 | 3 / 1 | 7.6 | No |
| 460747 | 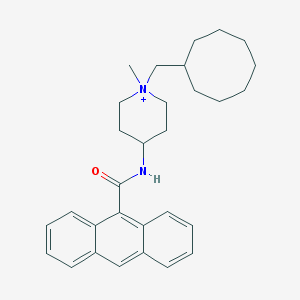 CHEMBL289682 CHEMBL289682 | C30H39N2O+ | 443.655 | 1 / 1 | 7.8 | No |
| 172063 | 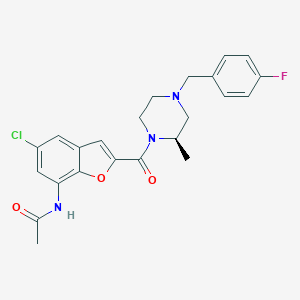 CHEMBL198966 CHEMBL198966 | C23H23ClFN3O3 | 443.903 | 5 / 1 | 3.8 | Yes |
| 173162 | 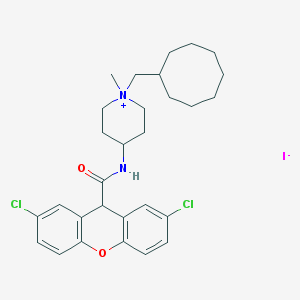 CHEMBL33929 CHEMBL33929 | C29H37Cl2IN2O2 | 643.431 | 3 / 1 | N/A | No |
| 173342 | 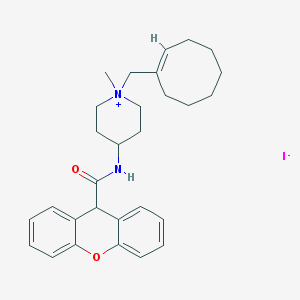 CHEMBL32257 CHEMBL32257 | C29H37IN2O2 | 572.531 | 3 / 1 | N/A | No |
| 176022 | 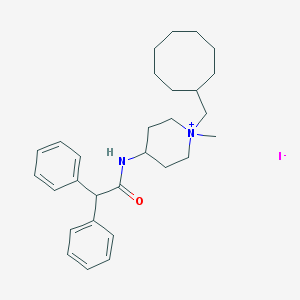 CHEMBL33761 CHEMBL33761 | C29H41IN2O | 560.564 | 2 / 1 | N/A | No |
| 183486 | 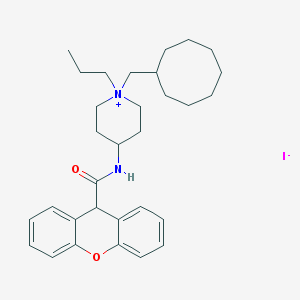 CHEMBL33614 CHEMBL33614 | C31H43IN2O2 | 602.601 | 3 / 1 | N/A | No |
| 195811 | 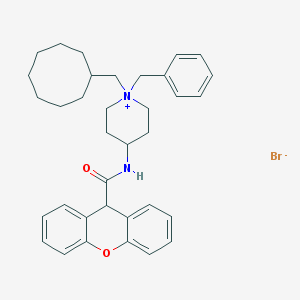 CHEMBL33586 CHEMBL33586 | C35H43BrN2O2 | 603.645 | 3 / 1 | N/A | No |
| 197279 | 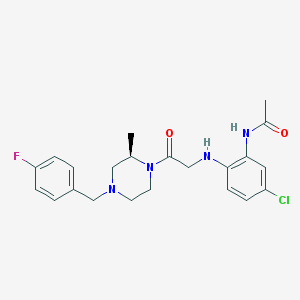 CHEMBL370162 CHEMBL370162 | C22H26ClFN4O2 | 432.924 | 5 / 2 | 3.1 | Yes |
| 198055 | 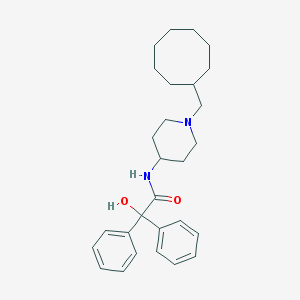 CHEMBL33159 CHEMBL33159 | C28H38N2O2 | 434.624 | 3 / 2 | 6.1 | No |
| 202879 | 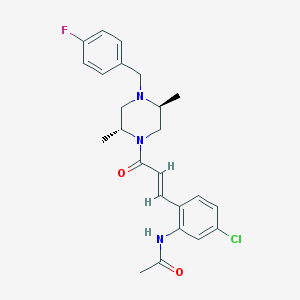 CHEMBL197375 CHEMBL197375 | C24H27ClFN3O2 | 443.947 | 4 / 1 | 3.9 | Yes |
| 202880 | 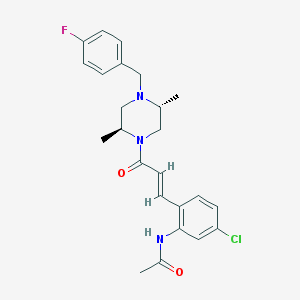 CHEMBL197345 CHEMBL197345 | C24H27ClFN3O2 | 443.947 | 4 / 1 | 3.9 | Yes |
| 203318 | 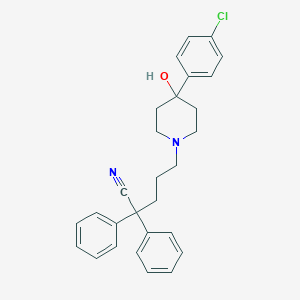 VUF-2274 VUF-2274 | C28H29ClN2O | 445.003 | 3 / 1 | 5.6 | No |
| 461010 | 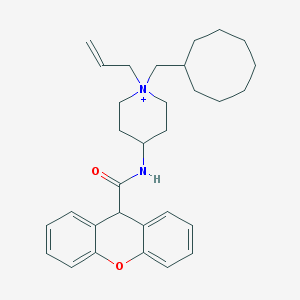 CHEMBL33104 CHEMBL33104 | C31H41N2O2+ | 473.681 | 2 / 1 | 7.1 | No |
| 213883 | 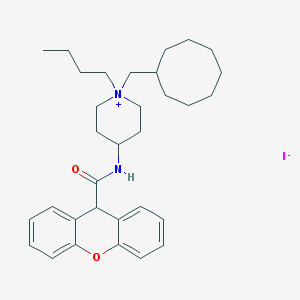 CHEMBL32713 CHEMBL32713 | C32H45IN2O2 | 616.628 | 3 / 1 | N/A | No |
| 215057 | 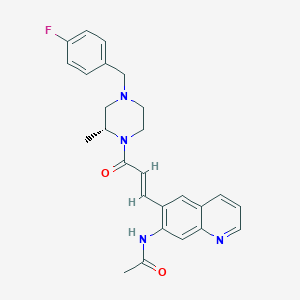 CHEMBL199412 CHEMBL199412 | C26H27FN4O2 | 446.526 | 5 / 1 | 3.1 | Yes |
| 224801 | 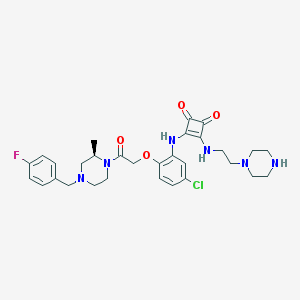 CHEMBL387608 CHEMBL387608 | C30H36ClFN6O4 | 599.104 | 10 / 3 | 2.9 | No |
| 461204 | 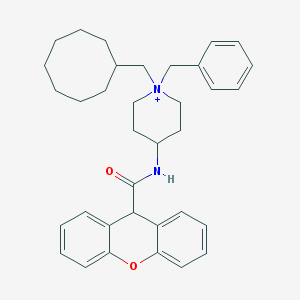 CHEMBL33586 CHEMBL33586 | C35H43N2O2+ | 523.741 | 2 / 1 | 8.0 | No |
| 238001 | 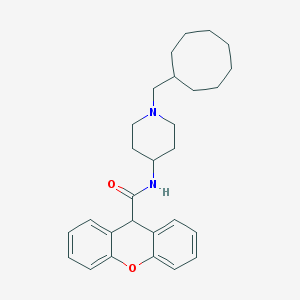 CHEMBL33074 CHEMBL33074 | C28H36N2O2 | 432.608 | 3 / 1 | 6.5 | No |
| 243802 | 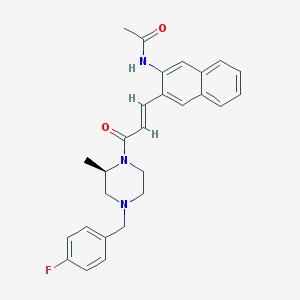 CHEMBL370596 CHEMBL370596 | C27H28FN3O2 | 445.538 | 4 / 1 | 4.1 | Yes |
| 461342 | 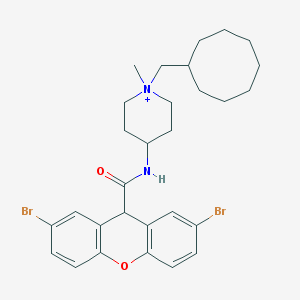 CHEMBL33223 CHEMBL33223 | C29H37Br2N2O2+ | 605.435 | 2 / 1 | 7.9 | No |
| 251968 | 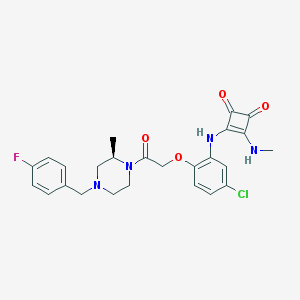 CHEMBL231829 CHEMBL231829 | C25H26ClFN4O4 | 500.955 | 8 / 2 | 3.5 | No |
| 254945 | 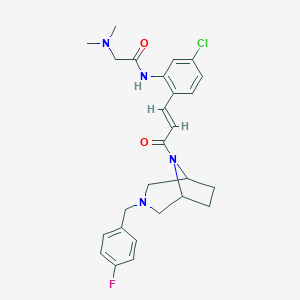 CHEMBL200242 CHEMBL200242 | C26H30ClFN4O2 | 485.0 | 5 / 1 | 3.7 | Yes |
| 256125 | 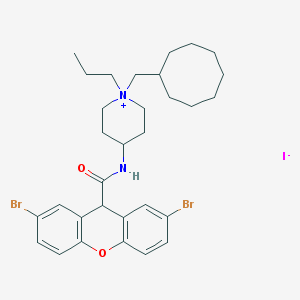 CHEMBL418589 CHEMBL418589 | C31H41Br2IN2O2 | 760.393 | 3 / 1 | N/A | No |
| 257183 | 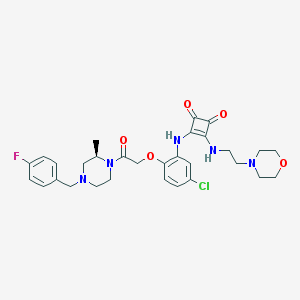 CHEMBL234295 CHEMBL234295 | C30H35ClFN5O5 | 600.088 | 10 / 2 | 3.1 | No |
| 268366 | 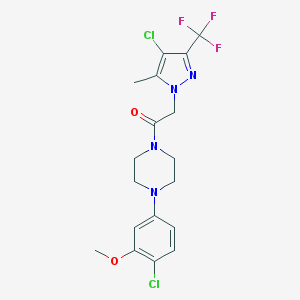 CHEMBL2322875 CHEMBL2322875 | C18H19Cl2F3N4O2 | 451.271 | 7 / 0 | 4.0 | Yes |
| 461565 | 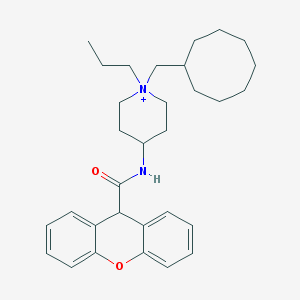 CHEMBL33614 CHEMBL33614 | C31H43N2O2+ | 475.697 | 2 / 1 | 7.4 | No |
| 275334 | 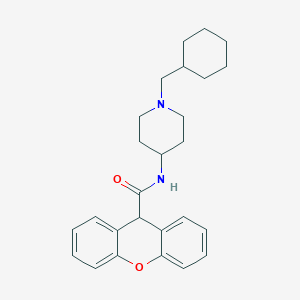 CHEMBL34053 CHEMBL34053 | C26H32N2O2 | 404.554 | 3 / 1 | 4.9 | Yes |
| 277261 | 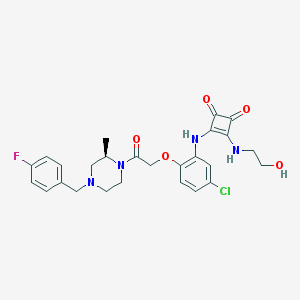 CHEMBL232638 CHEMBL232638 | C26H28ClFN4O5 | 530.981 | 9 / 3 | 2.8 | No |
| 461677 | 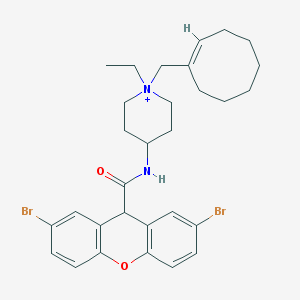 CHEMBL289902 CHEMBL289902 | C30H37Br2N2O2+ | 617.446 | 2 / 1 | 7.5 | No |
| 304073 | 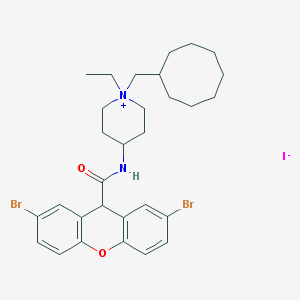 CHEMBL33340 CHEMBL33340 | C30H39Br2IN2O2 | 746.366 | 3 / 1 | N/A | No |
| 461913 | 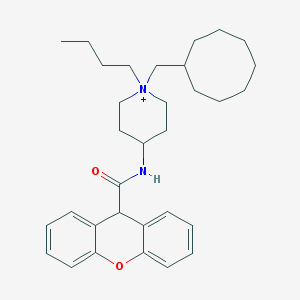 CHEMBL32713 CHEMBL32713 | C32H45N2O2+ | 489.724 | 2 / 1 | 7.8 | No |
| 461935 | 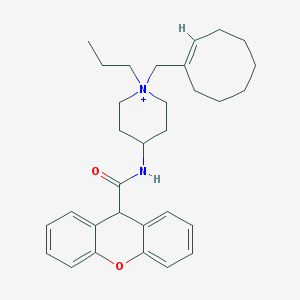 CHEMBL33050 CHEMBL33050 | C31H41N2O2+ | 473.681 | 2 / 1 | 6.6 | No |
| 318209 | 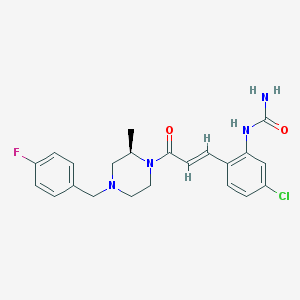 CHEMBL196860 CHEMBL196860 | C22H24ClFN4O2 | 430.908 | 4 / 2 | 2.9 | Yes |
| 324653 | 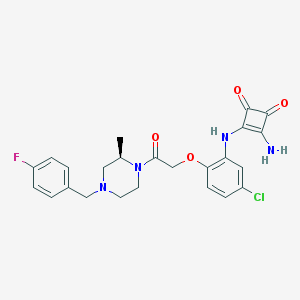 CHEMBL232441 CHEMBL232441 | C24H24ClFN4O4 | 486.928 | 8 / 2 | 2.8 | Yes |
| 330692 | 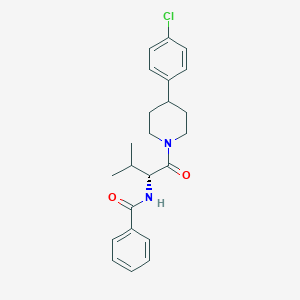 CHEMBL2180528 CHEMBL2180528 | C23H27ClN2O2 | 398.931 | 2 / 1 | 5.0 | Yes |
| 338256 | 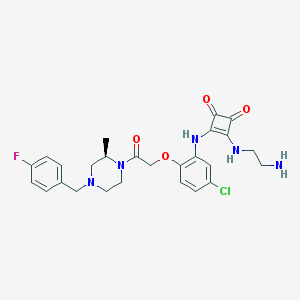 CHEMBL232837 CHEMBL232837 | C26H29ClFN5O4 | 529.997 | 9 / 3 | 2.5 | No |
| 347555 | 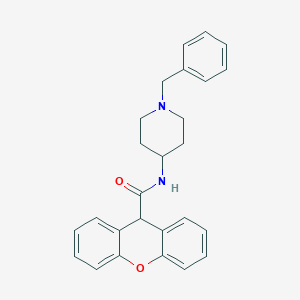 CHEMBL289464 CHEMBL289464 | C26H26N2O2 | 398.506 | 3 / 1 | 4.1 | Yes |
| 462285 | 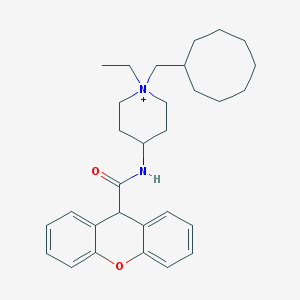 CHEMBL280751 CHEMBL280751 | C30H41N2O2+ | 461.67 | 2 / 1 | 6.9 | No |
| 362323 | 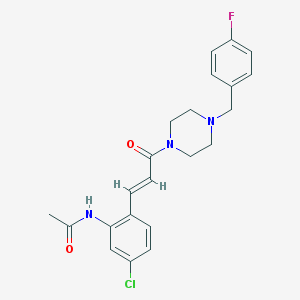 CHEMBL198852 CHEMBL198852 | C22H23ClFN3O2 | 415.893 | 4 / 1 | 3.0 | Yes |
| 391944 | 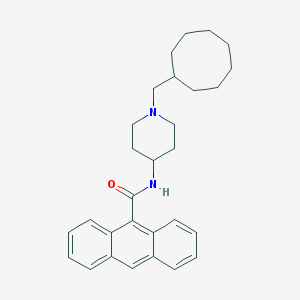 CHEMBL286701 CHEMBL286701 | C29H36N2O | 428.62 | 2 / 1 | 7.7 | No |
| 398741 | 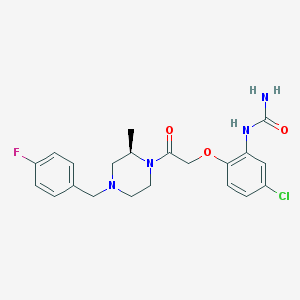 217645-70-0 217645-70-0 | C21H24ClFN4O3 | 434.896 | 5 / 2 | 2.5 | Yes |
| 398818 | 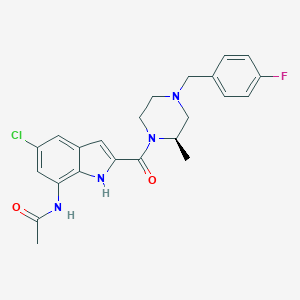 CHEMBL200794 CHEMBL200794 | C23H24ClFN4O2 | 442.919 | 4 / 2 | 3.5 | Yes |
| 462625 | 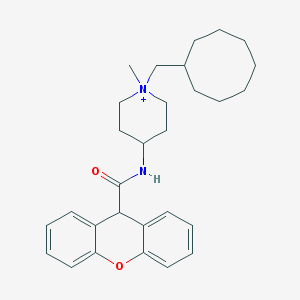 CHEMBL34165 CHEMBL34165 | C29H39N2O2+ | 447.643 | 2 / 1 | 6.5 | No |
| 420734 | 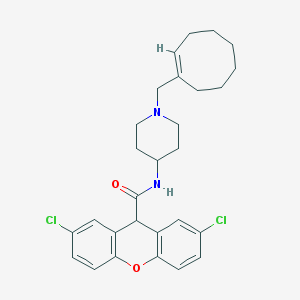 CHEMBL33639 CHEMBL33639 | C28H32Cl2N2O2 | 499.476 | 3 / 1 | 7.0 | No |
| 436146 | 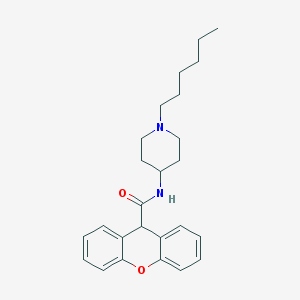 CHEMBL34226 CHEMBL34226 | C25H32N2O2 | 392.543 | 3 / 1 | 4.9 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218