

You can:
| Name | C-C chemokine receptor type 8 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | CCR8 |
| Synonym | GPRCY6 GPR-CY6 CMKBRL2 CKR-L1 ChemR1 [ Show all ] |
| Disease | Psoriasis |
| Length | 355 |
| Amino acid sequence | MDYTLDLSVTTVTDYYYPDIFSSPCDAELIQTNGKLLLAVFYCLLFVFSLLGNSLVILVLVVCKKLRSITDVYLLNLALSDLLFVFSFPFQTYYLLDQWVFGTVMCKVVSGFYYIGFYSSMFFITLMSVDRYLAVVHAVYALKVRTIRMGTTLCLAVWLTAIMATIPLLVFYQVASEDGVLQCYSFYNQQTLKWKIFTNFKMNILGLLIPFTIFMFCYIKILHQLKRCQNHNKTKAIRLVLIVVIASLLFWVPFNVVLFLTSLHSMHILDGCSISQQLTYATHVTEIISFTHCCVNPVIYAFVGEKFKKHLSEIFQKSCSQIFNYLGRQMPRESCEKSSSCQQHSSRSSSVDYIL |
| UniProt | P51685 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P51685 |
| 3D structure model | This predicted structure model is from GPCR-EXP P51685. |
| BioLiP | N/A |
| Therapeutic Target Database | T20575 |
| ChEMBL | CHEMBL4596 |
| IUPHAR | 65 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 3281 | 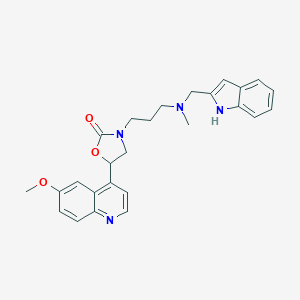 CHEMBL240249 CHEMBL240249 | C26H28N4O3 | 444.535 | 5 / 1 | 3.7 | Yes |
| 5473 | 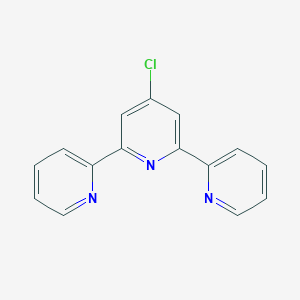 4'-Chloro-2,2':6',2''-terpyridine 4'-Chloro-2,2':6',2''-terpyridine | C15H10ClN3 | 267.716 | 3 / 0 | 3.0 | Yes |
| 6487 | 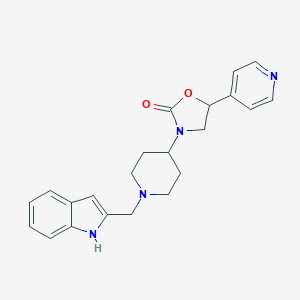 CHEMBL440956 CHEMBL440956 | C22H24N4O2 | 376.46 | 4 / 1 | 2.6 | Yes |
| 6631 | 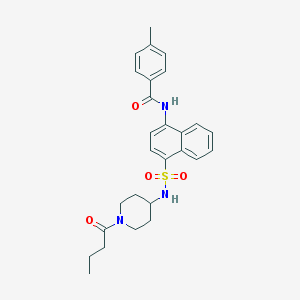 CHEMBL222062 CHEMBL222062 | C27H31N3O4S | 493.622 | 5 / 2 | 4.2 | Yes |
| 8631 | 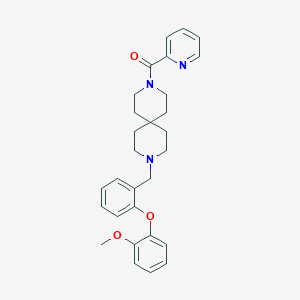 CHEMBL271127 CHEMBL271127 | C29H33N3O3 | 471.601 | 5 / 0 | 4.8 | Yes |
| 11956 | 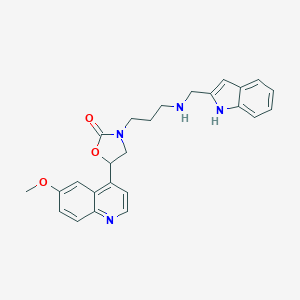 CHEMBL393411 CHEMBL393411 | C25H26N4O3 | 430.508 | 5 / 2 | 3.3 | Yes |
| 12101 | 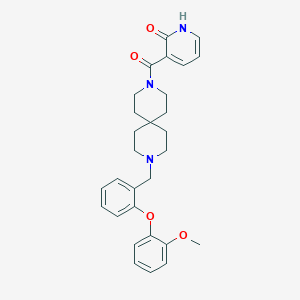 CHEMBL583843 CHEMBL583843 | C29H33N3O4 | 487.6 | 5 / 1 | 4.0 | Yes |
| 13932 | 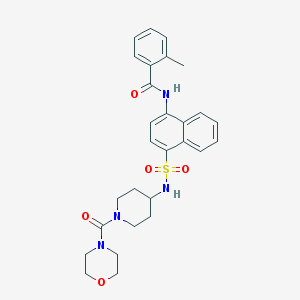 CHEMBL218218 CHEMBL218218 | C28H32N4O5S | 536.647 | 6 / 2 | 3.0 | No |
| 18700 | 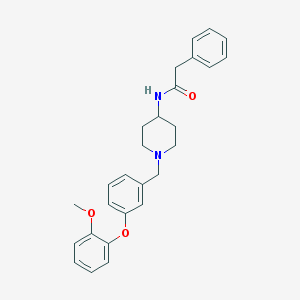 CHEMBL210322 CHEMBL210322 | C27H30N2O3 | 430.548 | 4 / 1 | 4.7 | Yes |
| 20399 | 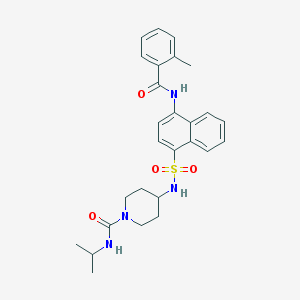 CHEMBL218350 CHEMBL218350 | C27H32N4O4S | 508.637 | 5 / 3 | 4.0 | No |
| 22002 | 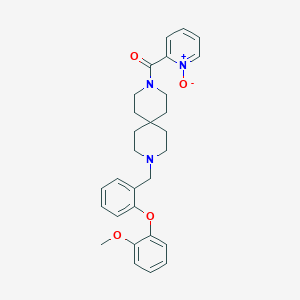 CHEMBL270969 CHEMBL270969 | C29H33N3O4 | 487.6 | 5 / 0 | 3.8 | Yes |
| 22787 | 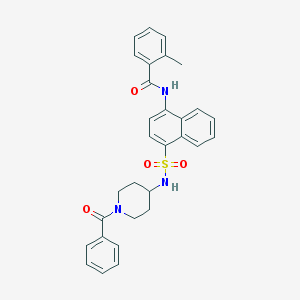 CHEMBL387451 CHEMBL387451 | C30H29N3O4S | 527.639 | 5 / 2 | 5.0 | No |
| 24240 | 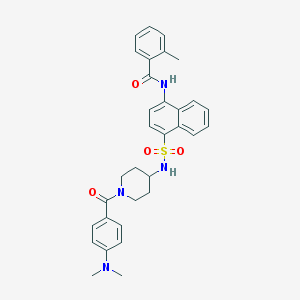 CHEMBL221078 CHEMBL221078 | C32H34N4O4S | 570.708 | 6 / 2 | 5.2 | No |
| 24443 | 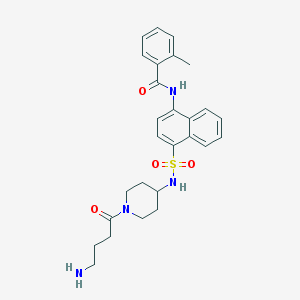 CHEMBL221867 CHEMBL221867 | C27H32N4O4S | 508.637 | 6 / 3 | 2.7 | No |
| 27676 | 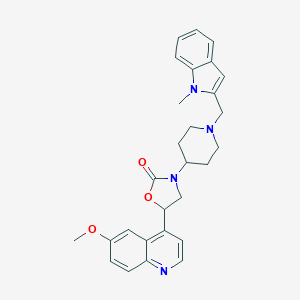 CHEMBL396102 CHEMBL396102 | C28H30N4O3 | 470.573 | 5 / 0 | 3.9 | Yes |
| 30950 | 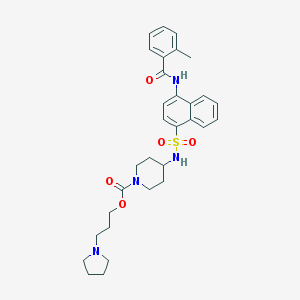 CHEMBL374527 CHEMBL374527 | C31H38N4O5S | 578.728 | 7 / 2 | 4.7 | No |
| 31774 | 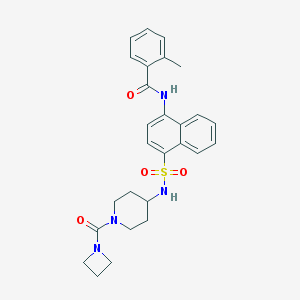 CHEMBL221130 CHEMBL221130 | C27H30N4O4S | 506.621 | 5 / 2 | 3.5 | No |
| 34464 | 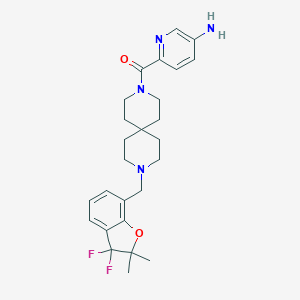 CHEMBL579072 CHEMBL579072 | C26H32F2N4O2 | 470.565 | 7 / 1 | 3.4 | Yes |
| 37254 | 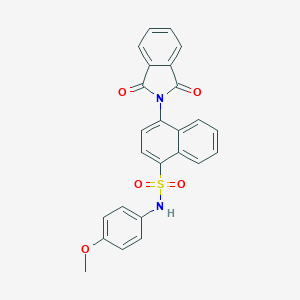 CHEMBL376069 CHEMBL376069 | C25H18N2O5S | 458.488 | 6 / 1 | 4.2 | Yes |
| 43277 | 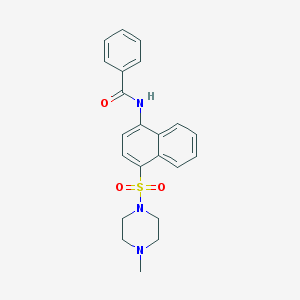 CHEMBL384534 CHEMBL384534 | C22H23N3O3S | 409.504 | 5 / 1 | 3.0 | Yes |
| 47361 | 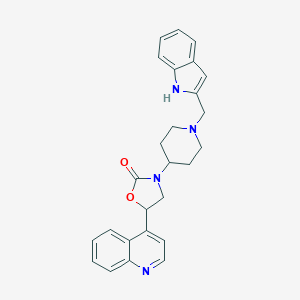 CHEMBL240462 CHEMBL240462 | C26H26N4O2 | 426.52 | 4 / 1 | 3.9 | Yes |
| 49579 | 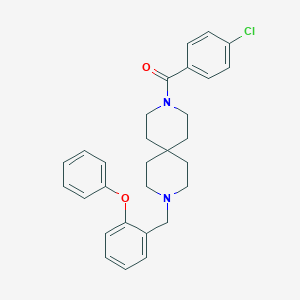 CHEMBL259243 CHEMBL259243 | C29H31ClN2O2 | 475.029 | 3 / 0 | 6.2 | No |
| 49634 | 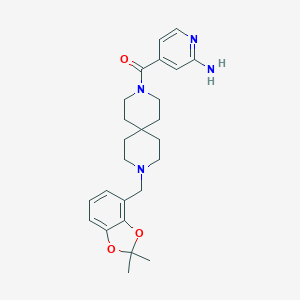 CHEMBL584087 CHEMBL584087 | C25H32N4O3 | 436.556 | 6 / 1 | 3.0 | Yes |
| 51226 | 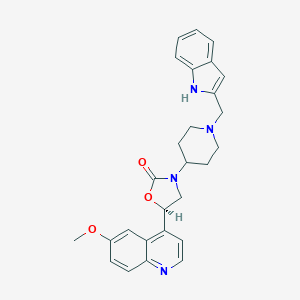 CHEMBL245568 CHEMBL245568 | C27H28N4O3 | 456.546 | 5 / 1 | 3.9 | Yes |
| 51227 | 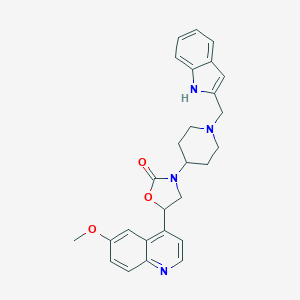 CHEMBL240848 CHEMBL240848 | C27H28N4O3 | 456.546 | 5 / 1 | 3.9 | Yes |
| 51349 | 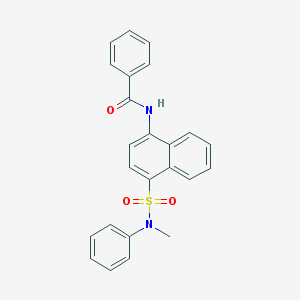 CHEMBL218502 CHEMBL218502 | C24H20N2O3S | 416.495 | 4 / 1 | 4.7 | Yes |
| 51378 | 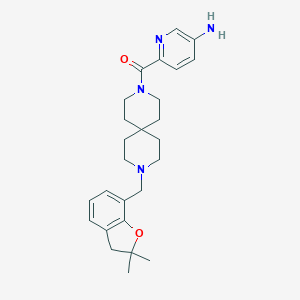 CHEMBL568523 CHEMBL568523 | C26H34N4O2 | 434.584 | 5 / 1 | 3.2 | Yes |
| 52621 | 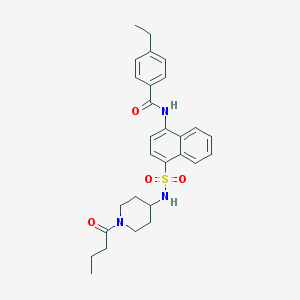 CHEMBL374978 CHEMBL374978 | C28H33N3O4S | 507.649 | 5 / 2 | 4.6 | No |
| 53018 | 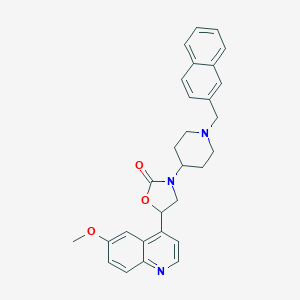 CHEMBL247762 CHEMBL247762 | C29H29N3O3 | 467.569 | 5 / 0 | 5.0 | Yes |
| 54007 | 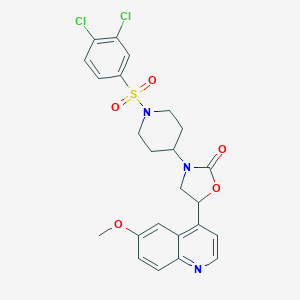 CHEMBL245365 CHEMBL245365 | C24H23Cl2N3O5S | 536.424 | 7 / 0 | 4.3 | No |
| 54488 | 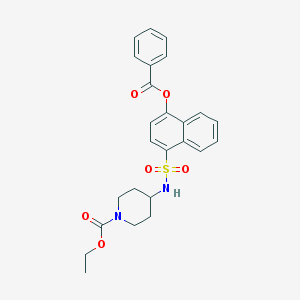 CHEMBL426207 CHEMBL426207 | C25H26N2O6S | 482.551 | 7 / 1 | 4.4 | Yes |
| 55813 | 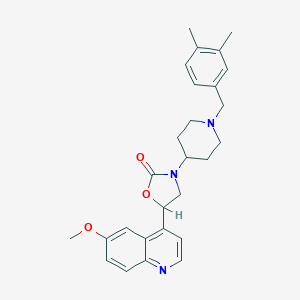 CHEMBL397395 CHEMBL397395 | C27H31N3O3 | 445.563 | 5 / 0 | 4.5 | Yes |
| 57457 | 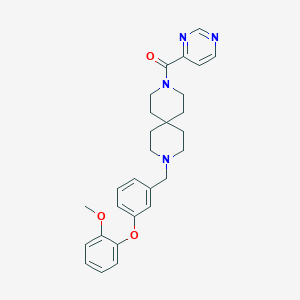 CHEMBL574655 CHEMBL574655 | C28H32N4O3 | 472.589 | 6 / 0 | 4.1 | Yes |
| 59141 | 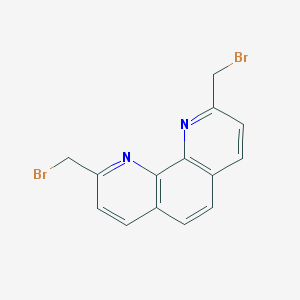 NSC602850 NSC602850 | C14H10Br2N2 | 366.056 | 2 / 0 | 3.6 | Yes |
| 59200 | 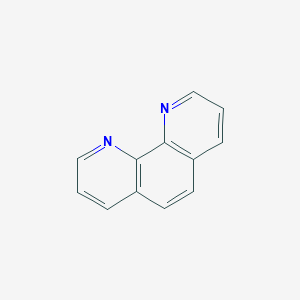 1,10-phenanthroline 1,10-phenanthroline | C12H8N2 | 180.21 | 2 / 0 | 1.8 | Yes |
| 60336 | 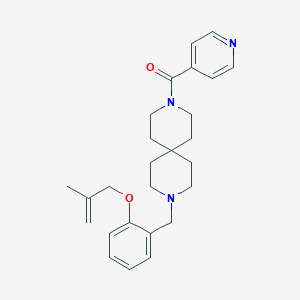 CHEMBL566759 CHEMBL566759 | C26H33N3O2 | 419.569 | 4 / 0 | 4.2 | Yes |
| 64828 | 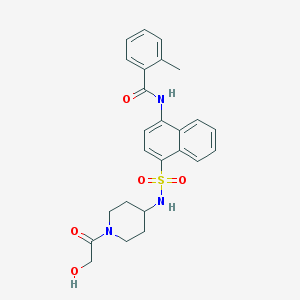 CHEMBL218036 CHEMBL218036 | C25H27N3O5S | 481.567 | 6 / 3 | 2.7 | Yes |
| 66925 | 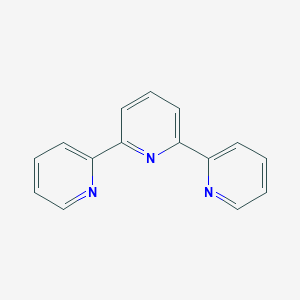 2,2':6',2''-Terpyridine 2,2':6',2''-Terpyridine | C15H11N3 | 233.274 | 3 / 0 | 2.4 | Yes |
| 70140 | 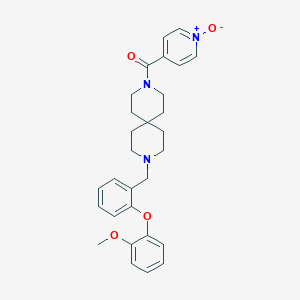 CHEMBL270971 CHEMBL270971 | C29H33N3O4 | 487.6 | 5 / 0 | 3.4 | Yes |
| 70717 | 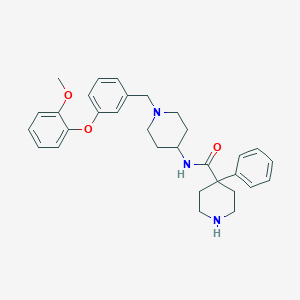 CHEMBL205447 CHEMBL205447 | C31H37N3O3 | 499.655 | 5 / 2 | 4.6 | Yes |
| 84597 | 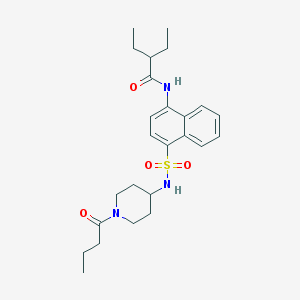 CHEMBL385654 CHEMBL385654 | C25H35N3O4S | 473.632 | 5 / 2 | 3.9 | Yes |
| 84777 | 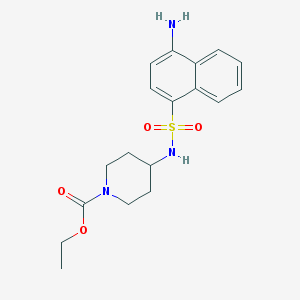 CHEMBL374916 CHEMBL374916 | C18H23N3O4S | 377.459 | 6 / 2 | 2.3 | Yes |
| 87444 | 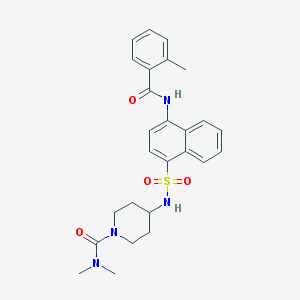 CHEMBL376910 CHEMBL376910 | C26H30N4O4S | 494.61 | 5 / 2 | 3.4 | Yes |
| 87688 | 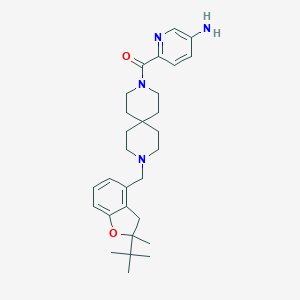 CHEMBL568294 CHEMBL568294 | C29H40N4O2 | 476.665 | 5 / 1 | 4.6 | Yes |
| 88190 | 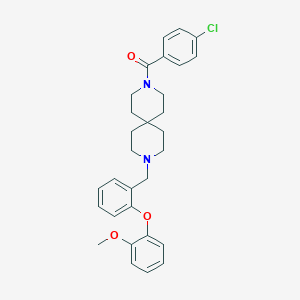 CHEMBL409224 CHEMBL409224 | C30H33ClN2O3 | 505.055 | 4 / 0 | 6.1 | No |
| 94136 | 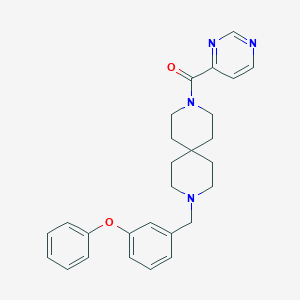 CHEMBL410885 CHEMBL410885 | C27H30N4O2 | 442.563 | 5 / 0 | 4.1 | Yes |
| 98566 | 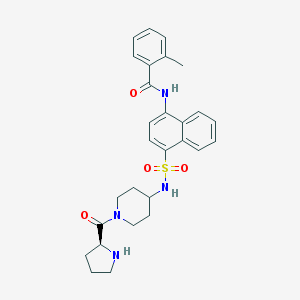 CHEMBL386308 CHEMBL386308 | C28H32N4O4S | 520.648 | 6 / 3 | 3.5 | No |
| 99376 | 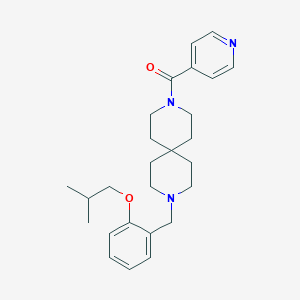 CHEMBL567419 CHEMBL567419 | C26H35N3O2 | 421.585 | 4 / 0 | 4.2 | Yes |
| 99780 | 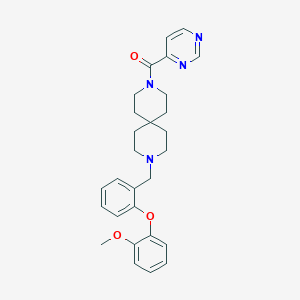 CHEMBL573216 CHEMBL573216 | C28H32N4O3 | 472.589 | 6 / 0 | 4.1 | Yes |
| 105123 | 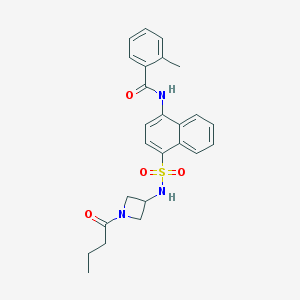 CHEMBL217979 CHEMBL217979 | C25H27N3O4S | 465.568 | 5 / 2 | 3.5 | Yes |
| 106833 | 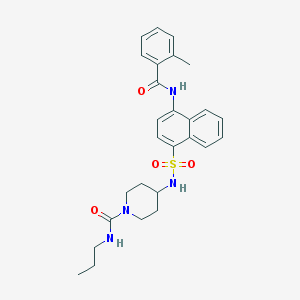 CHEMBL385288 CHEMBL385288 | C27H32N4O4S | 508.637 | 5 / 3 | 4.1 | No |
| 109351 | 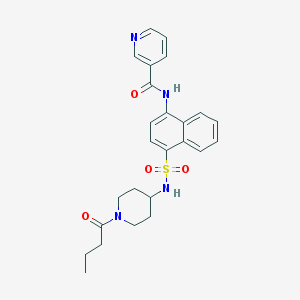 CHEMBL218555 CHEMBL218555 | C25H28N4O4S | 480.583 | 6 / 2 | 2.8 | Yes |
| 111354 | 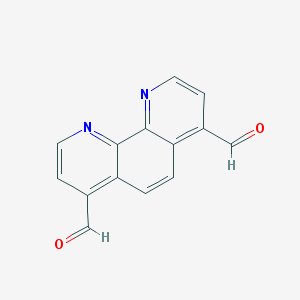 1,10-phenanthroline-4,7-dicarbaldehyde 1,10-phenanthroline-4,7-dicarbaldehyde | C14H8N2O2 | 236.23 | 4 / 0 | 1.4 | Yes |
| 112773 | 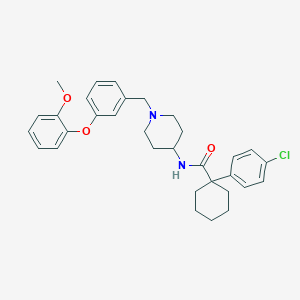 CHEMBL205457 CHEMBL205457 | C32H37ClN2O3 | 533.109 | 4 / 1 | 7.3 | No |
| 120237 | 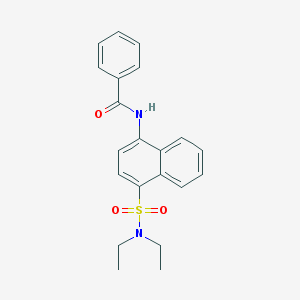 CHEMBL376846 CHEMBL376846 | C21H22N2O3S | 382.478 | 4 / 1 | 3.9 | Yes |
| 122259 | 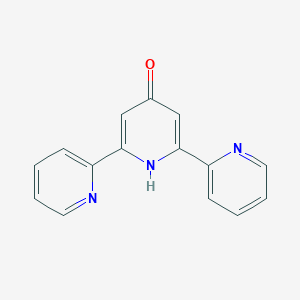 128143-88-4 128143-88-4 | C15H11N3O | 249.273 | 4 / 1 | 1.3 | Yes |
| 125451 | 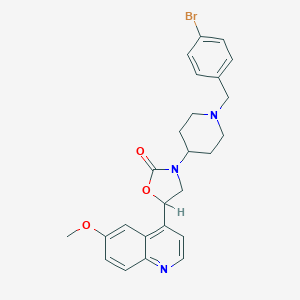 CHEMBL395424 CHEMBL395424 | C25H26BrN3O3 | 496.405 | 5 / 0 | 4.4 | Yes |
| 127586 | 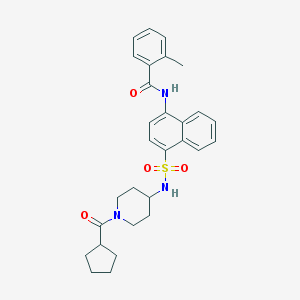 CHEMBL219433 CHEMBL219433 | C29H33N3O4S | 519.66 | 5 / 2 | 4.9 | No |
| 129640 | 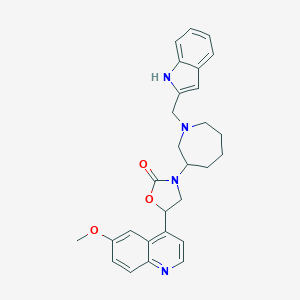 CHEMBL393721 CHEMBL393721 | C28H30N4O3 | 470.573 | 5 / 1 | 4.3 | Yes |
| 129749 | 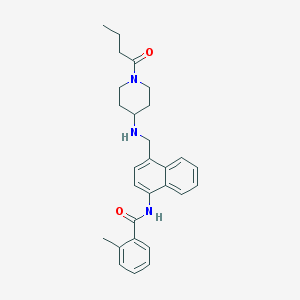 CHEMBL221841 CHEMBL221841 | C28H33N3O2 | 443.591 | 3 / 2 | 4.6 | Yes |
| 131400 | 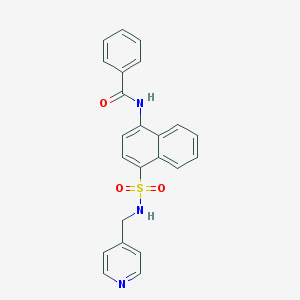 CHEMBL385287 CHEMBL385287 | C23H19N3O3S | 417.483 | 5 / 2 | 3.4 | Yes |
| 131423 | 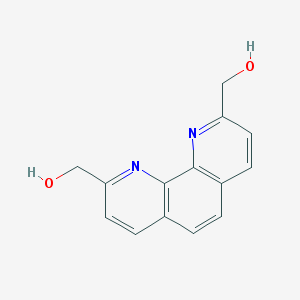 78831-36-4 78831-36-4 | C14H12N2O2 | 240.262 | 4 / 2 | 0.8 | Yes |
| 134057 | 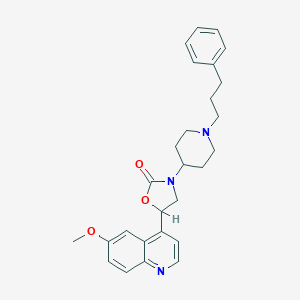 CHEMBL247965 CHEMBL247965 | C27H31N3O3 | 445.563 | 5 / 0 | 4.6 | Yes |
| 135394 | 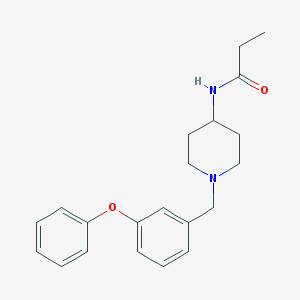 CHEMBL381354 CHEMBL381354 | C21H26N2O2 | 338.451 | 3 / 1 | 3.6 | Yes |
| 136566 | 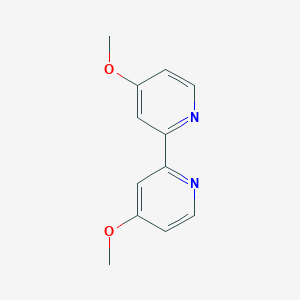 4,4'-Dimethoxy-2,2'-bipyridine 4,4'-Dimethoxy-2,2'-bipyridine | C12H12N2O2 | 216.24 | 4 / 0 | 1.4 | Yes |
| 136962 | 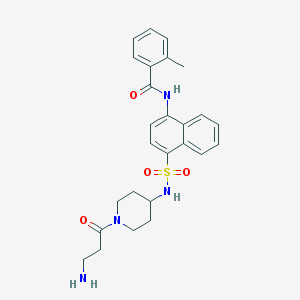 CHEMBL220703 CHEMBL220703 | C26H30N4O4S | 494.61 | 6 / 3 | 2.4 | Yes |
| 137652 | 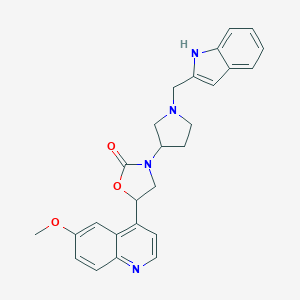 CHEMBL240460 CHEMBL240460 | C26H26N4O3 | 442.519 | 5 / 1 | 3.6 | Yes |
| 139866 | 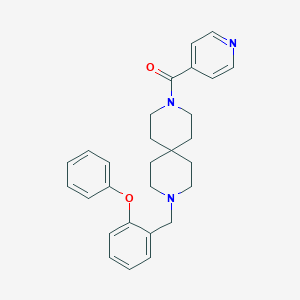 CHEMBL271768 CHEMBL271768 | C28H31N3O2 | 441.575 | 4 / 0 | 4.5 | Yes |
| 139968 | 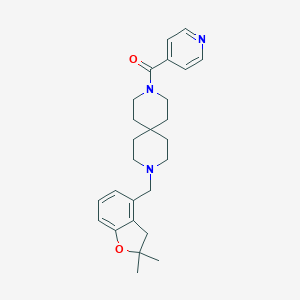 CHEMBL583621 CHEMBL583621 | C26H33N3O2 | 419.569 | 4 / 0 | 3.6 | Yes |
| 143238 | 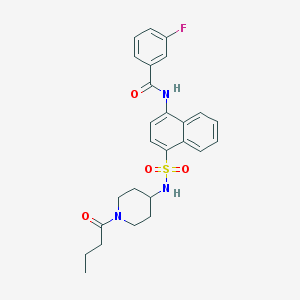 CHEMBL221639 CHEMBL221639 | C26H28FN3O4S | 497.585 | 6 / 2 | 4.0 | Yes |
| 144282 | 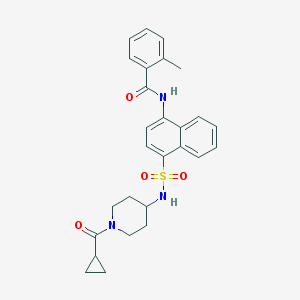 CHEMBL374979 CHEMBL374979 | C27H29N3O4S | 491.606 | 5 / 2 | 3.8 | Yes |
| 144951 | 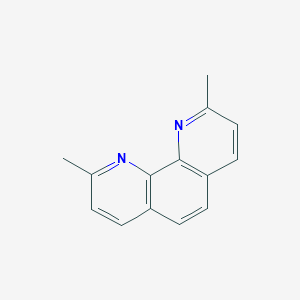 Neocuproine Neocuproine | C14H12N2 | 208.264 | 2 / 0 | 3.3 | Yes |
| 145205 | 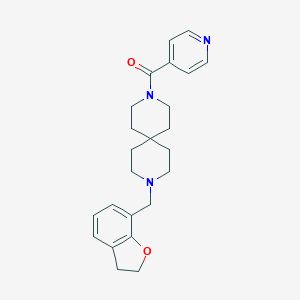 CHEMBL572537 CHEMBL572537 | C24H29N3O2 | 391.515 | 4 / 0 | 2.9 | Yes |
| 149889 | 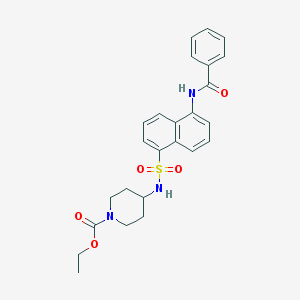 CHEMBL218037 CHEMBL218037 | C25H27N3O5S | 481.567 | 6 / 2 | 3.8 | Yes |
| 151984 | 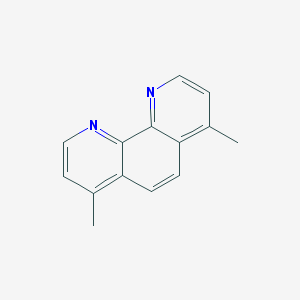 4,7-Dimethyl-1,10-phenanthroline 4,7-Dimethyl-1,10-phenanthroline | C14H12N2 | 208.264 | 2 / 0 | 3.2 | Yes |
| 554058 | 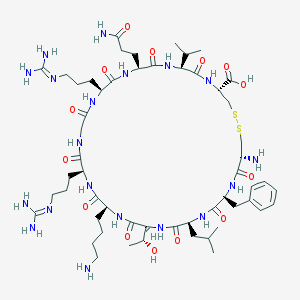 Viral macrophage inflammatory protein-II Viral macrophage inflammatory protein-II | C55H93N19O14S2 | 1308.59 | 20 / 19 | -6.2 | No |
| 157307 | 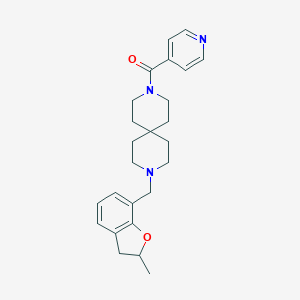 CHEMBL568296 CHEMBL568296 | C25H31N3O2 | 405.542 | 4 / 0 | 3.4 | Yes |
| 159400 | 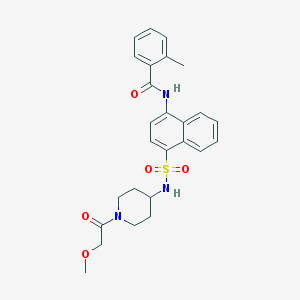 CHEMBL221961 CHEMBL221961 | C26H29N3O5S | 495.594 | 6 / 2 | 3.3 | Yes |
| 166309 | 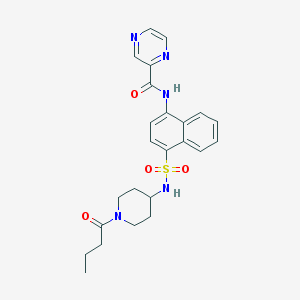 CHEMBL222112 CHEMBL222112 | C24H27N5O4S | 481.571 | 7 / 2 | 2.0 | Yes |
| 169521 | 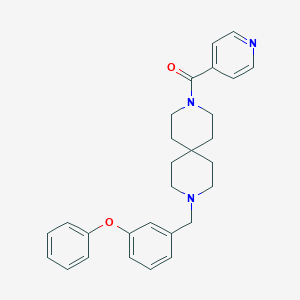 CHEMBL271801 CHEMBL271801 | C28H31N3O2 | 441.575 | 4 / 0 | 4.5 | Yes |
| 169553 | 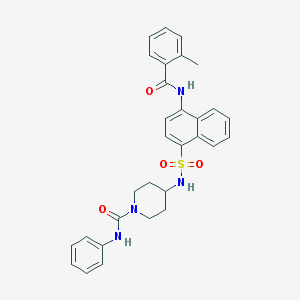 CHEMBL387161 CHEMBL387161 | C30H30N4O4S | 542.654 | 5 / 3 | 4.8 | No |
| 171865 | 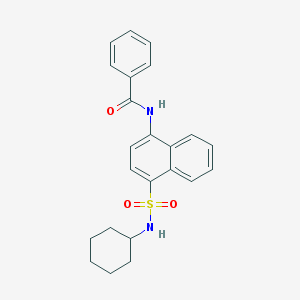 CHEMBL218503 CHEMBL218503 | C23H24N2O3S | 408.516 | 4 / 2 | 4.8 | Yes |
| 175508 | 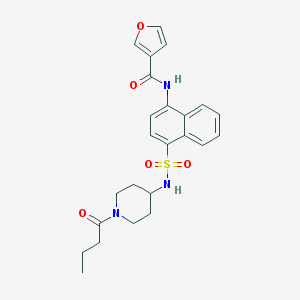 CHEMBL219432 CHEMBL219432 | C24H27N3O5S | 469.556 | 6 / 2 | 2.9 | Yes |
| 176848 | 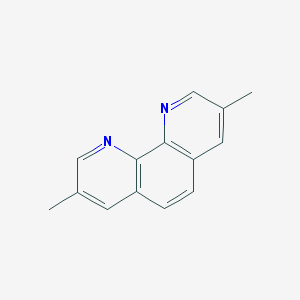 CHEMBL2205809 CHEMBL2205809 | C14H12N2 | 208.264 | 2 / 0 | 3.2 | Yes |
| 180583 | 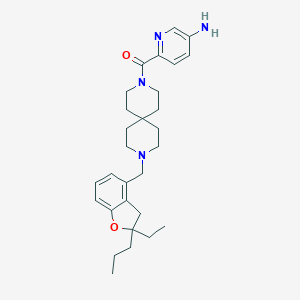 CHEMBL567417 CHEMBL567417 | C29H40N4O2 | 476.665 | 5 / 1 | 4.6 | Yes |
| 186589 | 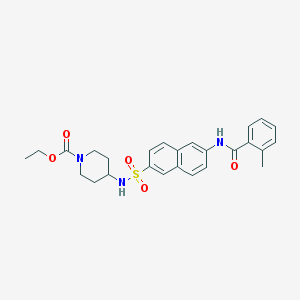 CHEMBL373893 CHEMBL373893 | C26H29N3O5S | 495.594 | 6 / 2 | 4.2 | Yes |
| 186756 | 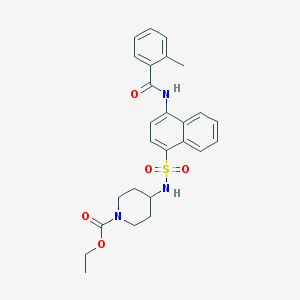 CHEMBL218375 CHEMBL218375 | C26H29N3O5S | 495.594 | 6 / 2 | 4.2 | Yes |
| 188563 | 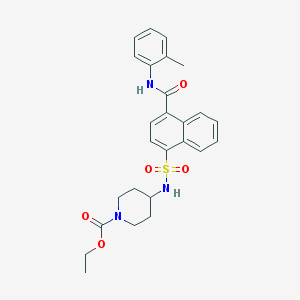 CHEMBL375315 CHEMBL375315 | C26H29N3O5S | 495.594 | 6 / 2 | 4.2 | Yes |
| 188877 | 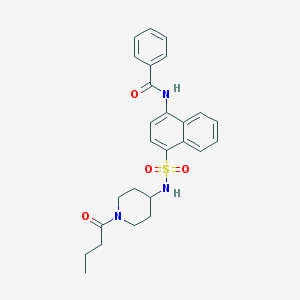 CHEMBL426551 CHEMBL426551 | C26H29N3O4S | 479.595 | 5 / 2 | 3.9 | Yes |
| 189047 | 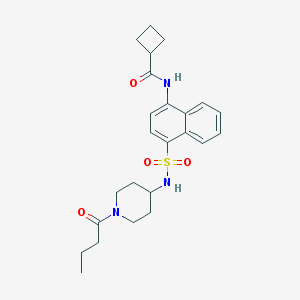 CHEMBL218554 CHEMBL218554 | C24H31N3O4S | 457.589 | 5 / 2 | 3.2 | Yes |
| 190170 | 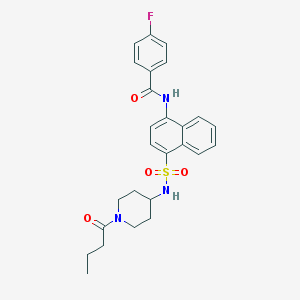 CHEMBL221978 CHEMBL221978 | C26H28FN3O4S | 497.585 | 6 / 2 | 4.0 | Yes |
| 190297 | 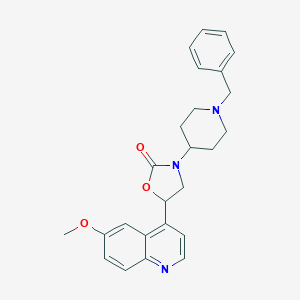 CHEMBL247964 CHEMBL247964 | C25H27N3O3 | 417.509 | 5 / 0 | 3.8 | Yes |
| 190745 | 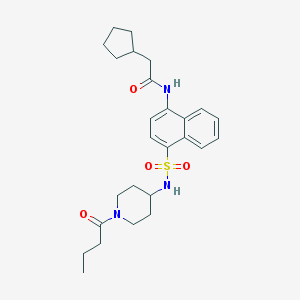 CHEMBL220650 CHEMBL220650 | C26H35N3O4S | 485.643 | 5 / 2 | 4.3 | Yes |
| 191175 | 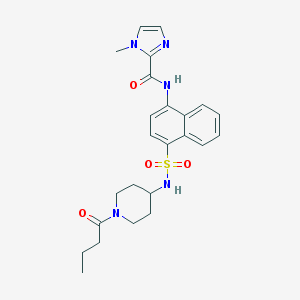 CHEMBL221842 CHEMBL221842 | C24H29N5O4S | 483.587 | 6 / 2 | 2.3 | Yes |
| 192188 | 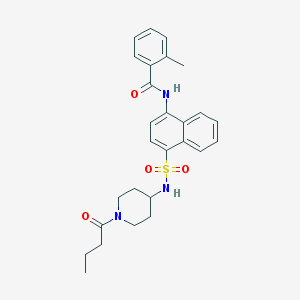 CHEMBL374939 CHEMBL374939 | C27H31N3O4S | 493.622 | 5 / 2 | 4.2 | Yes |
| 193958 | 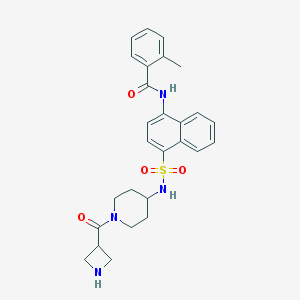 CHEMBL375854 CHEMBL375854 | C27H30N4O4S | 506.621 | 6 / 3 | 2.7 | No |
| 195059 | 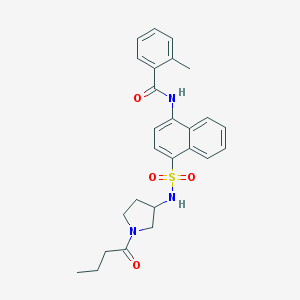 CHEMBL219327 CHEMBL219327 | C26H29N3O4S | 479.595 | 5 / 2 | 3.9 | Yes |
| 198272 | 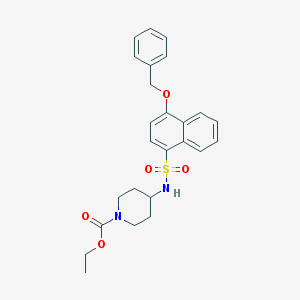 CHEMBL221789 CHEMBL221789 | C25H28N2O5S | 468.568 | 6 / 1 | 4.4 | Yes |
| 200301 | 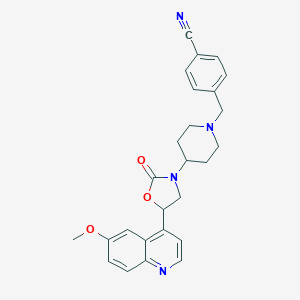 CHEMBL395423 CHEMBL395423 | C26H26N4O3 | 442.519 | 6 / 0 | 3.5 | Yes |
| 216981 | 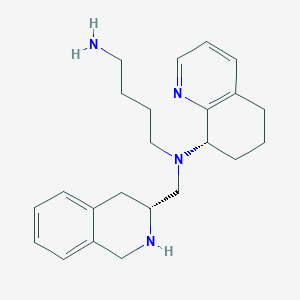 CHEMBL3091687 CHEMBL3091687 | C23H32N4 | 364.537 | 4 / 2 | 2.4 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218