

You can:
| Name | 5-hydroxytryptamine receptor 4 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | HTR4 |
| Synonym | 5-HT-4 5-hydroxytryptamine (serotonin) receptor 4, G protein-coupled 5-HT4 receptor 5-HT4 serotonin receptor 4 |
| Disease | N/A |
| Length | 388 |
| Amino acid sequence | MDKLDANVSSEEGFGSVEKVVLLTFLSTVILMAILGNLLVMVAVCWDRQLRKIKTNYFIVSLAFADLLVSVLVMPFGAIELVQDIWIYGEVFCLVRTSLDVLLTTASIFHLCCISLDRYYAICCQPLVYRNKMTPLRIALMLGGCWVIPTFISFLPIMQGWNNIGIIDLIEKRKFNQNSNSTYCVFMVNKPYAITCSVVAFYIPFLLMVLAYYRIYVTAKEHAHQIQMLQRAGASSESRPQSADQHSTHRMRTETKAAKTLCIIMGCFCLCWAPFFVTNIVDPFIDYTVPGQVWTAFLWLGYINSGLNPFLYAFLNKSFRRAFLIILCCDDERYRRPSILGQTVPCSTTTINGSTHVLRDAVECGGQWESQCHPPATSPLVAAQPSDT |
| UniProt | Q13639 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q13639 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q13639. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL1875 |
| IUPHAR | 9 |
| DrugBank | BE0000084 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 521442 | 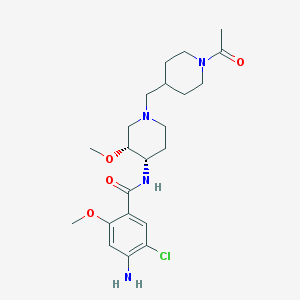 CHEMBL3759362 CHEMBL3759362 | C22H33ClN4O4 | 452.98 | 6 / 2 | 1.5 | Yes |
| 509 | 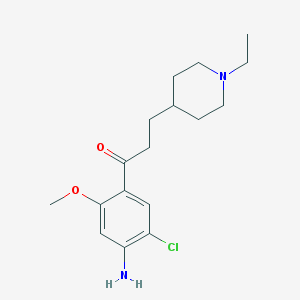 CHEMBL82360 CHEMBL82360 | C17H25ClN2O2 | 324.849 | 4 / 1 | 3.0 | Yes |
| 1617 | 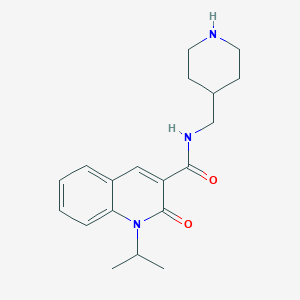 CHEMBL551535 CHEMBL551535 | C19H25N3O2 | 327.428 | 3 / 2 | 2.7 | Yes |
| 1687 | 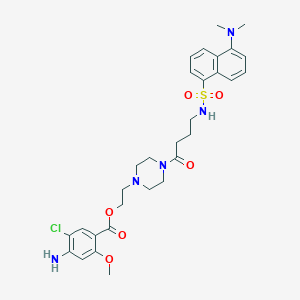 CHEMBL156206 CHEMBL156206 | C30H38ClN5O6S | 632.173 | 10 / 2 | 3.3 | No |
| 3718 | 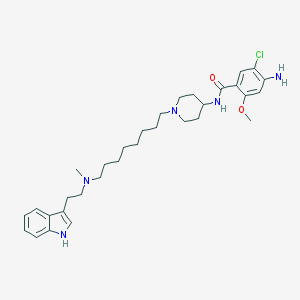 CHEMBL552484 CHEMBL552484 | C32H46ClN5O2 | 568.203 | 5 / 3 | 6.3 | No |
| 3844 | 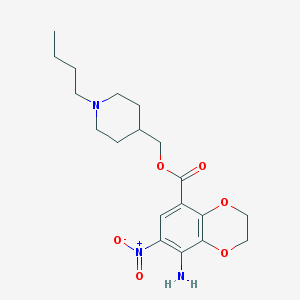 CHEMBL1255606 CHEMBL1255606 | C19H27N3O6 | 393.44 | 8 / 1 | 3.2 | Yes |
| 557397 | 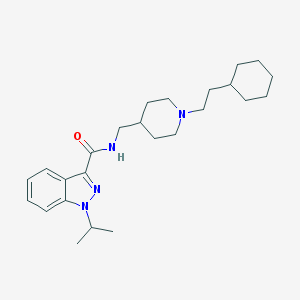 CHEMBL2179699 CHEMBL2179699 | C25H38N4O | 410.606 | 3 / 1 | 5.8 | No |
| 4100 | 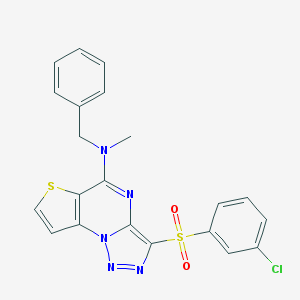 CHEMBL1173578 CHEMBL1173578 | C21H16ClN5O2S2 | 469.962 | 7 / 0 | 4.6 | Yes |
| 553264 | 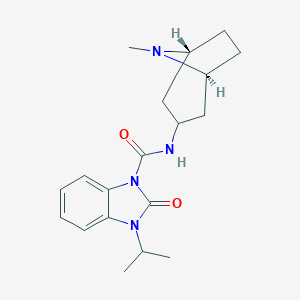 BIMU-8 BIMU-8 | C19H26N4O2 | 342.443 | 3 / 1 | 2.5 | Yes |
| 555498 | 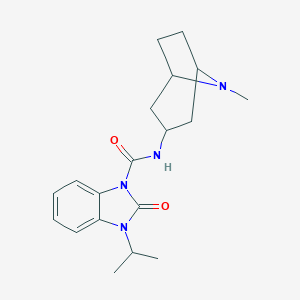 CHEMBL59834 CHEMBL59834 | C19H26N4O2 | 342.443 | 3 / 1 | 2.5 | Yes |
| 5019 | 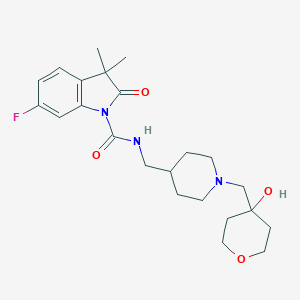 UNII-QPE81T4KH0 UNII-QPE81T4KH0 | C23H32FN3O4 | 433.524 | 6 / 2 | 2.5 | Yes |
| 555503 | 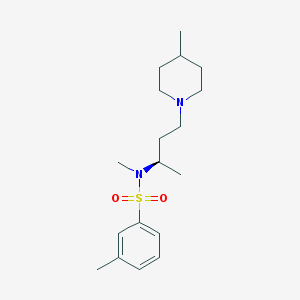 SB-258719 SB-258719 | C18H30N2O2S | 338.51 | 4 / 0 | 3.5 | Yes |
| 521616 | 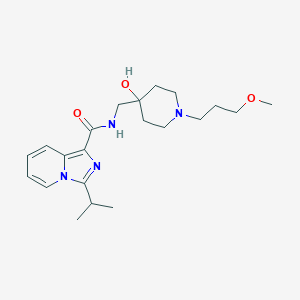 CHEMBL3740030 CHEMBL3740030 | C21H32N4O3 | 388.512 | 5 / 2 | 2.6 | Yes |
| 5862 | 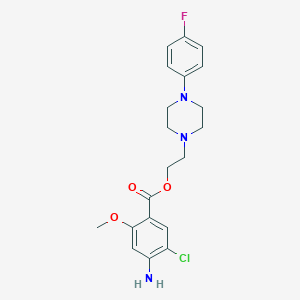 CHEMBL333028 CHEMBL333028 | C20H23ClFN3O3 | 407.87 | 7 / 1 | 3.4 | Yes |
| 521635 | 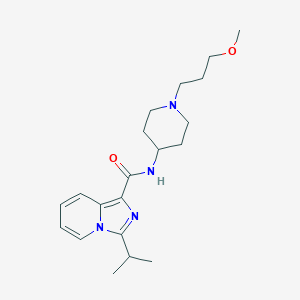 CHEMBL3741776 CHEMBL3741776 | C20H30N4O2 | 358.486 | 4 / 1 | 3.4 | Yes |
| 6110 | 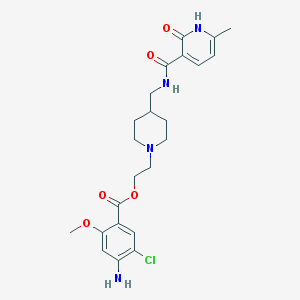 CHEMBL489412 CHEMBL489412 | C23H29ClN4O5 | 476.958 | 7 / 3 | 2.9 | Yes |
| 6446 | 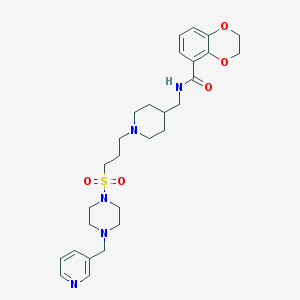 CHEMBL181415 CHEMBL181415 | C28H39N5O5S | 557.71 | 9 / 1 | 1.6 | No |
| 6710 | 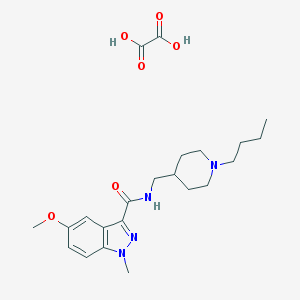 CHEMBL2179689 CHEMBL2179689 | C22H32N4O6 | 448.52 | 8 / 3 | N/A | N/A |
| 536126 | 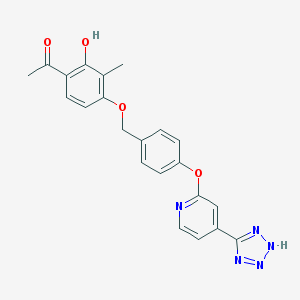 CHEMBL3899832 CHEMBL3899832 | C22H19N5O4 | 417.425 | 8 / 2 | 3.6 | Yes |
| 9592 | 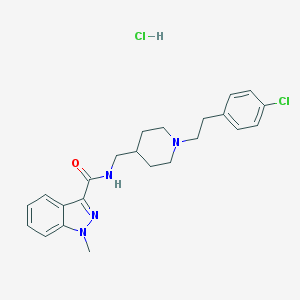 CHEMBL2179686 CHEMBL2179686 | C23H28Cl2N4O | 447.404 | 3 / 2 | N/A | N/A |
| 9861 | 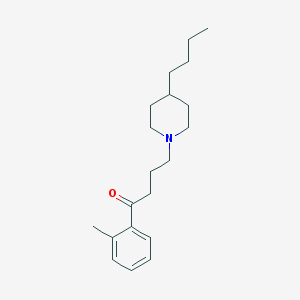 AC-42 AC-42 | C20H31NO | 301.474 | 2 / 0 | 5.2 | No |
| 10434 | 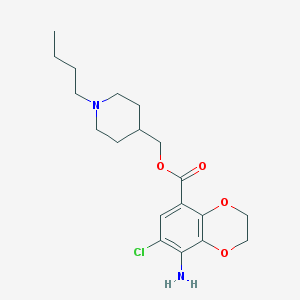 SB 204070 hydrochloride SB 204070 hydrochloride | C19H27ClN2O4 | 382.885 | 6 / 1 | 3.5 | Yes |
| 10872 | 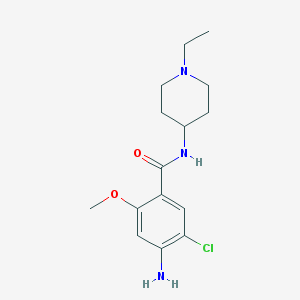 CHEMBL570074 CHEMBL570074 | C15H22ClN3O2 | 311.81 | 4 / 2 | 2.8 | Yes |
| 10960 | 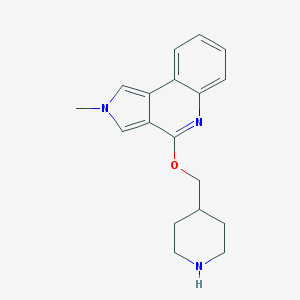 CHEMBL2179680 CHEMBL2179680 | C18H21N3O | 295.386 | 3 / 1 | 2.8 | Yes |
| 14238 | 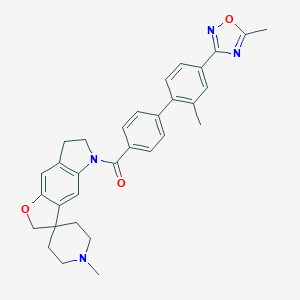 180083-23-2 180083-23-2 | C32H32N4O3 | 520.633 | 6 / 0 | 5.6 | No |
| 519753 | 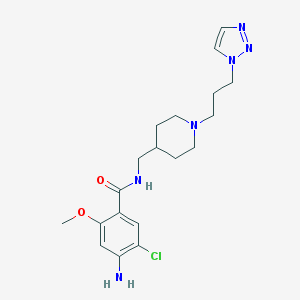 UNII-J6388YJ4YR UNII-J6388YJ4YR | C19H27ClN6O2 | 406.915 | 6 / 2 | 1.7 | Yes |
| 15052 | 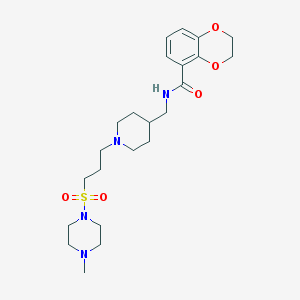 5-HT4 antagonist 1 5-HT4 antagonist 1 | C23H36N4O5S | 480.624 | 8 / 1 | 1.2 | Yes |
| 15075 | 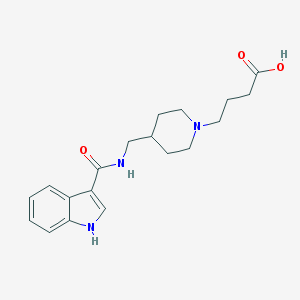 857650-83-0 857650-83-0 | C19H25N3O3 | 343.427 | 4 / 3 | 0.1 | Yes |
| 15574 | 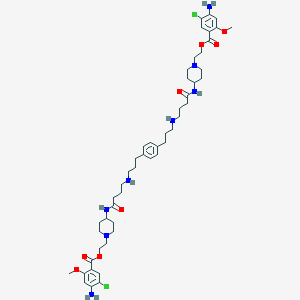 ML10302 scaffold, 14 ML10302 scaffold, 14 | C50H72Cl2N8O8 | 984.074 | 14 / 6 | 5.6 | No |
| 521951 | 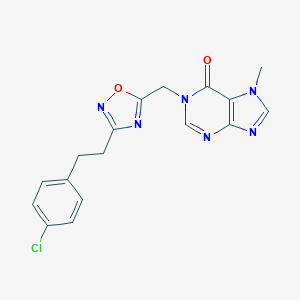 AM-0902 AM-0902 | C17H15ClN6O2 | 370.797 | 6 / 0 | 2.3 | Yes |
| 17043 | 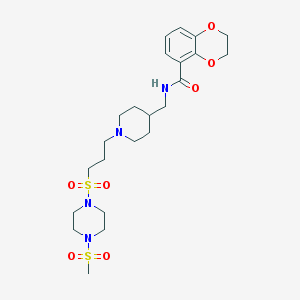 CHEMBL179799 CHEMBL179799 | C23H36N4O7S2 | 544.682 | 10 / 1 | 0.4 | No |
| 17382 | 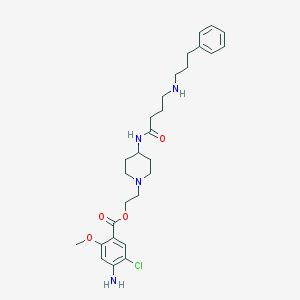 ML10302 scaffold, 19 ML10302 scaffold, 19 | C28H39ClN4O4 | 531.094 | 7 / 3 | 3.8 | No |
| 442380 | 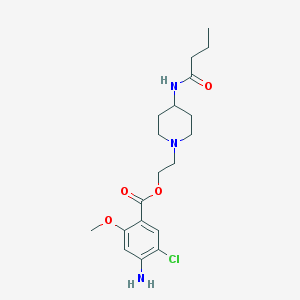 CHEMBL490432 CHEMBL490432 | C19H28ClN3O4 | 397.9 | 6 / 2 | 2.4 | Yes |
| 519778 | 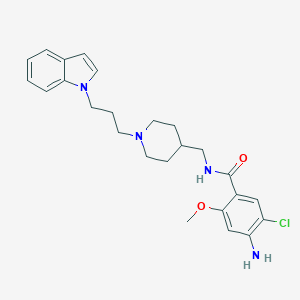 SCHEMBL725615 SCHEMBL725615 | C25H31ClN4O2 | 454.999 | 4 / 2 | 4.0 | Yes |
| 19445 | 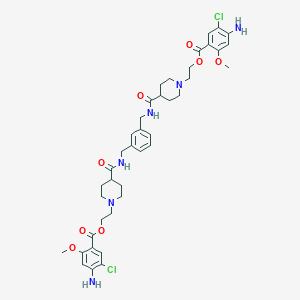 CHEMBL198668 CHEMBL198668 | C40H50Cl2N6O8 | 813.774 | 12 / 4 | 4.1 | No |
| 19838 | 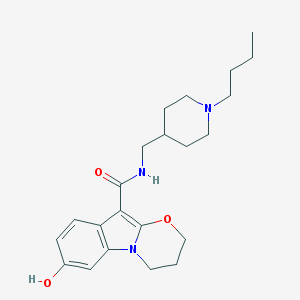 CHEMBL2440451 CHEMBL2440451 | C22H31N3O3 | 385.508 | 4 / 2 | 3.3 | Yes |
| 442497 | 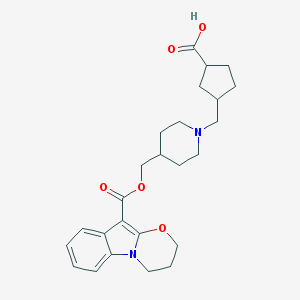 CHEMBL3329811 CHEMBL3329811 | C25H32N2O5 | 440.54 | 6 / 1 | 1.4 | Yes |
| 20939 | 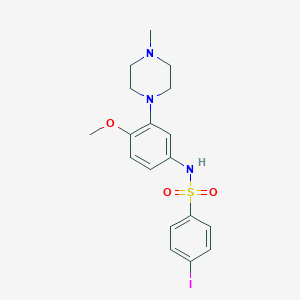 SB-258585 SB-258585 | C18H22IN3O3S | 487.356 | 6 / 1 | 3.0 | Yes |
| 21426 | 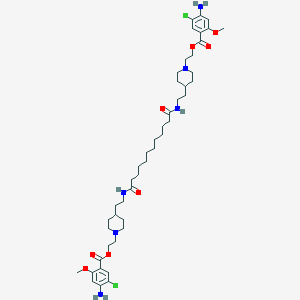 CHEMBL371718 CHEMBL371718 | C46H70Cl2N6O8 | 906.0 | 12 / 4 | 7.9 | No |
| 21818 | 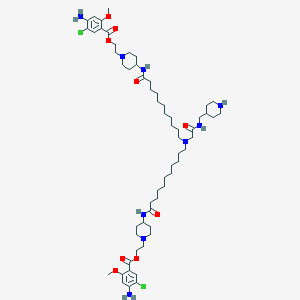 CHEMBL408379 CHEMBL408379 | C60H97Cl2N9O9 | 1159.39 | 15 / 6 | 10.1 | No |
| 522100 | 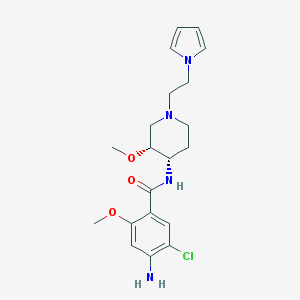 CHEMBL3758721 CHEMBL3758721 | C20H27ClN4O3 | 406.911 | 5 / 2 | 1.6 | Yes |
| 23437 | 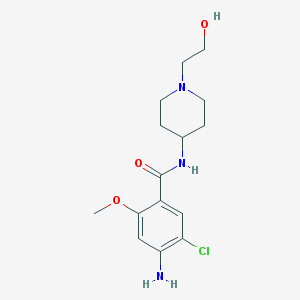 CHEMBL562477 CHEMBL562477 | C15H22ClN3O3 | 327.809 | 5 / 3 | 1.8 | Yes |
| 23771 | 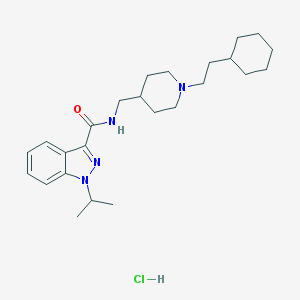 CHEMBL2179699 CHEMBL2179699 | C25H39ClN4O | 447.064 | 3 / 2 | N/A | N/A |
| 23971 | 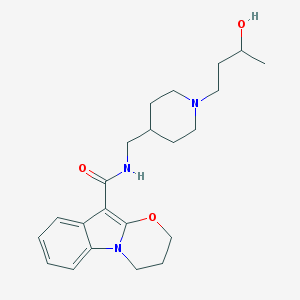 CHEMBL2440453 CHEMBL2440453 | C22H31N3O3 | 385.508 | 4 / 2 | 2.5 | Yes |
| 522291 | 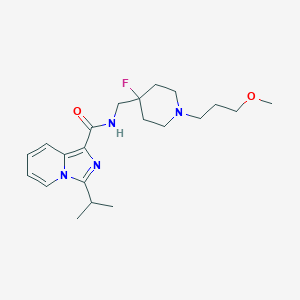 CHEMBL3740864 CHEMBL3740864 | C21H31FN4O2 | 390.503 | 5 / 1 | 3.6 | Yes |
| 24105 | 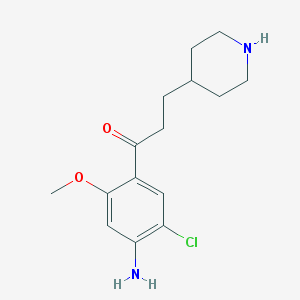 CHEMBL82833 CHEMBL82833 | C15H21ClN2O2 | 296.795 | 4 / 2 | 2.1 | Yes |
| 24282 | 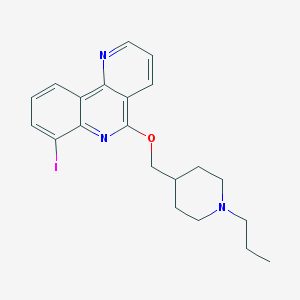 CHEMBL2181168 CHEMBL2181168 | C21H24IN3O | 461.347 | 4 / 0 | 5.1 | No |
| 536619 | 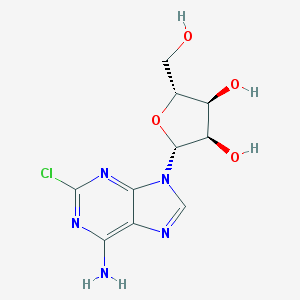 2-Chloroadenosine 2-Chloroadenosine | C10H12ClN5O4 | 301.687 | 8 / 4 | -0.1 | Yes |
| 558054 | 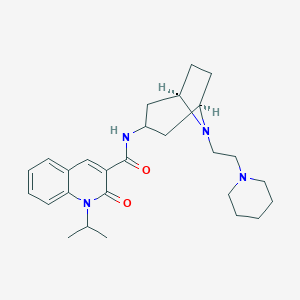 CHEMBL558259 CHEMBL558259 | C27H38N4O2 | 450.627 | 4 / 1 | 3.9 | Yes |
| 25961 | 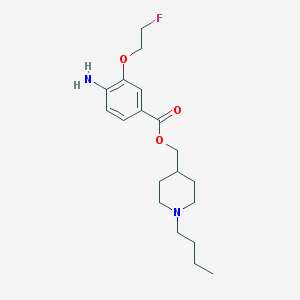 CHEMBL1258786 CHEMBL1258786 | C19H29FN2O3 | 352.45 | 6 / 1 | 3.4 | Yes |
| 26923 | 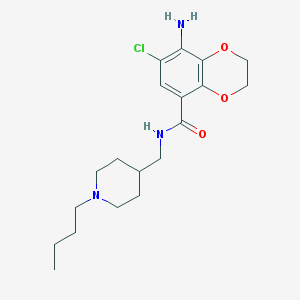 CHEMBL323475 CHEMBL323475 | C19H28ClN3O3 | 381.901 | 5 / 2 | 2.9 | Yes |
| 27053 | 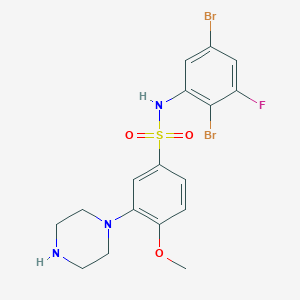 SB-357134 SB-357134 | C17H18Br2FN3O3S | 523.215 | 7 / 2 | 3.4 | No |
| 27275 | 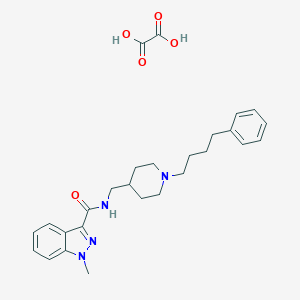 CHEMBL2179687 CHEMBL2179687 | C27H34N4O5 | 494.592 | 7 / 3 | N/A | N/A |
| 27663 | 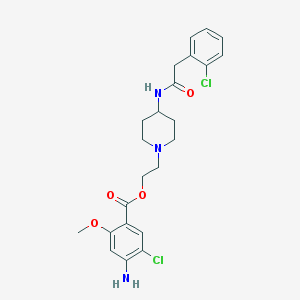 CHEMBL490852 CHEMBL490852 | C23H27Cl2N3O4 | 480.386 | 6 / 2 | 3.8 | Yes |
| 558125 | 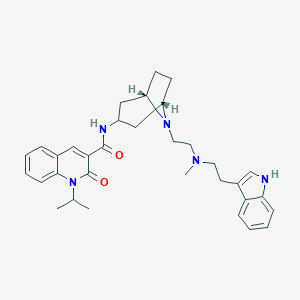 CHEMBL541223 CHEMBL541223 | C33H41N5O2 | 539.724 | 4 / 2 | 5.1 | No |
| 28792 | 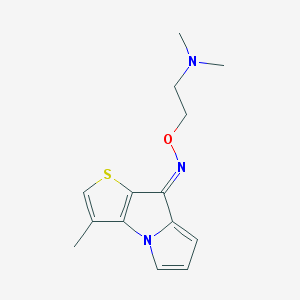 CHEMBL334923 CHEMBL334923 | C14H17N3OS | 275.37 | 4 / 0 | 3.2 | Yes |
| 29924 | 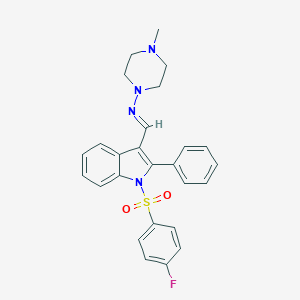 CHEMBL2414713 CHEMBL2414713 | C26H25FN4O2S | 476.57 | 6 / 0 | 5.0 | Yes |
| 31485 | 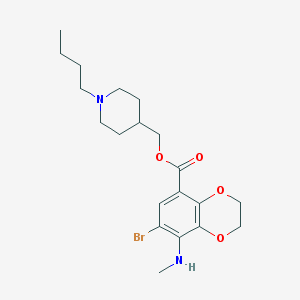 CHEMBL1258338 CHEMBL1258338 | C20H29BrN2O4 | 441.366 | 6 / 1 | 4.2 | Yes |
| 548238 | 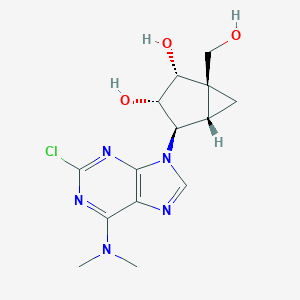 CHEMBL3959210 CHEMBL3959210 | C14H18ClN5O3 | 339.78 | 7 / 3 | 0.3 | Yes |
| 31935 | 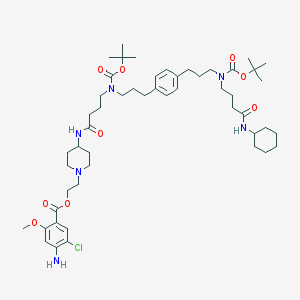 CHEMBL2113291 CHEMBL2113291 | C51H79ClN6O9 | 955.676 | 11 / 3 | 8.0 | No |
| 32013 | 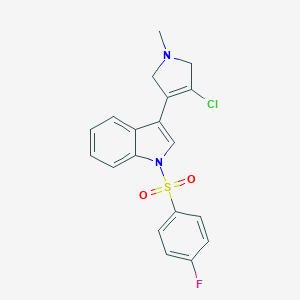 SCHEMBL6530503 SCHEMBL6530503 | C19H16ClFN2O2S | 390.857 | 4 / 0 | 3.4 | Yes |
| 536842 | 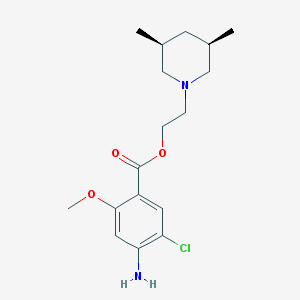 CHEMBL50660 CHEMBL50660 | C17H25ClN2O3 | 340.848 | 5 / 1 | 3.4 | Yes |
| 553418 | 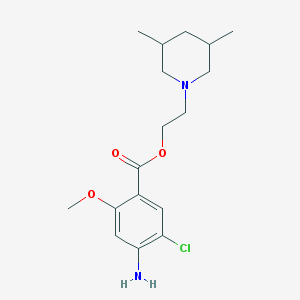 CHEMBL155917 CHEMBL155917 | C17H25ClN2O3 | 340.848 | 5 / 1 | 3.4 | Yes |
| 33636 | 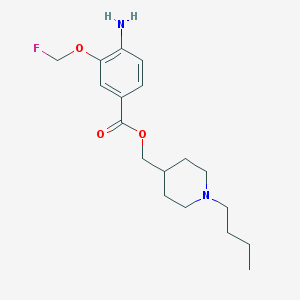 CHEMBL1258672 CHEMBL1258672 | C18H27FN2O3 | 338.423 | 6 / 1 | 3.5 | Yes |
| 33824 | 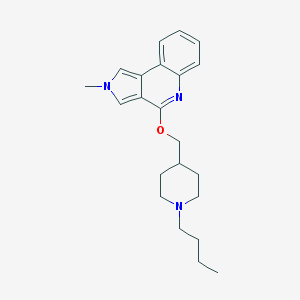 CHEMBL2179679 CHEMBL2179679 | C22H29N3O | 351.494 | 3 / 0 | 4.5 | Yes |
| 536867 | 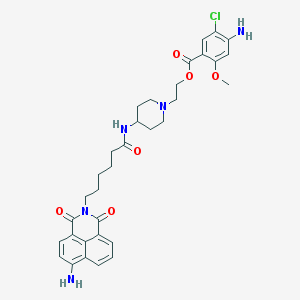 BDBM29528 BDBM29528 | C33H38ClN5O6 | 636.146 | 9 / 3 | 3.8 | No |
| 36069 | 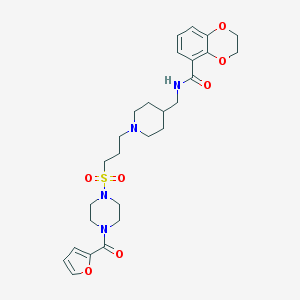 CHEMBL180830 CHEMBL180830 | C27H36N4O7S | 560.666 | 9 / 1 | 1.7 | No |
| 36549 | 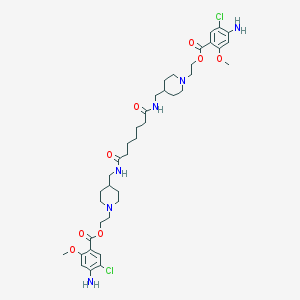 CHEMBL382543 CHEMBL382543 | C39H56Cl2N6O8 | 807.811 | 12 / 4 | 4.5 | No |
| 36588 | 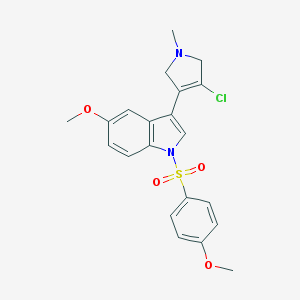 SCHEMBL6530751 SCHEMBL6530751 | C21H21ClN2O4S | 432.919 | 5 / 0 | 3.3 | Yes |
| 37686 | 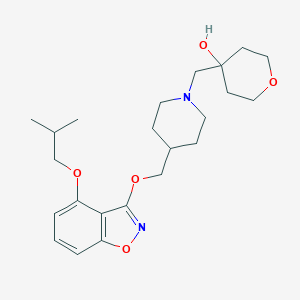 CHEMBL2179589 CHEMBL2179589 | C23H34N2O5 | 418.534 | 7 / 1 | 3.6 | Yes |
| 38003 | 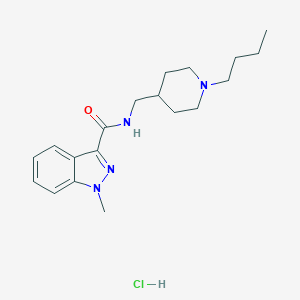 CHEMBL2179682 CHEMBL2179682 | C19H29ClN4O | 364.918 | 3 / 2 | N/A | N/A |
| 39393 | 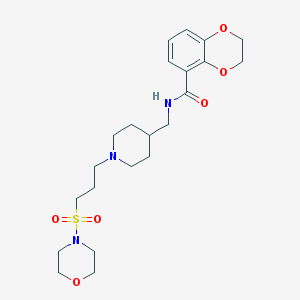 CHEMBL362582 CHEMBL362582 | C22H33N3O6S | 467.581 | 8 / 1 | 1.0 | Yes |
| 522750 | 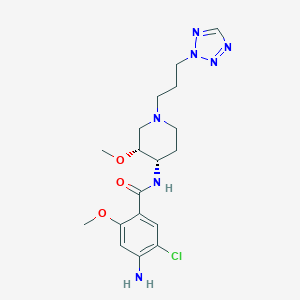 CHEMBL3758523 CHEMBL3758523 | C18H26ClN7O3 | 423.902 | 8 / 2 | 1.5 | Yes |
| 42997 | 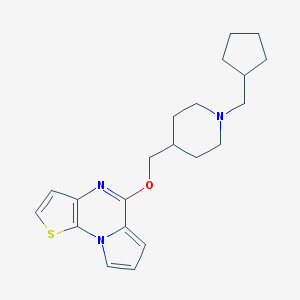 CHEMBL3218878 CHEMBL3218878 | C21H27N3OS | 369.527 | 4 / 0 | 5.8 | No |
| 43616 | 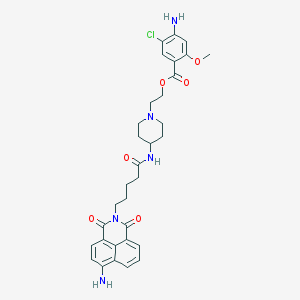 CHEMBL452353 CHEMBL452353 | C32H36ClN5O6 | 622.119 | 9 / 3 | 3.5 | No |
| 45120 | 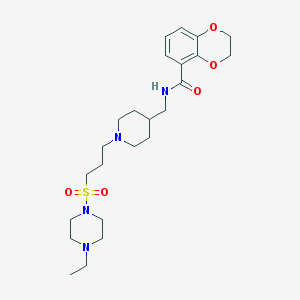 CHEMBL424790 CHEMBL424790 | C24H38N4O5S | 494.651 | 8 / 1 | 1.5 | Yes |
| 45166 | 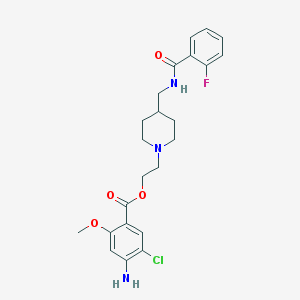 CHEMBL491648 CHEMBL491648 | C23H27ClFN3O4 | 463.934 | 7 / 2 | 3.5 | Yes |
| 558680 | 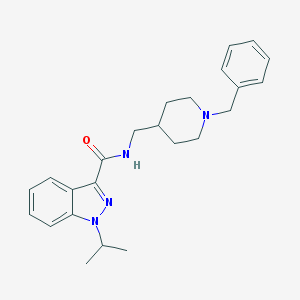 CHEMBL2179696 CHEMBL2179696 | C24H30N4O | 390.531 | 3 / 1 | 4.2 | Yes |
| 45380 | 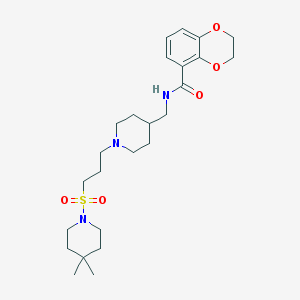 CHEMBL178285 CHEMBL178285 | C25H39N3O5S | 493.663 | 7 / 1 | 3.0 | Yes |
| 45409 | 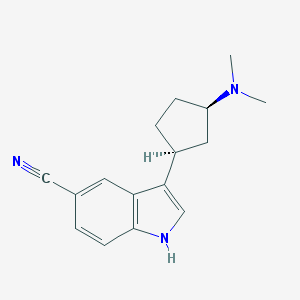 CHEMBL1275709 CHEMBL1275709 | C16H19N3 | 253.349 | 2 / 1 | 2.8 | Yes |
| 48328 | 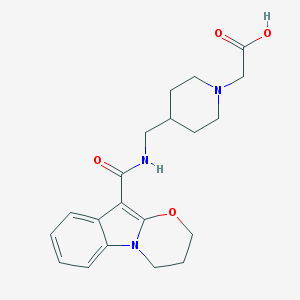 CHEMBL1632170 CHEMBL1632170 | C20H25N3O4 | 371.437 | 5 / 2 | -0.3 | Yes |
| 48854 | 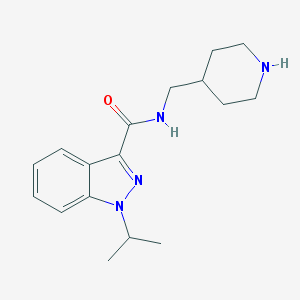 CHEMBL552796 CHEMBL552796 | C17H24N4O | 300.406 | 3 / 2 | 2.2 | Yes |
| 49386 | 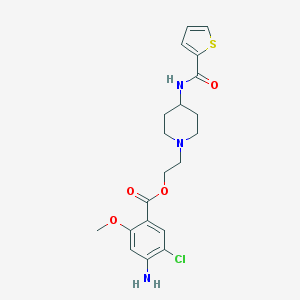 CHEMBL491855 CHEMBL491855 | C20H24ClN3O4S | 437.939 | 7 / 2 | 3.3 | Yes |
| 51388 | 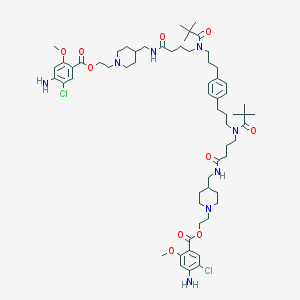 CHEMBL1162955 CHEMBL1162955 | C62H92Cl2N8O10 | 1180.36 | 14 / 4 | 8.7 | No |
| 51718 | 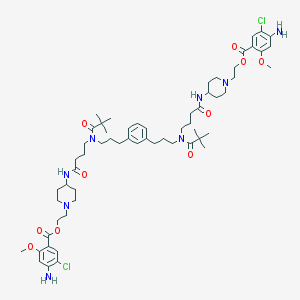 CHEMBL390202 CHEMBL390202 | C60H88Cl2N8O10 | 1152.31 | 14 / 4 | 8.3 | No |
| 52089 | 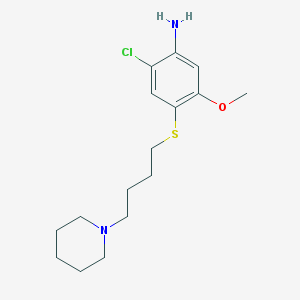 CHEMBL314210 CHEMBL314210 | C16H25ClN2OS | 328.899 | 4 / 1 | 3.9 | Yes |
| 558889 | 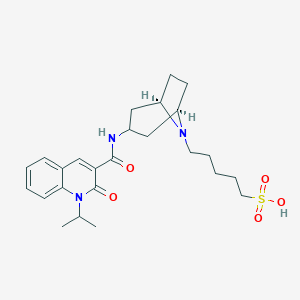 CHEMBL559746 CHEMBL559746 | C25H35N3O5S | 489.631 | 6 / 2 | 0.7 | Yes |
| 52576 | 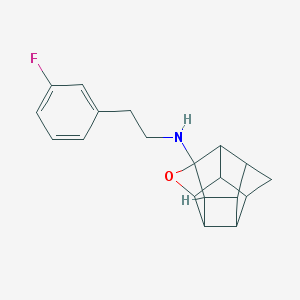 CHEMBL590615 CHEMBL590615 | C19H20FNO | 297.373 | 3 / 1 | 3.0 | Yes |
| 54067 | 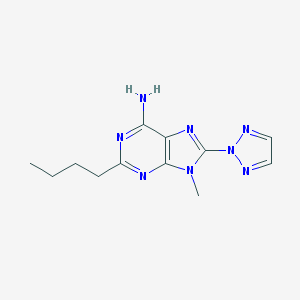 ST-1535 ST-1535 | C12H16N8 | 272.316 | 6 / 1 | 1.8 | Yes |
| 55244 | 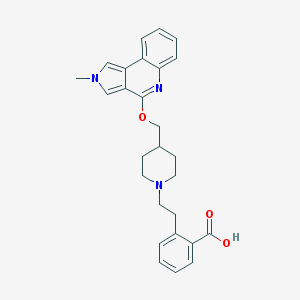 CHEMBL2179673 CHEMBL2179673 | C27H29N3O3 | 443.547 | 5 / 1 | 2.5 | Yes |
| 56030 | 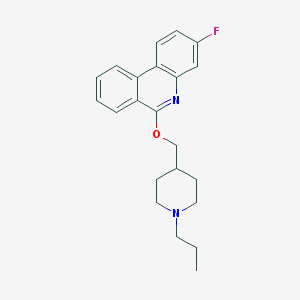 CHEMBL2181185 CHEMBL2181185 | C22H25FN2O | 352.453 | 4 / 0 | 5.5 | No |
| 56092 | 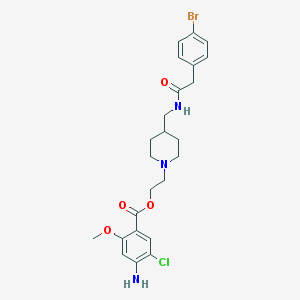 CHEMBL489642 CHEMBL489642 | C24H29BrClN3O4 | 538.867 | 6 / 2 | 4.1 | No |
| 523169 | 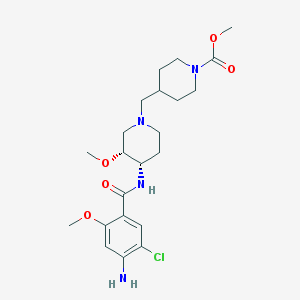 CHEMBL3759969 CHEMBL3759969 | C22H33ClN4O5 | 468.979 | 7 / 2 | 1.9 | Yes |
| 56461 | 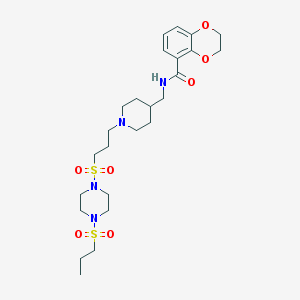 CHEMBL360245 CHEMBL360245 | C25H40N4O7S2 | 572.736 | 10 / 1 | 1.3 | No |
| 56663 | 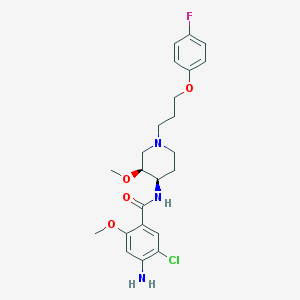 cisapride cisapride | C23H29ClFN3O4 | 465.95 | 7 / 2 | 3.4 | Yes |
| 56664 | 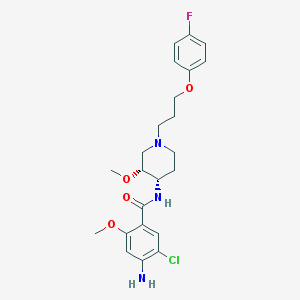 cisapride cisapride | C23H29ClFN3O4 | 465.95 | 7 / 2 | 3.4 | Yes |
| 56677 | 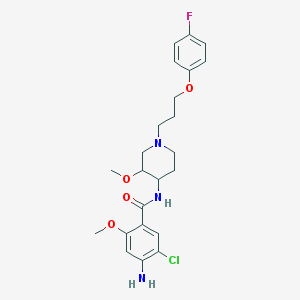 cisapride cisapride | C23H29ClFN3O4 | 465.95 | 7 / 2 | 3.4 | Yes |
| 559018 | 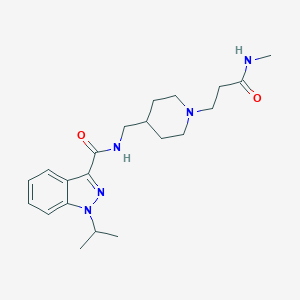 CHEMBL2179703 CHEMBL2179703 | C21H31N5O2 | 385.512 | 4 / 2 | 1.9 | Yes |
| 56732 | 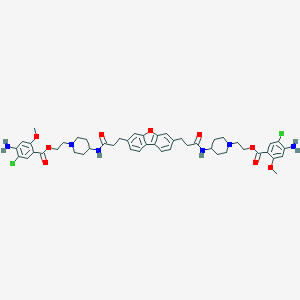 CHEMBL198226 CHEMBL198226 | C48H56Cl2N6O9 | 931.909 | 13 / 4 | 6.8 | No |
| 523194 | 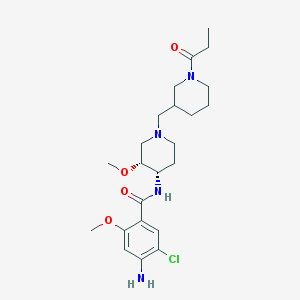 CHEMBL3759348 CHEMBL3759348 | C23H35ClN4O4 | 467.007 | 6 / 2 | 2.0 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218