

You can:
| Name | Proteinase-activated receptor 2 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | F2RL1 |
| Synonym | Protease-activated receptor-2 PAR2 PAR-2 GPR11 G-protein coupled receptor 11 [ Show all ] |
| Disease | N/A |
| Length | 397 |
| Amino acid sequence | MRSPSAAWLLGAAILLAASLSCSGTIQGTNRSSKGRSLIGKVDGTSHVTGKGVTVETVFSVDEFSASVLTGKLTTVFLPIVYTIVFVVGLPSNGMALWVFLFRTKKKHPAVIYMANLALADLLSVIWFPLKIAYHIHGNNWIYGEALCNVLIGFFYGNMYCSILFMTCLSVQRYWVIVNPMGHSRKKANIAIGISLAIWLLILLVTIPLYVVKQTIFIPALNITTCHDVLPEQLLVGDMFNYFLSLAIGVFLFPAFLTASAYVLMIRMLRSSAMDENSEKKRKRAIKLIVTVLAMYLICFTPSNLLLVVHYFLIKSQGQSHVYALYIVALCLSTLNSCIDPFVYYFVSHDFRDHAKNALLCRSVRTVKQMQVSLTSKKHSRKSSSYSSSSTTVKTSY |
| UniProt | P55085 |
| Protein Data Bank | 5ndz, 5ndd |
| GPCR-HGmod model | P55085 |
| 3D structure model | This structure is from PDB ID 5ndz. |
| BioLiP | BL0377325, BL0377326 |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL5963 |
| IUPHAR | 348 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 2177 | 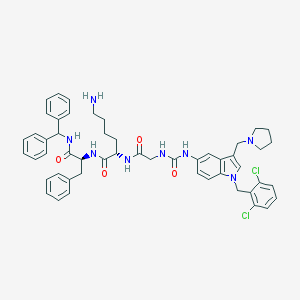 CHEMBL2431727 CHEMBL2431727 | C51H56Cl2N8O4 | 915.961 | 6 / 6 | 7.6 | No |
| 521511 | 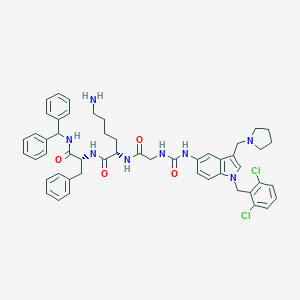 CHEMBL3735057 CHEMBL3735057 | C51H56Cl2N8O4 | 915.961 | 6 / 6 | 7.6 | No |
| 535984 | 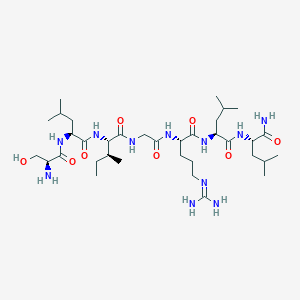 CHEMBL236457 CHEMBL236457 | C35H67N11O8 | 769.99 | 10 / 11 | -0.3 | No |
| 521548 | 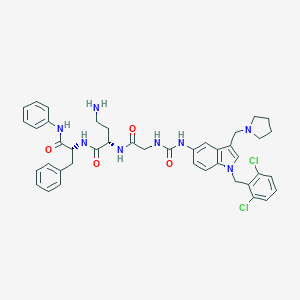 CHEMBL3735405 CHEMBL3735405 | C42H46Cl2N8O4 | 797.782 | 6 / 6 | 5.2 | No |
| 536026 | 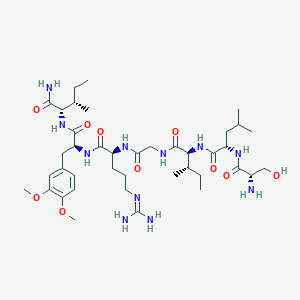 CHEMBL398099 CHEMBL398099 | C40H69N11O10 | 864.059 | 12 / 11 | 0.0 | No |
| 5495 | 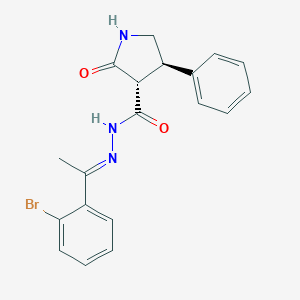 CHEMBL493494 CHEMBL493494 | C19H18BrN3O2 | 400.276 | 3 / 2 | 3.6 | Yes |
| 536110 | 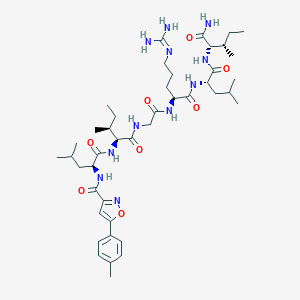 CHEMBL236194 CHEMBL236194 | C43H69N11O8 | 868.094 | 10 / 9 | 3.8 | No |
| 9075 | 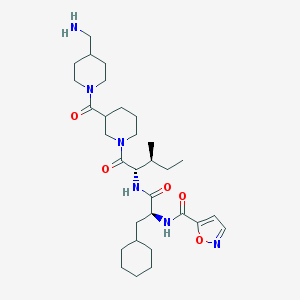 CHEMBL1269138 CHEMBL1269138 | C31H50N6O5 | 586.778 | 7 / 3 | 3.5 | No |
| 548032 | 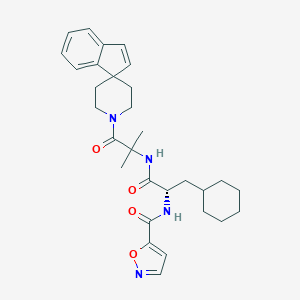 CHEMBL3924586 CHEMBL3924586 | C30H38N4O4 | 518.658 | 5 / 2 | 5.3 | No |
| 16051 | 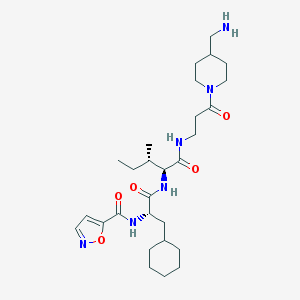 CHEMBL1271444 CHEMBL1271444 | C28H46N6O5 | 546.713 | 7 / 4 | 2.8 | No |
| 522007 | 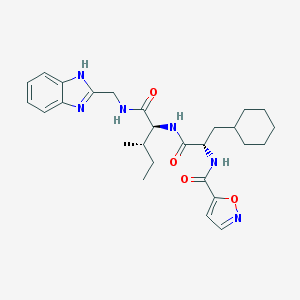 CHEMBL3752716 CHEMBL3752716 | C27H36N6O4 | 508.623 | 6 / 4 | 4.6 | No |
| 548099 | 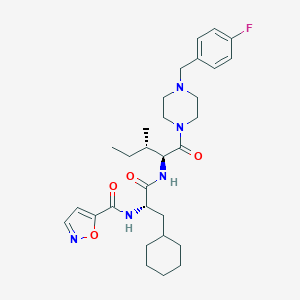 CHEMBL3895574 CHEMBL3895574 | C30H42FN5O4 | 555.695 | 7 / 2 | 5.1 | No |
| 536466 | 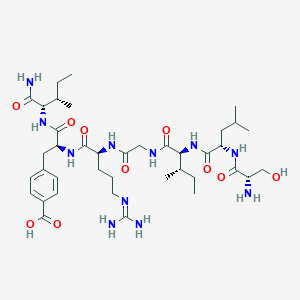 CHEMBL395357 CHEMBL395357 | C39H65N11O10 | 848.016 | 12 / 12 | -2.7 | No |
| 22132 | 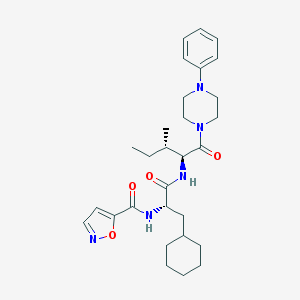 CHEMBL1270427 CHEMBL1270427 | C29H41N5O4 | 523.678 | 6 / 2 | 5.3 | No |
| 23446 | 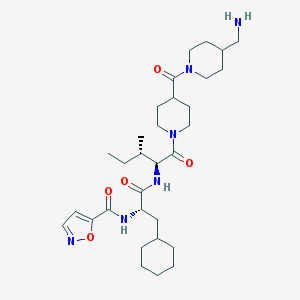 CHEMBL1269134 CHEMBL1269134 | C31H50N6O5 | 586.778 | 7 / 3 | 3.5 | No |
| 536635 | 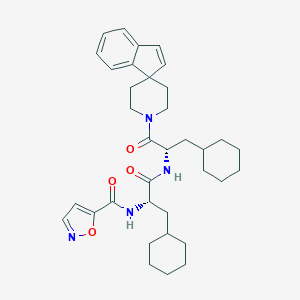 CHEMBL3902619 CHEMBL3902619 | C35H46N4O4 | 586.777 | 5 / 2 | 7.9 | No |
| 25438 | 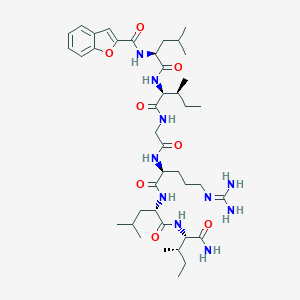 CHEMBL236292 CHEMBL236292 | C41H66N10O8 | 827.041 | 9 / 9 | 3.7 | No |
| 30132 | 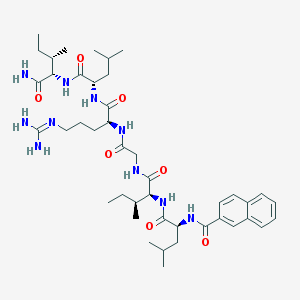 CHEMBL236293 CHEMBL236293 | C43H68N10O7 | 837.08 | 8 / 9 | 4.2 | No |
| 30631 | 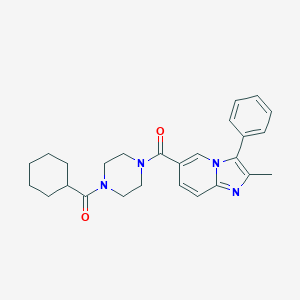 CHEMBL2431715 CHEMBL2431715 | C26H30N4O2 | 430.552 | 3 / 0 | 4.6 | Yes |
| 30796 | 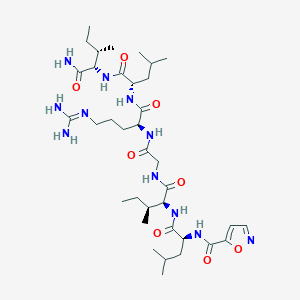 CHEMBL238396 CHEMBL238396 | C36H63N11O8 | 777.969 | 10 / 9 | 1.7 | No |
| 536855 | 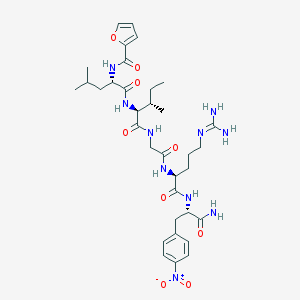 CHEMBL410365 CHEMBL410365 | C34H50N10O9 | 742.835 | 10 / 8 | 1.4 | No |
| 522749 | 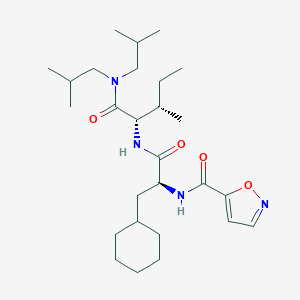 CHEMBL3754768 CHEMBL3754768 | C27H46N4O4 | 490.689 | 5 / 2 | 6.4 | No |
| 46442 | 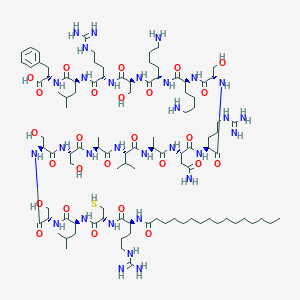 CID 73353878 CID 73353878 | C100H178N30O26S | 2248.77 | 32 / 37 | -4.3 | No |
| 522884 | 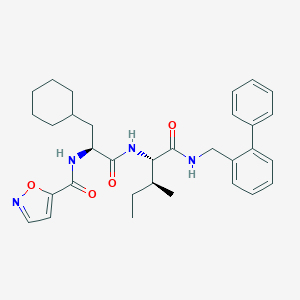 CHEMBL3751999 CHEMBL3751999 | C32H40N4O4 | 544.696 | 5 / 3 | 6.6 | No |
| 522925 | 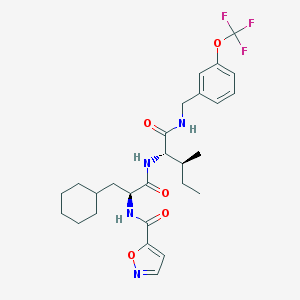 CHEMBL3752901 CHEMBL3752901 | C27H35F3N4O5 | 552.595 | 9 / 3 | 6.2 | No |
| 548468 | 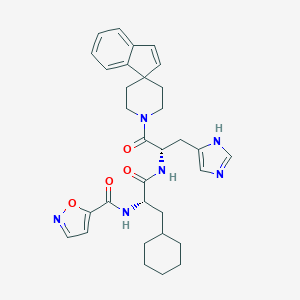 CHEMBL3919505 CHEMBL3919505 | C32H38N6O4 | 570.694 | 6 / 3 | 4.9 | No |
| 51508 | 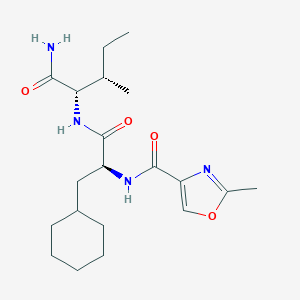 CHEMBL1271236 CHEMBL1271236 | C20H32N4O4 | 392.5 | 5 / 3 | 3.5 | Yes |
| 51862 | 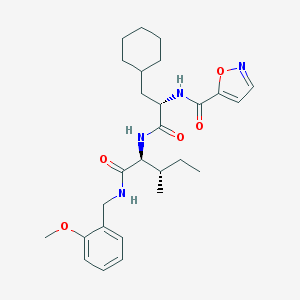 CHEMBL2431717 CHEMBL2431717 | C27H38N4O5 | 498.624 | 6 / 3 | 5.0 | Yes |
| 537360 | 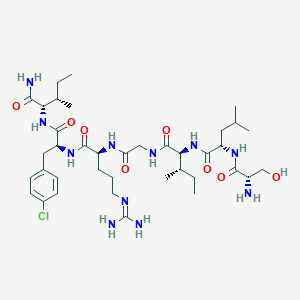 CHEMBL396184 CHEMBL396184 | C38H64ClN11O8 | 838.449 | 10 / 11 | -0.6 | No |
| 56734 | 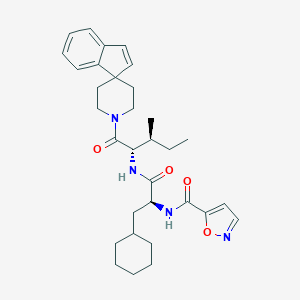 CHEMBL2431617 CHEMBL2431617 | C32H42N4O4 | 546.712 | 5 / 2 | 6.5 | No |
| 553519 | 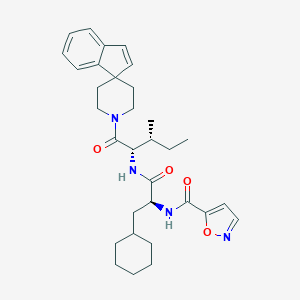 GB88 GB88 | C32H42N4O4 | 546.712 | 5 / 2 | 6.5 | No |
| 58340 | 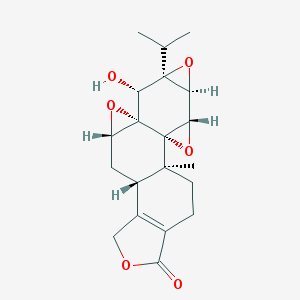 triptolide triptolide | C20H24O6 | 360.406 | 6 / 1 | 0.2 | Yes |
| 60417 | 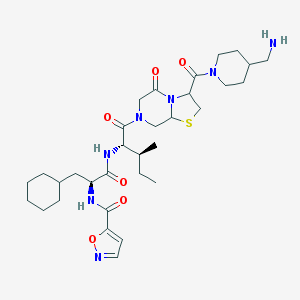 CHEMBL1269141 CHEMBL1269141 | C32H49N7O6S | 659.847 | 9 / 3 | 3.0 | No |
| 444257 | 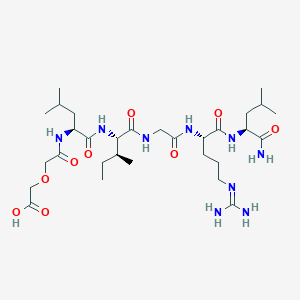 CHEMBL1689570 CHEMBL1689570 | C30H55N9O9 | 685.824 | 10 / 9 | -0.3 | No |
| 67377 | 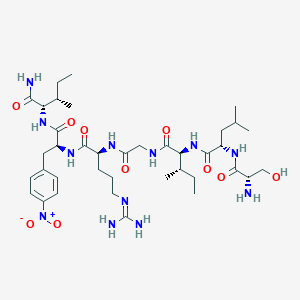 CHEMBL396183 CHEMBL396183 | C38H64N12O10 | 849.004 | 12 / 11 | -0.2 | No |
| 548687 | 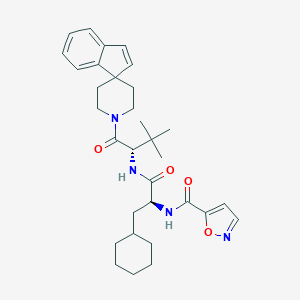 CHEMBL3897838 CHEMBL3897838 | C32H42N4O4 | 546.712 | 5 / 2 | 6.5 | No |
| 75658 | 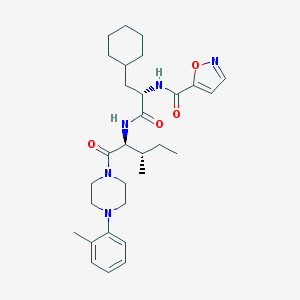 CHEMBL1270521 CHEMBL1270521 | C30H43N5O4 | 537.705 | 6 / 2 | 5.7 | No |
| 76201 | 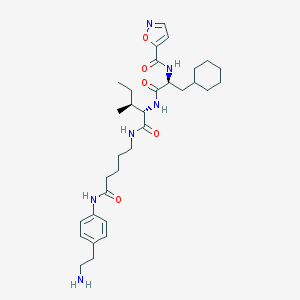 CHEMBL1269132 CHEMBL1269132 | C32H48N6O5 | 596.773 | 7 / 5 | 4.4 | No |
| 523786 | 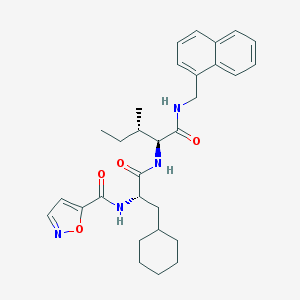 CHEMBL3753844 CHEMBL3753844 | C30H38N4O4 | 518.658 | 5 / 3 | 6.3 | No |
| 553665 | 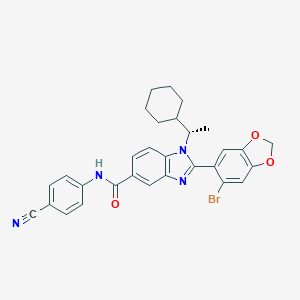 AZ3451 AZ3451 | C30H27BrN4O3 | 571.475 | 5 / 1 | 6.9 | No |
| 538027 | 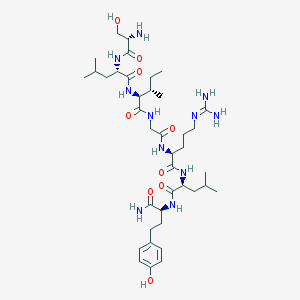 CHEMBL237961 CHEMBL237961 | C39H67N11O9 | 834.033 | 11 / 12 | 0.0 | No |
| 538112 | 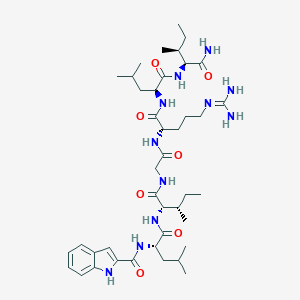 CHEMBL439563 CHEMBL439563 | C41H67N11O7 | 826.057 | 8 / 10 | 3.4 | No |
| 445148 | 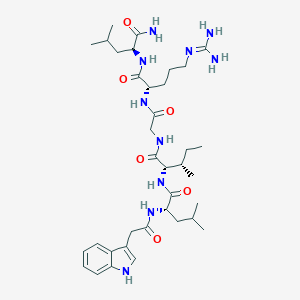 CHEMBL1689567 CHEMBL1689567 | C36H58N10O6 | 726.924 | 7 / 9 | 2.0 | No |
| 473768 | 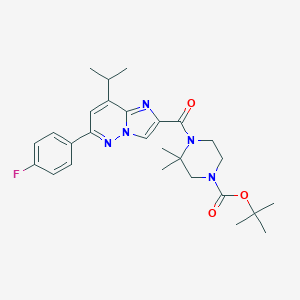 CHEMBL3582248 CHEMBL3582248 | C27H34FN5O3 | 495.599 | 6 / 0 | 4.3 | Yes |
| 524092 | 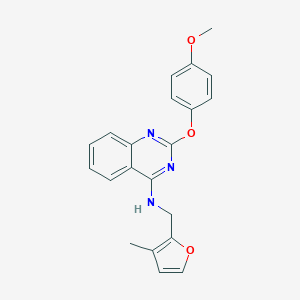 CHEMBL3736475 CHEMBL3736475 | C21H19N3O3 | 361.401 | 6 / 1 | 4.7 | Yes |
| 88982 | 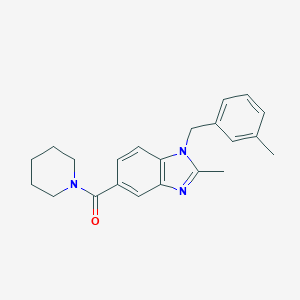 UNII-Z9T72Z578T UNII-Z9T72Z578T | C22H25N3O | 347.462 | 2 / 0 | 4.1 | Yes |
| 90920 | 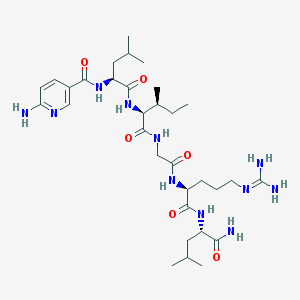 CHEMBL1689561 CHEMBL1689561 | C32H55N11O6 | 689.863 | 9 / 9 | 0.5 | No |
| 91109 | 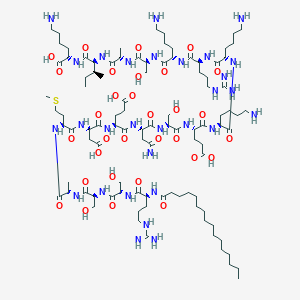 CID 73356960 CID 73356960 | C99H179N29O31S | 2303.75 | 38 / 37 | -14.8 | No |
| 91963 | 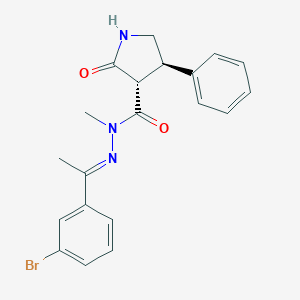 CHEMBL459724 CHEMBL459724 | C20H20BrN3O2 | 414.303 | 3 / 1 | 3.3 | Yes |
| 93773 | 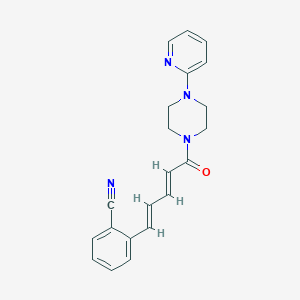 UNII-V423P4PL29 UNII-V423P4PL29 | C21H20N4O | 344.418 | 4 / 0 | 3.0 | Yes |
| 553722 | 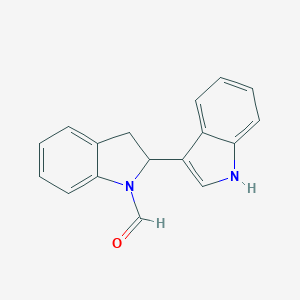 GTPL9586 GTPL9586 | C17H14N2O | 262.312 | 1 / 1 | 3.0 | Yes |
| 549037 | 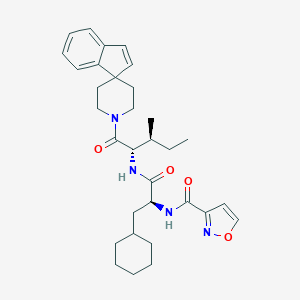 CHEMBL3935419 CHEMBL3935419 | C32H42N4O4 | 546.712 | 5 / 2 | 6.5 | No |
| 97653 | 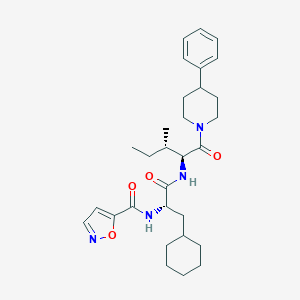 CHEMBL2431716 CHEMBL2431716 | C30H42N4O4 | 522.69 | 5 / 2 | 5.9 | No |
| 524390 | 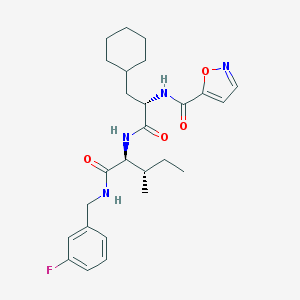 CHEMBL3752956 CHEMBL3752956 | C26H35FN4O4 | 486.588 | 6 / 3 | 5.1 | No |
| 538419 | 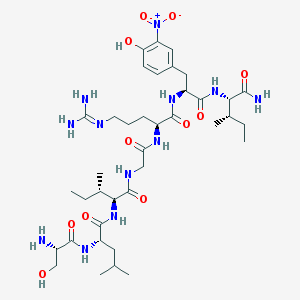 CHEMBL397818 CHEMBL397818 | C38H64N12O11 | 865.003 | 13 / 12 | 0.0 | No |
| 100529 | 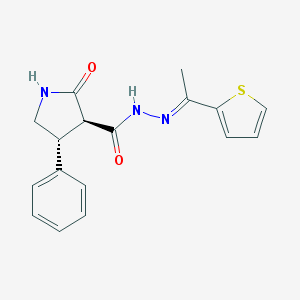 CHEMBL494303 CHEMBL494303 | C17H17N3O2S | 327.402 | 4 / 2 | 3.0 | Yes |
| 102054 | 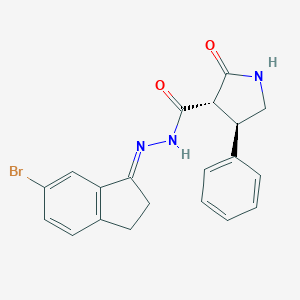 CHEMBL459509 CHEMBL459509 | C20H18BrN3O2 | 412.287 | 3 / 2 | 3.7 | Yes |
| 524587 | 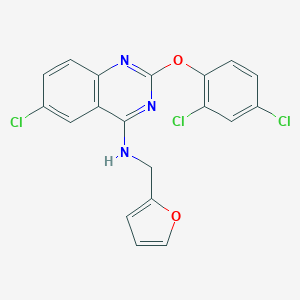 CHEMBL3735017 CHEMBL3735017 | C19H12Cl3N3O2 | 420.674 | 5 / 1 | 6.2 | No |
| 538608 | 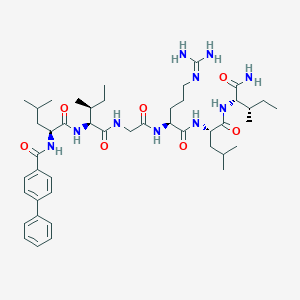 CHEMBL236294 CHEMBL236294 | C45H70N10O7 | 863.118 | 8 / 9 | 4.6 | No |
| 107130 | 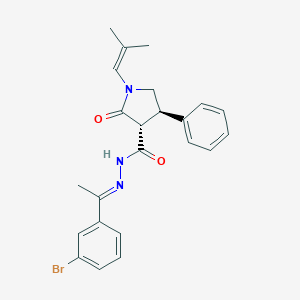 CHEMBL461639 CHEMBL461639 | C23H24BrN3O2 | 454.368 | 3 / 1 | 5.2 | No |
| 538623 | 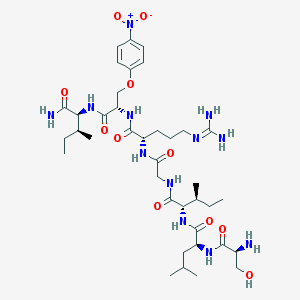 CHEMBL396797 CHEMBL396797 | C38H64N12O11 | 865.003 | 13 / 11 | -0.4 | No |
| 109210 | 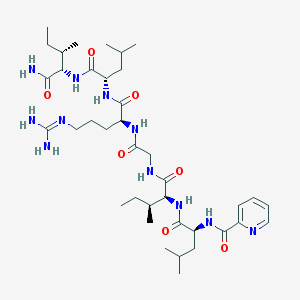 CHEMBL414319 CHEMBL414319 | C38H65N11O7 | 788.008 | 9 / 9 | 2.2 | No |
| 111676 | 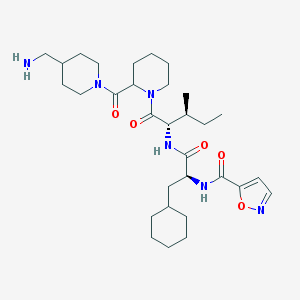 CHEMBL1269140 CHEMBL1269140 | C31H50N6O5 | 586.778 | 7 / 3 | 3.9 | No |
| 538801 | 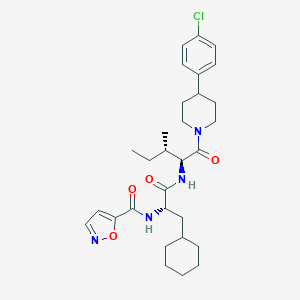 CHEMBL3914140 CHEMBL3914140 | C30H41ClN4O4 | 557.132 | 5 / 2 | 6.6 | No |
| 115937 | 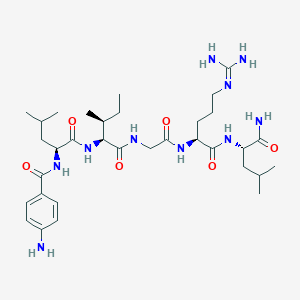 CHEMBL1689563 CHEMBL1689563 | C33H56N10O6 | 688.875 | 8 / 9 | 1.2 | No |
| 525069 | 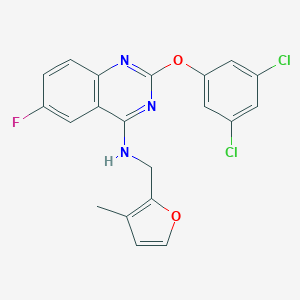 CHEMBL3735493 CHEMBL3735493 | C20H14Cl2FN3O2 | 418.249 | 6 / 1 | 6.0 | No |
| 539117 | 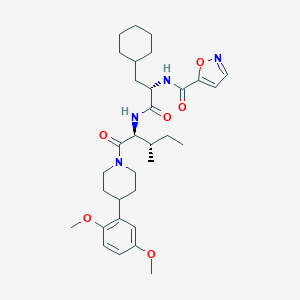 CHEMBL3923129 CHEMBL3923129 | C32H46N4O6 | 582.742 | 7 / 2 | 5.9 | No |
| 553925 | 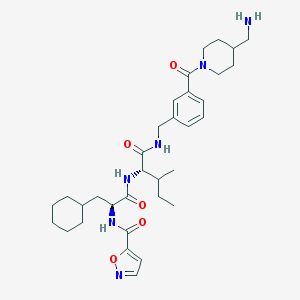 GB110 GB110 | C33H48N6O5 | 608.784 | 7 / 4 | 4.3 | No |
| 126382 | 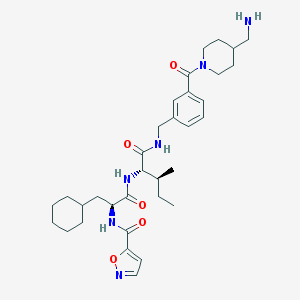 CHEMBL1269133 CHEMBL1269133 | C33H48N6O5 | 608.784 | 7 / 4 | 4.3 | No |
| 525224 | 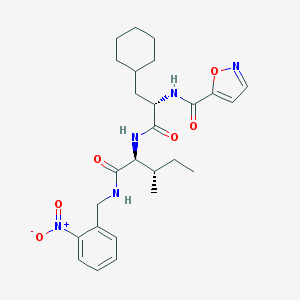 CHEMBL3754518 CHEMBL3754518 | C26H35N5O6 | 513.595 | 7 / 3 | 4.8 | No |
| 553936 | 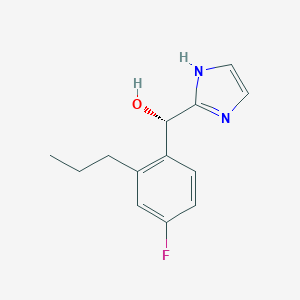 GTPL9585 GTPL9585 | C13H15FN2O | 234.274 | 3 / 2 | 2.3 | Yes |
| 539287 | 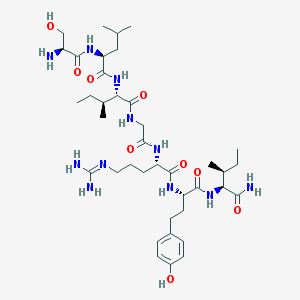 CHEMBL238406 CHEMBL238406 | C39H67N11O9 | 834.033 | 11 / 12 | 0.0 | No |
| 539294 | 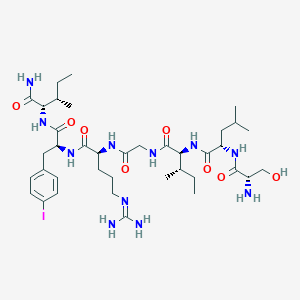 CHEMBL237968 CHEMBL237968 | C38H64IN11O8 | 929.903 | 10 / 11 | 0.7 | No |
| 133135 | 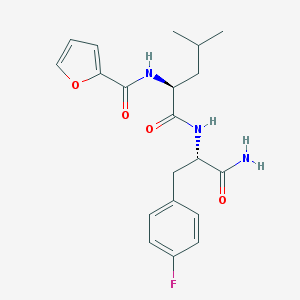 CHEMBL1271131 CHEMBL1271131 | C20H24FN3O4 | 389.427 | 5 / 3 | 1.8 | Yes |
| 525386 | 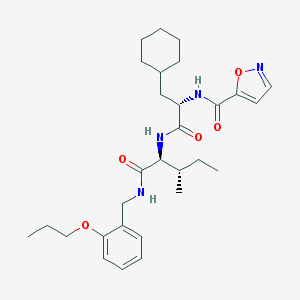 CHEMBL3753815 CHEMBL3753815 | C29H42N4O5 | 526.678 | 6 / 3 | 5.9 | No |
| 549664 | 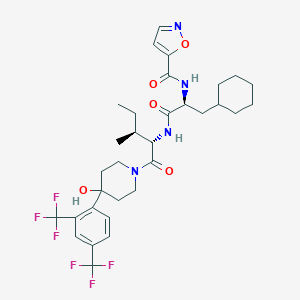 CHEMBL3932079 CHEMBL3932079 | C32H40F6N4O5 | 674.685 | 12 / 3 | 6.6 | No |
| 525604 | 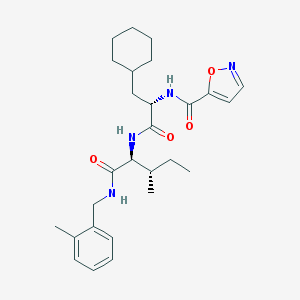 CHEMBL3754533 CHEMBL3754533 | C27H38N4O4 | 482.625 | 5 / 3 | 5.4 | No |
| 539585 | 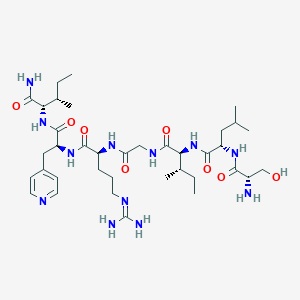 CHEMBL410759 CHEMBL410759 | C37H64N12O8 | 804.995 | 11 / 11 | -1.1 | No |
| 144347 | 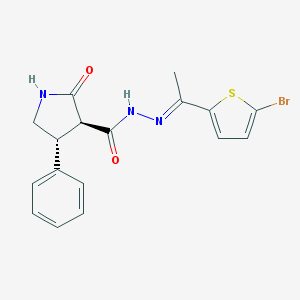 CHEMBL493704 CHEMBL493704 | C17H16BrN3O2S | 406.298 | 4 / 2 | 4.0 | Yes |
| 539675 | 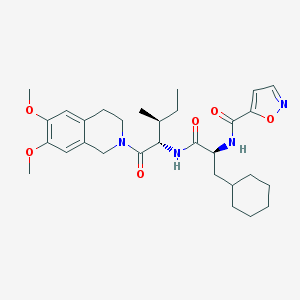 CHEMBL3905570 CHEMBL3905570 | C30H42N4O6 | 554.688 | 7 / 2 | 5.2 | No |
| 145582 | 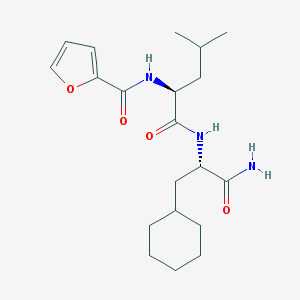 CHEMBL1271026 CHEMBL1271026 | C20H31N3O4 | 377.485 | 4 / 3 | 3.7 | Yes |
| 539721 | 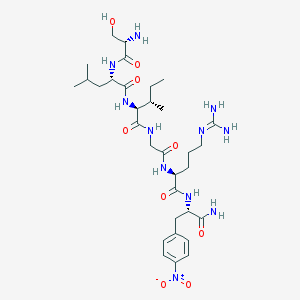 CHEMBL236099 CHEMBL236099 | C32H53N11O9 | 735.844 | 11 / 10 | -2.4 | No |
| 525770 | 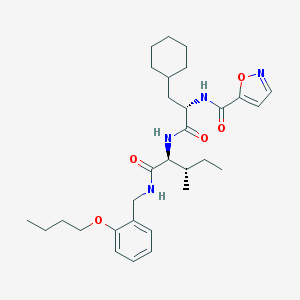 CHEMBL3752763 CHEMBL3752763 | C30H44N4O5 | 540.705 | 6 / 3 | 6.2 | No |
| 147869 | 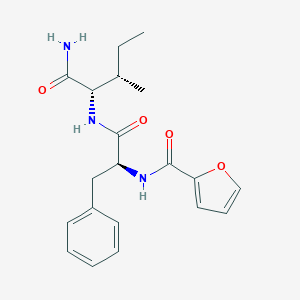 CHEMBL1270824 CHEMBL1270824 | C20H25N3O4 | 371.437 | 4 / 3 | 1.8 | Yes |
| 539816 | 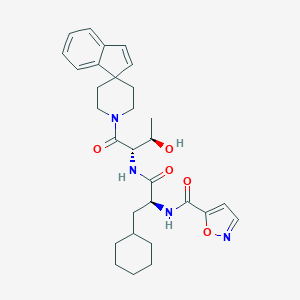 CHEMBL3901521 CHEMBL3901521 | C30H38N4O5 | 534.657 | 6 / 3 | 4.5 | No |
| 539868 | 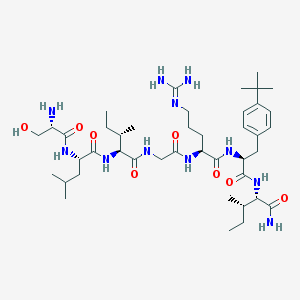 CHEMBL278937 CHEMBL278937 | C42H73N11O8 | 860.115 | 10 / 11 | 1.7 | No |
| 154539 | 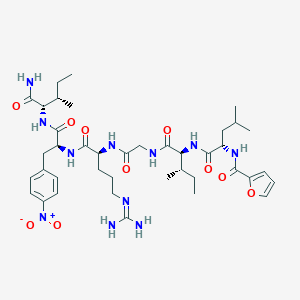 CHEMBL2431723 CHEMBL2431723 | C40H61N11O10 | 855.995 | 11 / 9 | 2.5 | No |
| 155490 | 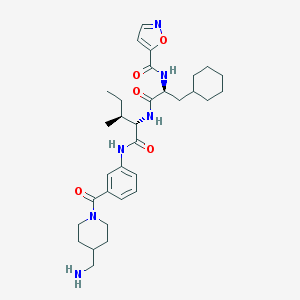 CHEMBL1269139 CHEMBL1269139 | C32H46N6O5 | 594.757 | 7 / 4 | 4.3 | No |
| 526151 | 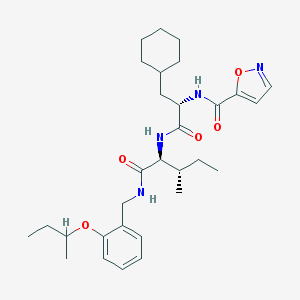 CHEMBL3752235 CHEMBL3752235 | C30H44N4O5 | 540.705 | 6 / 3 | 6.3 | No |
| 549976 | 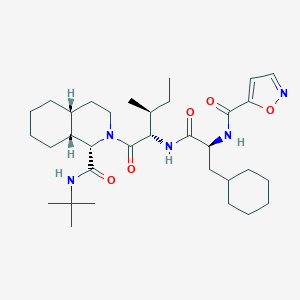 CHEMBL3976309 CHEMBL3976309 | C33H53N5O5 | 599.817 | 6 / 3 | 6.5 | No |
| 166107 | 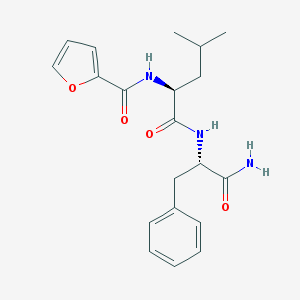 CHEMBL1271130 CHEMBL1271130 | C20H25N3O4 | 371.437 | 4 / 3 | 2.6 | Yes |
| 540296 | 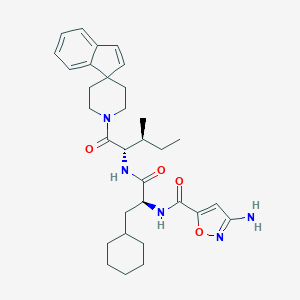 CHEMBL3902670 CHEMBL3902670 | C32H43N5O4 | 561.727 | 6 / 3 | 6.1 | No |
| 172087 | 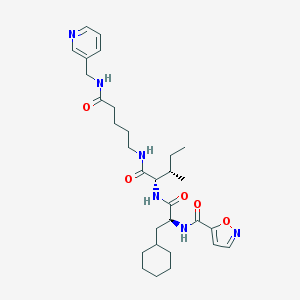 CHEMBL1270125 CHEMBL1270125 | C30H44N6O5 | 568.719 | 7 / 4 | 3.9 | No |
| 540407 | 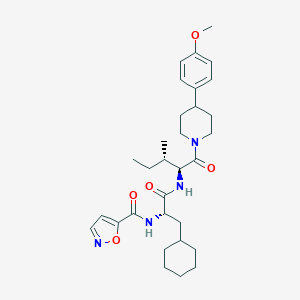 CHEMBL3895158 CHEMBL3895158 | C31H44N4O5 | 552.716 | 6 / 2 | 5.9 | No |
| 526474 | 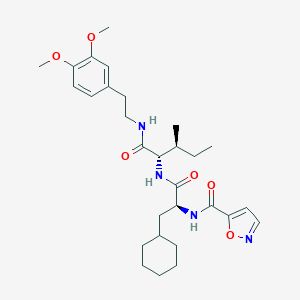 CHEMBL3753010 CHEMBL3753010 | C29H42N4O6 | 542.677 | 7 / 3 | 5.4 | No |
| 176826 | 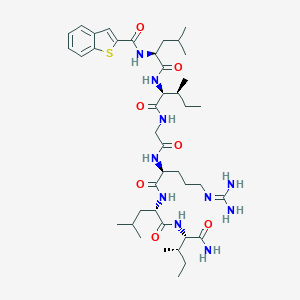 CHEMBL398128 CHEMBL398128 | C41H66N10O7S | 843.102 | 9 / 9 | 4.3 | No |
| 179153 | 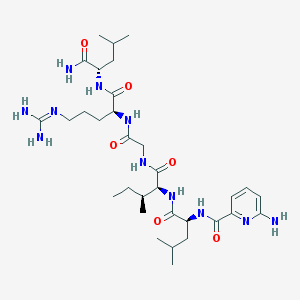 CHEMBL1689562 CHEMBL1689562 | C32H55N11O6 | 689.863 | 9 / 9 | 0.8 | No |
| 540607 | 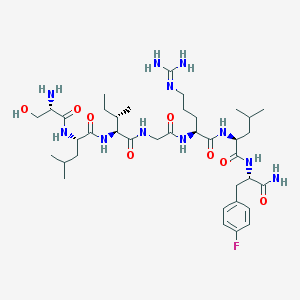 CHEMBL237960 CHEMBL237960 | C38H64FN11O8 | 821.997 | 11 / 11 | -1.1 | No |
| 540643 | 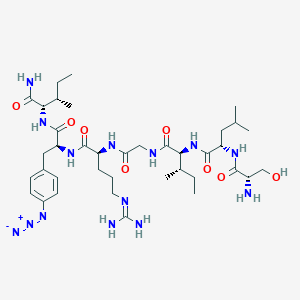 CHEMBL237969 CHEMBL237969 | C38H64N14O8 | 845.02 | 12 / 11 | 0.9 | No |
| 526797 | 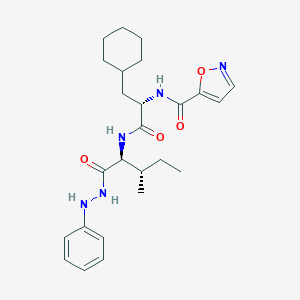 CHEMBL3752860 CHEMBL3752860 | C25H35N5O4 | 469.586 | 6 / 4 | 5.9 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218