

You can:
| Name | Thyrotropin-releasing hormone receptor |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Trhr |
| Synonym | Thyroliberin receptor TRH receptor TRH-R TRH-R1 TRH1 receptor |
| Disease | N/A for non-human GPCRs |
| Length | 412 |
| Amino acid sequence | MENETVSELNQTELPPQVAVALEYQVVTILLVVVICGLGIVGNIMVVLVVMRTKHMRTATNCYLVSLAVADLMVLVAAGLPNITDSIYGSWVYGYVGCLCITYLQYLGINASSCSITAFTIERYIAICHPIKAQFLCTFSRAKKIIIFVWAFTSIYCMLWFFLLDLNISTYKDAIVISCGYKISRNYYSPIYLMDFGVFYVMPMILATVLYGFIARILFLNPIPSDPKENSKTWKNDSTHQNKNMNLNTTNRCFNSTVSSRKQVTKMLAVVVILFALLWMPYRTLVVVNSFLSSPFQENWFLLFCRICIYLNSAINPVIYNLMSQKFRAAFRKLCNCKQKPTEKAANYSVALNYSVIKESDRFSTELDDITVTDTYVSTTKVSFDDTCLASEKNGPSSCTYGYSLTAKQEKI |
| UniProt | Q01717 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL4553 |
| IUPHAR | 363 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 553248 | 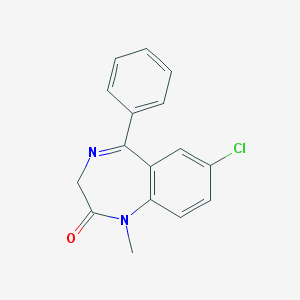 diazepam diazepam | C16H13ClN2O | 284.743 | 2 / 0 | 3.0 | Yes |
| 23706 | 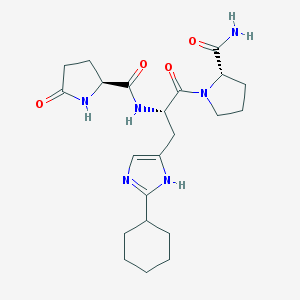 CHEMBL382080 CHEMBL382080 | C22H32N6O4 | 444.536 | 5 / 4 | -0.3 | Yes |
| 30095 | 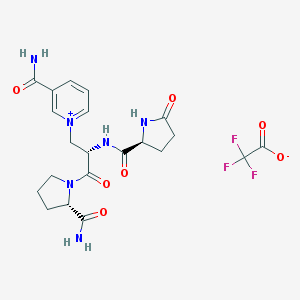 CHEMBL2371826 CHEMBL2371826 | C21H25F3N6O7 | 530.461 | 10 / 4 | N/A | No |
| 553522 | 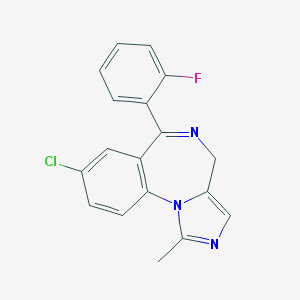 midazolam midazolam | C18H13ClFN3 | 325.771 | 3 / 0 | 2.5 | Yes |
| 553551 | 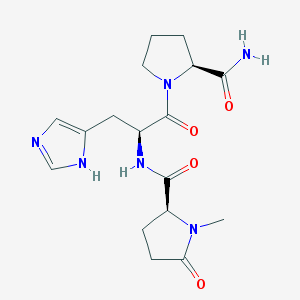 MeTRH MeTRH | C17H24N6O4 | 376.417 | 5 / 3 | -2.3 | Yes |
| 76347 | 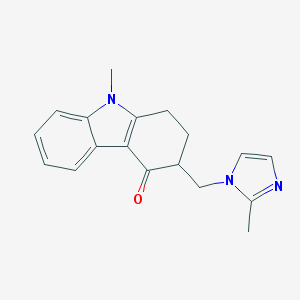 ondansetron ondansetron | C18H19N3O | 293.37 | 2 / 0 | 2.3 | Yes |
| 85926 | 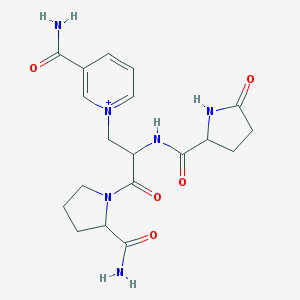 CHEMBL370700 CHEMBL370700 | C19H25N6O5+ | 417.446 | 5 / 4 | -3.0 | Yes |
| 549112 | 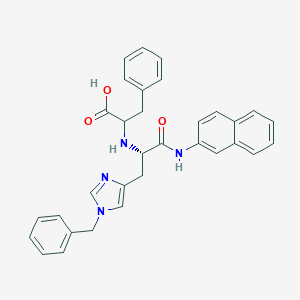 CHEMBL3897627 CHEMBL3897627 | C32H30N4O3 | 518.617 | 5 / 3 | 3.0 | No |
| 109034 | 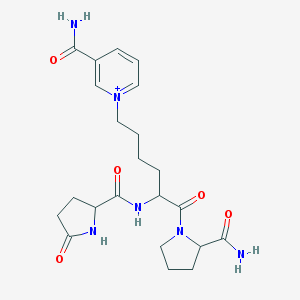 CHEMBL191346 CHEMBL191346 | C22H31N6O5+ | 459.527 | 5 / 4 | -2.0 | Yes |
| 109035 | 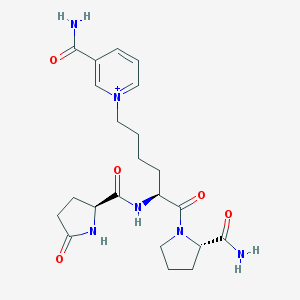 CHEMBL2371830 CHEMBL2371830 | C22H31N6O5+ | 459.527 | 5 / 4 | -2.0 | Yes |
| 118195 | 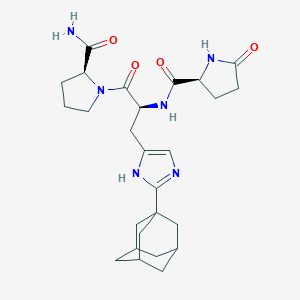 CHEMBL369944 CHEMBL369944 | C26H36N6O4 | 496.612 | 5 / 4 | 0.4 | Yes |
| 145298 | 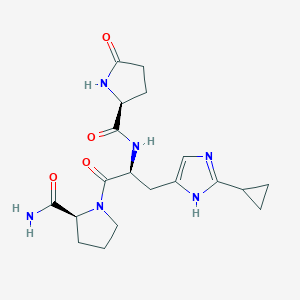 CHEMBL198263 CHEMBL198263 | C19H26N6O4 | 402.455 | 5 / 4 | -1.9 | Yes |
| 179802 | 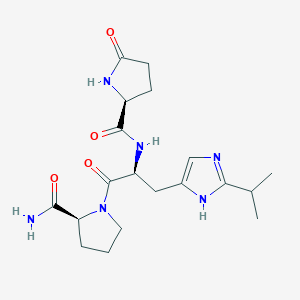 CHEMBL381666 CHEMBL381666 | C19H28N6O4 | 404.471 | 5 / 4 | -1.3 | Yes |
| 187991 | 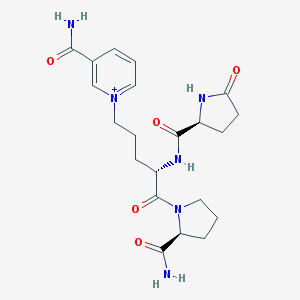 CHEMBL2371825 CHEMBL2371825 | C21H29N6O5+ | 445.5 | 5 / 4 | -2.3 | Yes |
| 187992 | 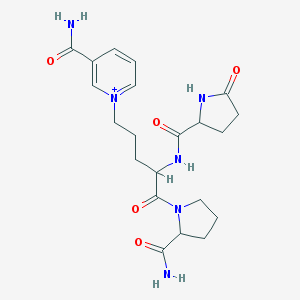 CHEMBL190578 CHEMBL190578 | C21H29N6O5+ | 445.5 | 5 / 4 | -2.3 | Yes |
| 540971 | 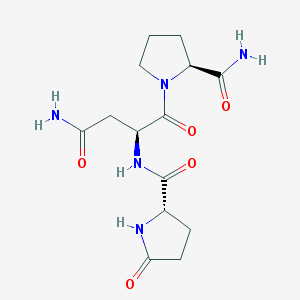 SCHEMBL3132248 SCHEMBL3132248 | C14H21N5O5 | 339.352 | 5 / 4 | -3.7 | Yes |
| 219975 | 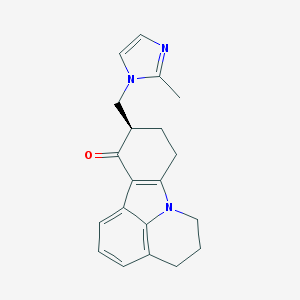 Cilansetron Cilansetron | C20H21N3O | 319.408 | 2 / 0 | 2.6 | Yes |
| 219985 | 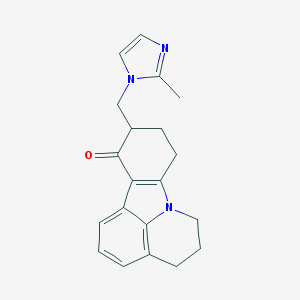 Imidazol-1-yl compound 1 Imidazol-1-yl compound 1 | C20H21N3O | 319.408 | 2 / 0 | 2.6 | Yes |
| 278824 | 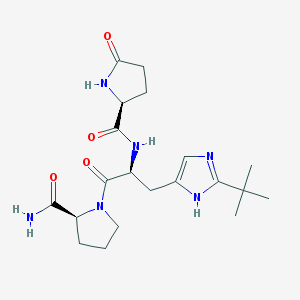 CHEMBL382714 CHEMBL382714 | C20H30N6O4 | 418.498 | 5 / 4 | -0.8 | Yes |
| 287019 | 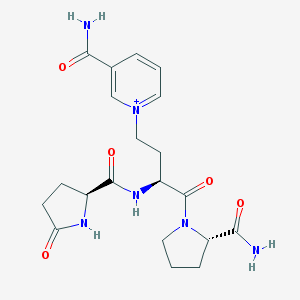 CHEMBL2371831 CHEMBL2371831 | C20H27N6O5+ | 431.473 | 5 / 4 | -2.7 | Yes |
| 287020 | 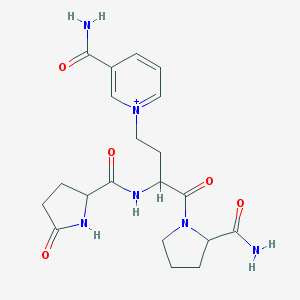 CHEMBL362358 CHEMBL362358 | C20H27N6O5+ | 431.473 | 5 / 4 | -2.7 | Yes |
| 351518 | 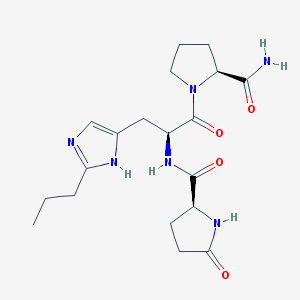 CHEMBL197149 CHEMBL197149 | C19H28N6O4 | 404.471 | 5 / 4 | -1.3 | Yes |
| 352654 | 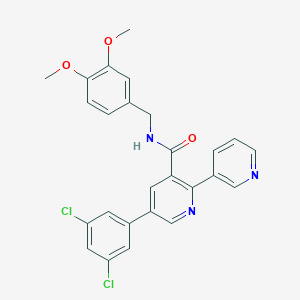 CHEMBL3099899 CHEMBL3099899 | C26H21Cl2N3O3 | 494.372 | 5 / 1 | 5.0 | Yes |
| 375708 | 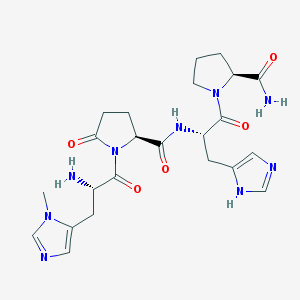 CHEMBL365775 CHEMBL365775 | C23H31N9O5 | 513.559 | 8 / 4 | -2.7 | No |
| 396601 | 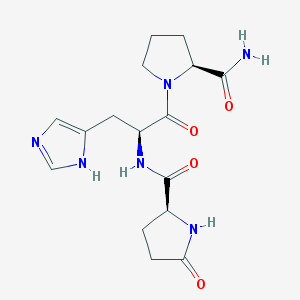 protirelin protirelin | C16H22N6O4 | 362.39 | 5 / 4 | -2.5 | Yes |
| 547231 | 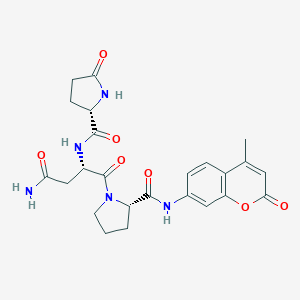 CHEMBL3906398 CHEMBL3906398 | C24H27N5O7 | 497.508 | 7 / 4 | -1.3 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218