

You can:
| Name | Metabotropic glutamate receptor 2 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | GRM2 |
| Synonym | mGluR2 mGlu2 receptor metabotropic glutamate receptor 2 GPRC1B glutamate receptor |
| Disease | Central nervous system disease Anxiety disorder Bipolar disorder Major depressive disorder Mood disorder [ Show all ] |
| Length | 872 |
| Amino acid sequence | MGSLLALLALLLLWGAVAEGPAKKVLTLEGDLVLGGLFPVHQKGGPAEDCGPVNEHRGIQRLEAMLFALDRINRDPHLLPGVRLGAHILDSCSKDTHALEQALDFVRASLSRGADGSRHICPDGSYATHGDAPTAITGVIGGSYSDVSIQVANLLRLFQIPQISYASTSAKLSDKSRYDYFARTVPPDFFQAKAMAEILRFFNWTYVSTVASEGDYGETGIEAFELEARARNICVATSEKVGRAMSRAAFEGVVRALLQKPSARVAVLFTRSEDARELLAASQRLNASFTWVASDGWGALESVVAGSEGAAEGAITIELASYPISDFASYFQSLDPWNNSRNPWFREFWEQRFRCSFRQRDCAAHSLRAVPFEQESKIMFVVNAVYAMAHALHNMHRALCPNTTRLCDAMRPVNGRRLYKDFVLNVKFDAPFRPADTHNEVRFDRFGDGIGRYNIFTYLRAGSGRYRYQKVGYWAEGLTLDTSLIPWASPSAGPLPASRCSEPCLQNEVKSVQPGEVCCWLCIPCQPYEYRLDEFTCADCGLGYWPNASLTGCFELPQEYIRWGDAWAVGPVTIACLGALATLFVLGVFVRHNATPVVKASGRELCYILLGGVFLCYCMTFIFIAKPSTAVCTLRRLGLGTAFSVCYSALLTKTNRIARIFGGAREGAQRPRFISPASQVAICLALISGQLLIVVAWLVVEAPGTGKETAPERREVVTLRCNHRDASMLGSLAYNVLLIALCTLYAFKTRKCPENFNEAKFIGFTMYTTCIIWLAFLPIFYVTSSDYRVQTTTMCVSVSLSGSVVLGCLFAPKLHIILFQPQKNVVSHRAPTSRFGSAAARASSSLGQGSGSQFVPTVCNGREVVDSTTSSL |
| UniProt | Q14416 |
| Protein Data Bank | 5cnj, 5kzn, 5kzq, 5cni, 4xas, 4xaq |
| GPCR-HGmod model | N/A |
| 3D structure model | This structure is from PDB ID 5cnj. |
| BioLiP | BL0365442, BL0324559,BL0324560, BL0324557,BL0324558, BL0306697,BL0306698, BL0306695,BL0306696, BL0365443 |
| Therapeutic Target Database | T62820 |
| ChEMBL | CHEMBL5137 |
| IUPHAR | 290 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 557316 | 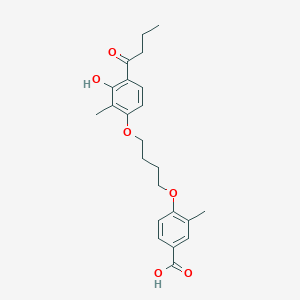 CHEMBL3287722 CHEMBL3287722 | C23H28O6 | 400.471 | 6 / 2 | 5.2 | No |
| 535924 | 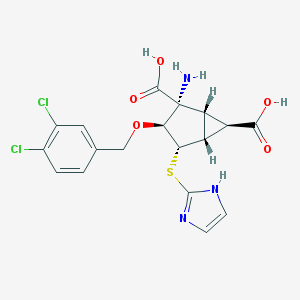 CHEMBL3977508 CHEMBL3977508 | C18H17Cl2N3O5S | 458.31 | 8 / 4 | -0.5 | Yes |
| 521452 | 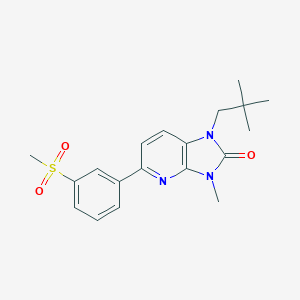 CHEMBL3765337 CHEMBL3765337 | C19H23N3O3S | 373.471 | 4 / 0 | 2.8 | Yes |
| 679 | 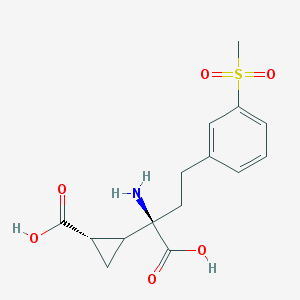 CHEMBL2111822 CHEMBL2111822 | C15H19NO6S | 341.378 | 7 / 3 | -2.1 | Yes |
| 557335 | 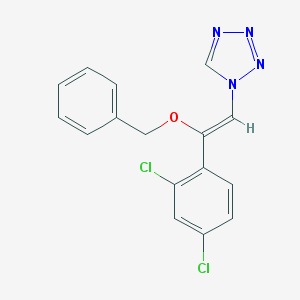 CHEMBL69204 CHEMBL69204 | C16H12Cl2N4O | 347.199 | 4 / 0 | 4.2 | Yes |
| 557342 | 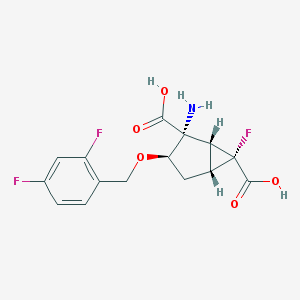 CHEMBL188511 CHEMBL188511 | C15H14F3NO5 | 345.274 | 9 / 3 | -1.7 | Yes |
| 1872 | 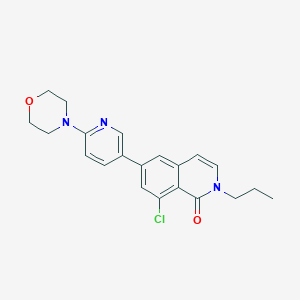 CHEMBL1669389 CHEMBL1669389 | C21H22ClN3O2 | 383.876 | 4 / 0 | 3.7 | Yes |
| 535965 | 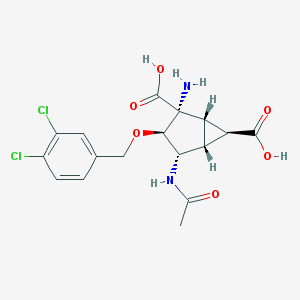 CHEMBL3913414 CHEMBL3913414 | C17H18Cl2N2O6 | 417.239 | 7 / 4 | -1.9 | Yes |
| 555487 | 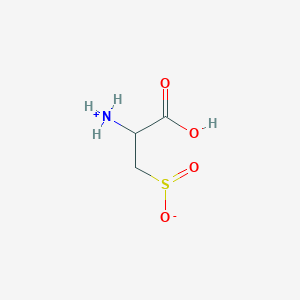 AC1N0YGA AC1N0YGA | C3H7NO4S | 153.152 | 5 / 2 | -4.7 | Yes |
| 536009 | 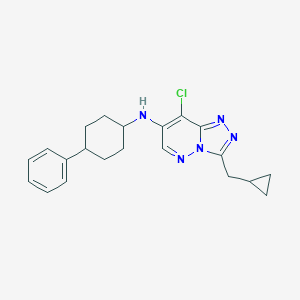 SCHEMBL16755980 SCHEMBL16755980 | C21H24ClN5 | 381.908 | 4 / 1 | 4.8 | Yes |
| 536017 | 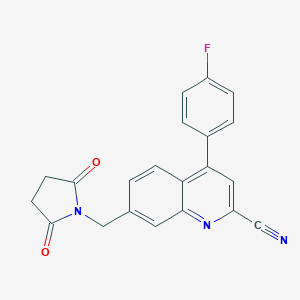 CHEMBL3904529 CHEMBL3904529 | C21H14FN3O2 | 359.36 | 5 / 0 | 2.7 | Yes |
| 557408 | 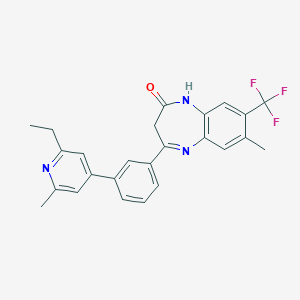 CHEMBL1629856 CHEMBL1629856 | C25H22F3N3O | 437.466 | 6 / 1 | 5.5 | No |
| 4839 | 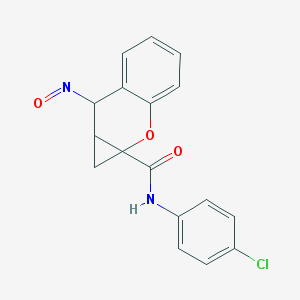 BDBM50332632 BDBM50332632 | C17H13ClN2O3 | 328.752 | 4 / 1 | 2.8 | Yes |
| 536062 | 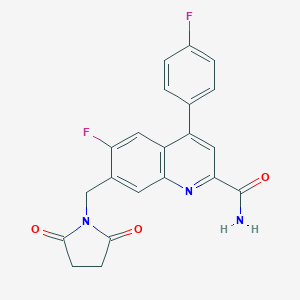 CHEMBL3951983 CHEMBL3951983 | C21H15F2N3O3 | 395.366 | 6 / 1 | 1.9 | Yes |
| 4942 | 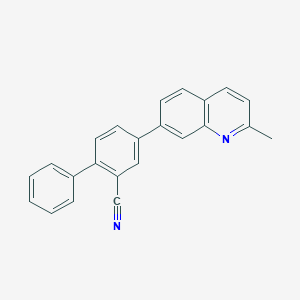 CHEMBL1099113 CHEMBL1099113 | C23H16N2 | 320.395 | 2 / 0 | 5.6 | No |
| 4965 | 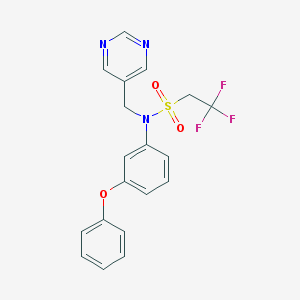 CHEMBL364784 CHEMBL364784 | C19H16F3N3O3S | 423.41 | 9 / 0 | 3.5 | Yes |
| 557434 | 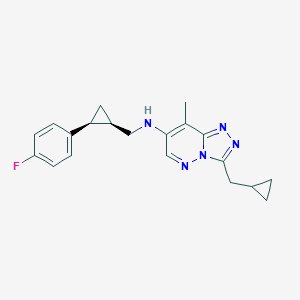 BDBM160615 BDBM160615 | C20H22FN5 | 351.429 | 5 / 1 | 3.7 | Yes |
| 557435 | 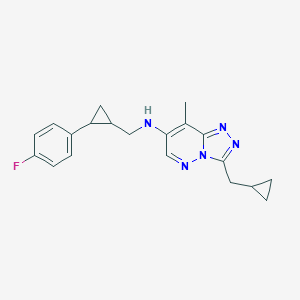 BDBM160605 BDBM160605 | C20H22FN5 | 351.429 | 5 / 1 | 3.7 | Yes |
| 557436 | 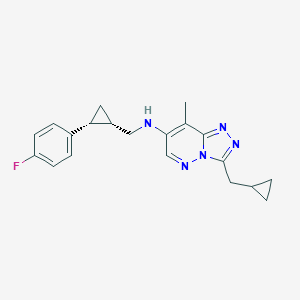 BDBM160641 BDBM160641 | C20H22FN5 | 351.429 | 5 / 1 | 3.7 | Yes |
| 557443 | 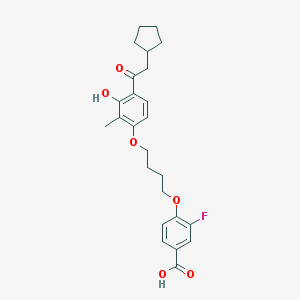 CHEMBL3287694 CHEMBL3287694 | C25H29FO6 | 444.499 | 7 / 2 | 6.3 | No |
| 536096 | 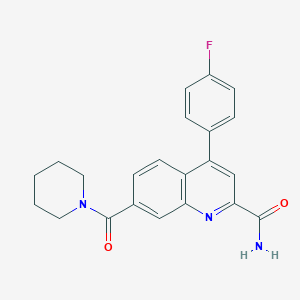 CHEMBL3970177 CHEMBL3970177 | C22H20FN3O2 | 377.419 | 4 / 1 | 3.4 | Yes |
| 547968 | 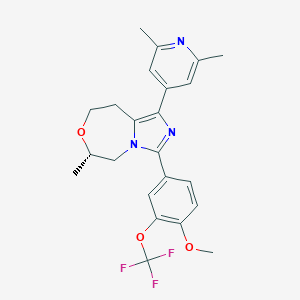 CHEMBL3972079 CHEMBL3972079 | C23H24F3N3O3 | 447.458 | 8 / 0 | 4.5 | Yes |
| 557469 | 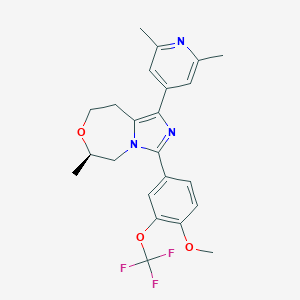 BDBM252757 BDBM252757 | C23H24F3N3O3 | 447.458 | 8 / 0 | 4.5 | Yes |
| 557474 | 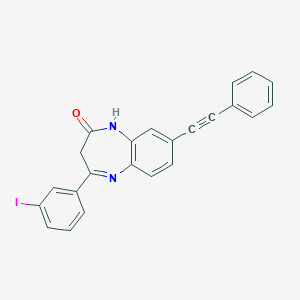 CHEMBL236035 CHEMBL236035 | C23H15IN2O | 462.29 | 2 / 1 | 5.1 | No |
| 441957 | 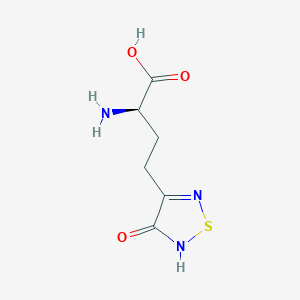 CHEMBL334842 CHEMBL334842 | C6H9N3O3S | 203.216 | 6 / 3 | -3.4 | Yes |
| 441960 | 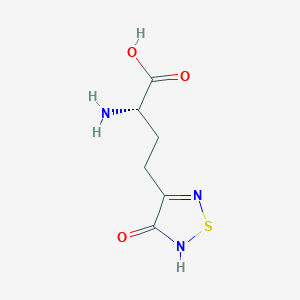 CHEMBL131922 CHEMBL131922 | C6H9N3O3S | 203.216 | 6 / 3 | -3.4 | Yes |
| 536137 | 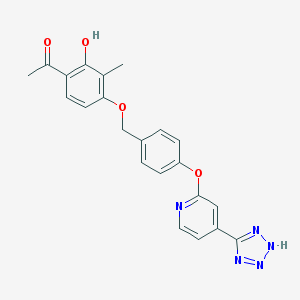 CHEMBL3899832 CHEMBL3899832 | C22H19N5O4 | 417.425 | 8 / 2 | 3.6 | Yes |
| 536159 | 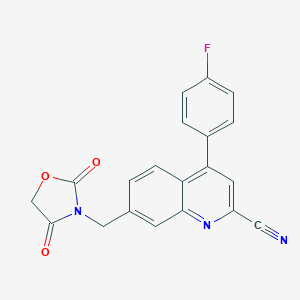 CHEMBL3986465 CHEMBL3986465 | C20H12FN3O3 | 361.332 | 6 / 0 | 3.3 | Yes |
| 557504 | 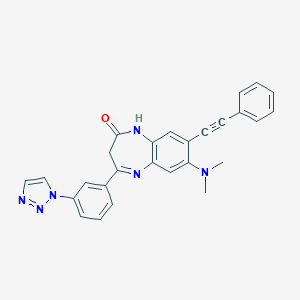 CHEMBL270402 CHEMBL270402 | C27H22N6O | 446.514 | 5 / 1 | 4.0 | Yes |
| 8265 | 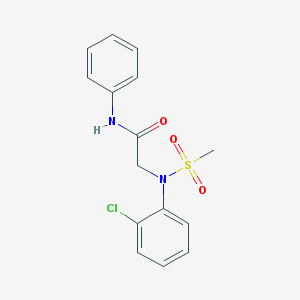 CHEMBL2208408 CHEMBL2208408 | C15H15ClN2O3S | 338.806 | 4 / 1 | 2.6 | Yes |
| 8380 | 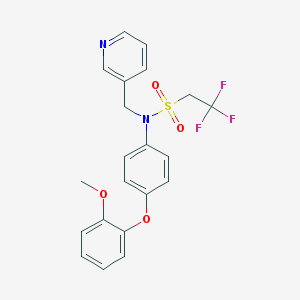 353231-17-1 353231-17-1 | C21H19F3N2O4S | 452.448 | 9 / 0 | 4.1 | Yes |
| 8473 | 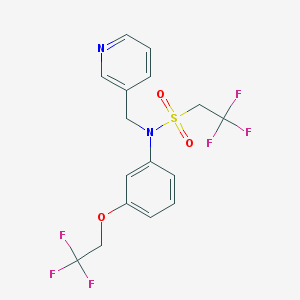 CHEMBL322465 CHEMBL322465 | C16H14F6N2O3S | 428.349 | 11 / 0 | 3.7 | No |
| 536190 | 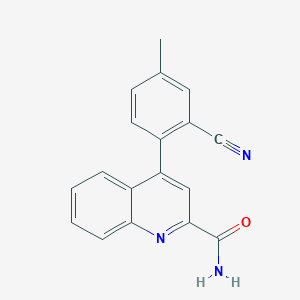 CHEMBL3907349 CHEMBL3907349 | C18H13N3O | 287.322 | 3 / 1 | 3.1 | Yes |
| 8695 | 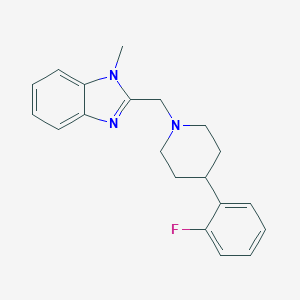 CHEMBL607689 CHEMBL607689 | C20H22FN3 | 323.415 | 3 / 0 | 3.6 | Yes |
| 8732 | 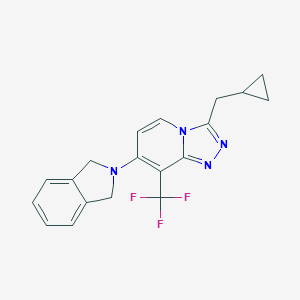 CHEMBL2179324 CHEMBL2179324 | C19H17F3N4 | 358.368 | 6 / 0 | 4.7 | Yes |
| 536246 | 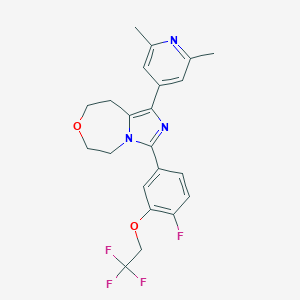 CHEMBL3984622 CHEMBL3984622 | C22H21F4N3O2 | 435.423 | 8 / 0 | 4.1 | Yes |
| 10163 | 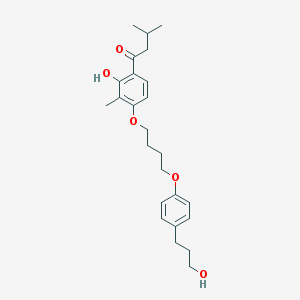 CHEMBL193707 CHEMBL193707 | C25H34O5 | 414.542 | 5 / 2 | 5.7 | No |
| 548002 | 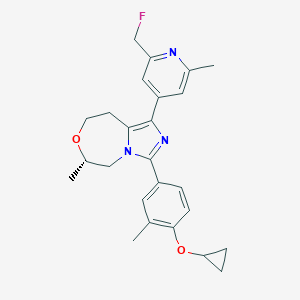 CHEMBL3938708 CHEMBL3938708 | C25H28FN3O2 | 421.516 | 5 / 0 | 4.0 | Yes |
| 557579 | 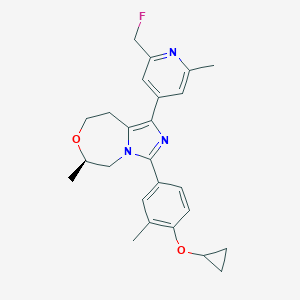 BDBM252811 BDBM252811 | C25H28FN3O2 | 421.516 | 5 / 0 | 4.0 | Yes |
| 10404 | 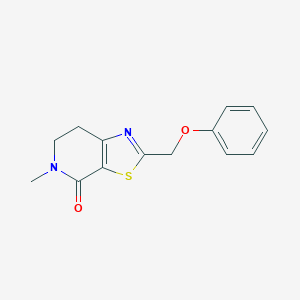 CHEMBL2426616 CHEMBL2426616 | C14H14N2O2S | 274.338 | 4 / 0 | 2.3 | Yes |
| 536255 | 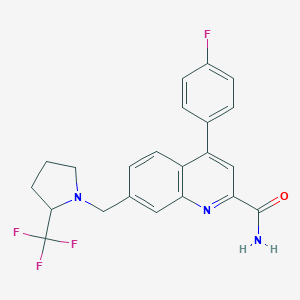 CHEMBL3913806 CHEMBL3913806 | C22H19F4N3O | 417.408 | 7 / 1 | 4.6 | Yes |
| 557595 | 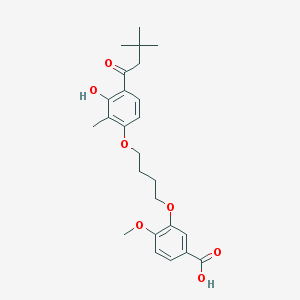 CHEMBL3287723 CHEMBL3287723 | C25H32O7 | 444.524 | 7 / 2 | 5.7 | No |
| 10773 | 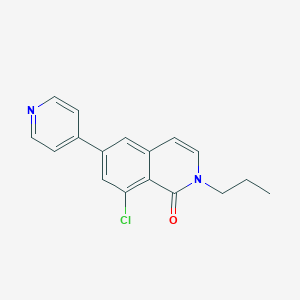 CHEMBL1669380 CHEMBL1669380 | C17H15ClN2O | 298.77 | 2 / 0 | 3.6 | Yes |
| 548014 | 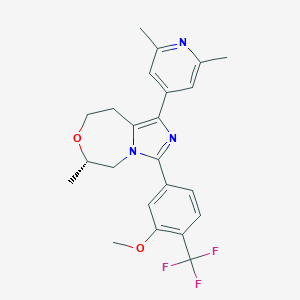 CHEMBL3892017 CHEMBL3892017 | C23H24F3N3O2 | 431.459 | 7 / 0 | 4.2 | Yes |
| 557608 | 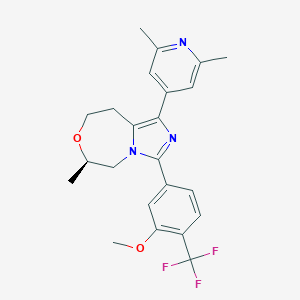 SCHEMBL15969131 SCHEMBL15969131 | C23H24F3N3O2 | 431.459 | 7 / 0 | 4.2 | Yes |
| 11484 | 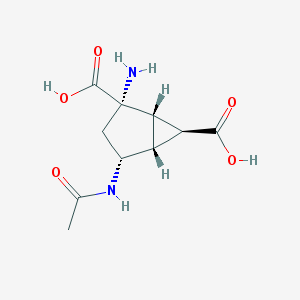 CHEMBL2381648 CHEMBL2381648 | C10H14N2O5 | 242.231 | 6 / 4 | -4.2 | Yes |
| 11487 | 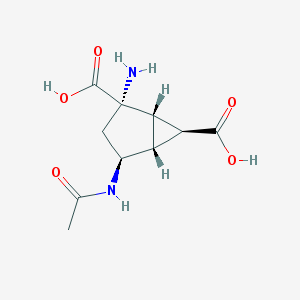 CHEMBL2381649 CHEMBL2381649 | C10H14N2O5 | 242.231 | 6 / 4 | -4.2 | Yes |
| 557620 | 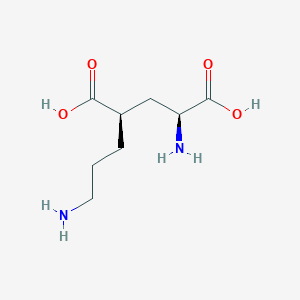 CHEMBL2312685 CHEMBL2312685 | C8H16N2O4 | 204.226 | 6 / 4 | -6.8 | Yes |
| 557639 | 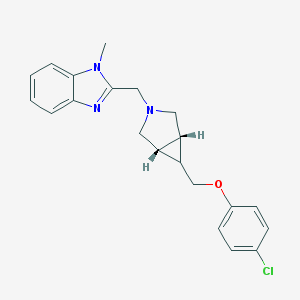 CHEMBL477170 CHEMBL477170 | C21H22ClN3O | 367.877 | 3 / 0 | 3.9 | Yes |
| 12370 | 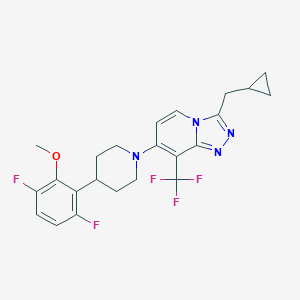 CHEMBL2206449 CHEMBL2206449 | C23H23F5N4O | 466.456 | 9 / 0 | 6.0 | No |
| 555529 | 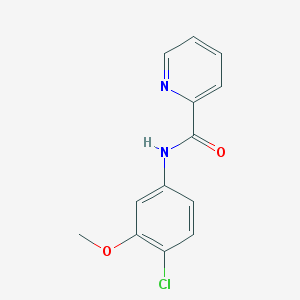 VU 0361737 VU 0361737 | C13H11ClN2O2 | 262.693 | 3 / 1 | 2.6 | Yes |
| 13313 | 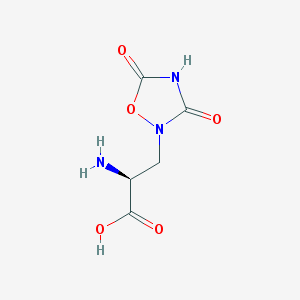 QUISQUALIC ACID QUISQUALIC ACID | C5H7N3O5 | 189.127 | 6 / 3 | -3.9 | Yes |
| 13333 | 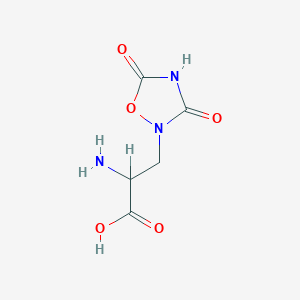 2-amino-3-(3,5-dioxo-1,2,4-oxadiazolidin-2-yl)propanoic acid 2-amino-3-(3,5-dioxo-1,2,4-oxadiazolidin-2-yl)propanoic acid | C5H7N3O5 | 189.127 | 6 / 3 | -3.9 | Yes |
| 13373 | 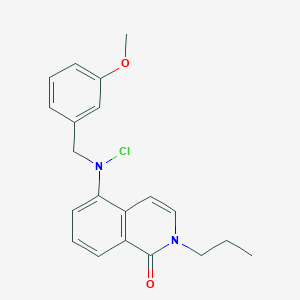 CHEMBL3221847 CHEMBL3221847 | C20H21ClN2O2 | 356.85 | 3 / 0 | 4.7 | Yes |
| 536355 | 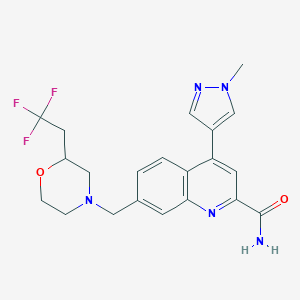 CHEMBL3979288 CHEMBL3979288 | C21H22F3N5O2 | 433.435 | 8 / 1 | 2.3 | Yes |
| 557662 | 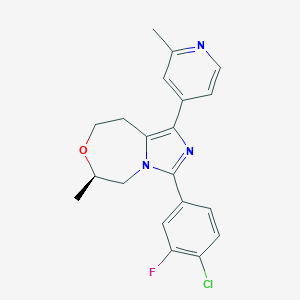 BDBM252661 BDBM252661 | C20H19ClFN3O | 371.84 | 4 / 0 | 3.7 | Yes |
| 548038 | 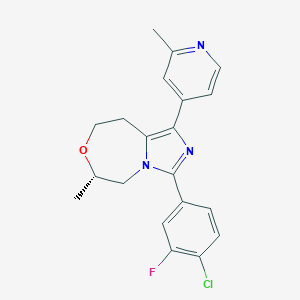 CHEMBL3945633 CHEMBL3945633 | C20H19ClFN3O | 371.84 | 4 / 0 | 3.7 | Yes |
| 14207 | 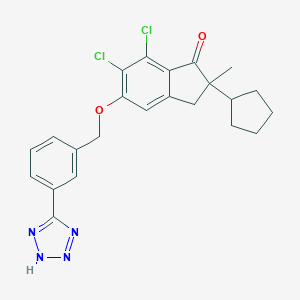 CHEMBL181869 CHEMBL181869 | C23H22Cl2N4O2 | 457.355 | 5 / 1 | 6.2 | No |
| 548041 | 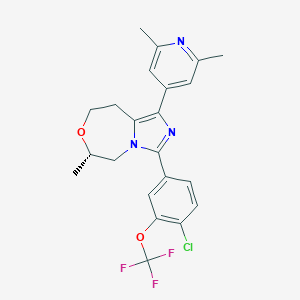 CHEMBL3893853 CHEMBL3893853 | C22H21ClF3N3O2 | 451.874 | 7 / 0 | 5.2 | No |
| 557667 | 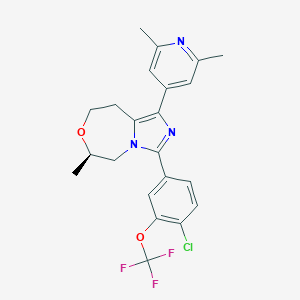 BDBM252671 BDBM252671 | C22H21ClF3N3O2 | 451.874 | 7 / 0 | 5.2 | No |
| 14352 | 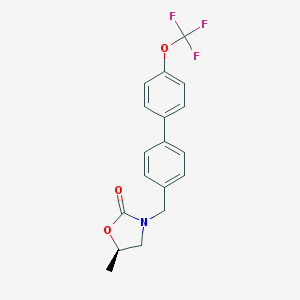 CHEMBL483561 CHEMBL483561 | C18H16F3NO3 | 351.325 | 6 / 0 | 4.6 | Yes |
| 15035 | 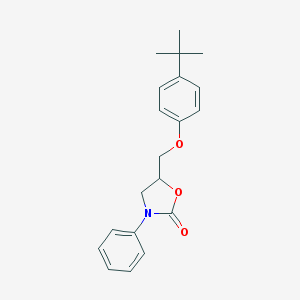 CHEMBL1096713 CHEMBL1096713 | C20H23NO3 | 325.408 | 3 / 0 | 4.9 | Yes |
| 557688 | 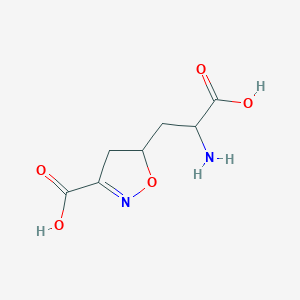 CHEMBL197629 CHEMBL197629 | C7H10N2O5 | 202.166 | 7 / 3 | -3.4 | Yes |
| 15208 | 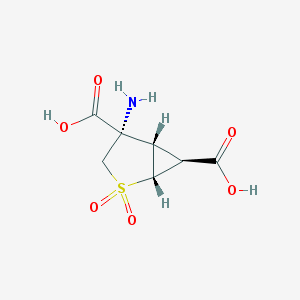 635318-11-5 635318-11-5 | C7H9NO6S | 235.21 | 7 / 3 | -4.8 | Yes |
| 521916 | 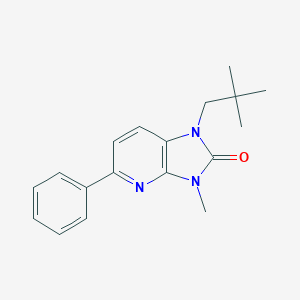 CHEMBL3763395 CHEMBL3763395 | C18H21N3O | 295.386 | 2 / 0 | 3.6 | Yes |
| 16005 | 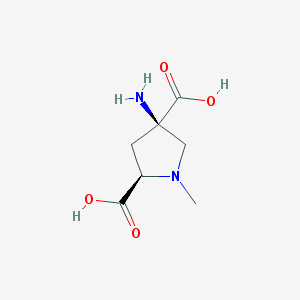 CHEMBL300330 CHEMBL300330 | C7H12N2O4 | 188.183 | 6 / 3 | -6.0 | Yes |
| 557722 | 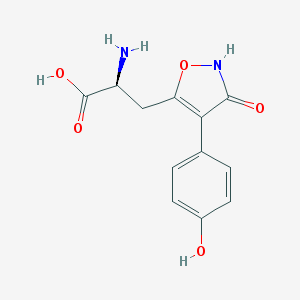 CHEMBL367027 CHEMBL367027 | C12H12N2O5 | 264.237 | 6 / 4 | -2.3 | Yes |
| 557739 | 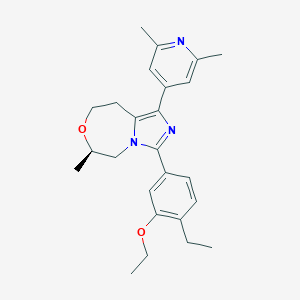 BDBM252823 BDBM252823 | C25H31N3O2 | 405.542 | 4 / 0 | 4.5 | Yes |
| 548067 | 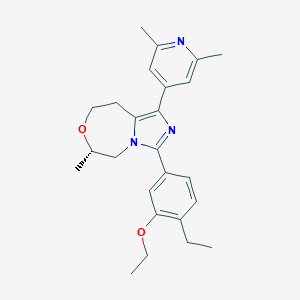 CHEMBL3889702 CHEMBL3889702 | C25H31N3O2 | 405.542 | 4 / 0 | 4.5 | Yes |
| 557741 | 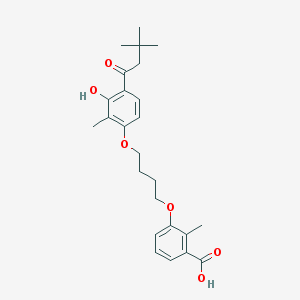 CHEMBL3287725 CHEMBL3287725 | C25H32O6 | 428.525 | 6 / 2 | 6.1 | No |
| 536423 | 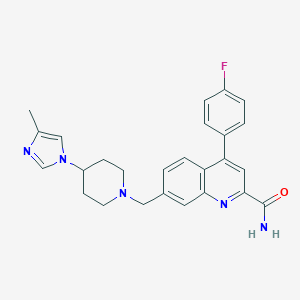 CHEMBL3936050 CHEMBL3936050 | C26H26FN5O | 443.526 | 5 / 1 | 3.6 | Yes |
| 548079 | 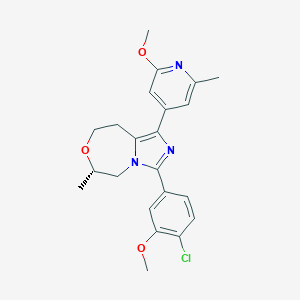 CHEMBL3921250 CHEMBL3921250 | C22H24ClN3O3 | 413.902 | 5 / 0 | 3.9 | Yes |
| 557758 | 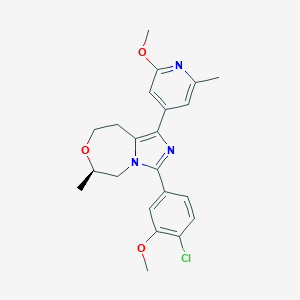 BDBM252660 BDBM252660 | C22H24ClN3O3 | 413.902 | 5 / 0 | 3.9 | Yes |
| 536433 | 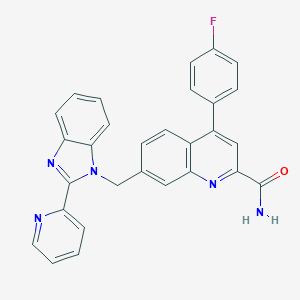 CHEMBL3909260 CHEMBL3909260 | C29H20FN5O | 473.511 | 5 / 1 | 4.9 | Yes |
| 557776 | 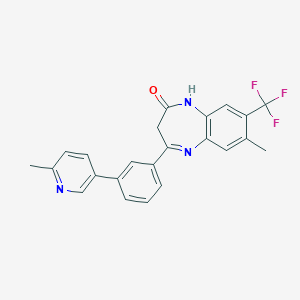 CHEMBL1629845 CHEMBL1629845 | C23H18F3N3O | 409.412 | 6 / 1 | 4.6 | Yes |
| 557810 | 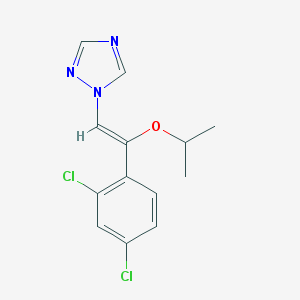 CHEMBL304285 CHEMBL304285 | C13H13Cl2N3O | 298.167 | 3 / 0 | 3.9 | Yes |
| 18230 | 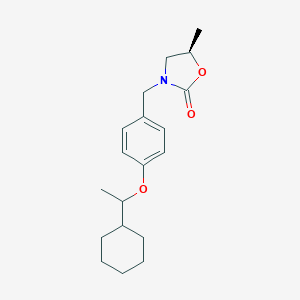 CHEMBL484340 CHEMBL484340 | C19H27NO3 | 317.429 | 3 / 0 | 4.6 | Yes |
| 18231 | 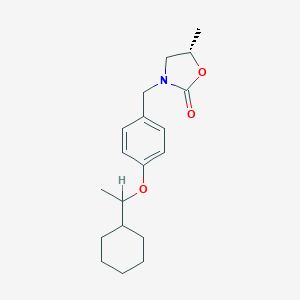 CHEMBL485336 CHEMBL485336 | C19H27NO3 | 317.429 | 3 / 0 | 4.6 | Yes |
| 18232 | 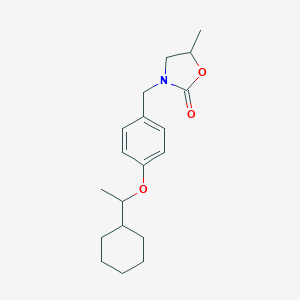 CHEMBL484731 CHEMBL484731 | C19H27NO3 | 317.429 | 3 / 0 | 4.6 | Yes |
| 18336 | 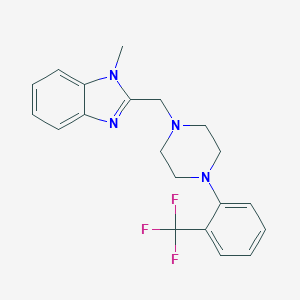 CHEMBL603443 CHEMBL603443 | C20H21F3N4 | 374.411 | 6 / 0 | 3.8 | Yes |
| 557838 | 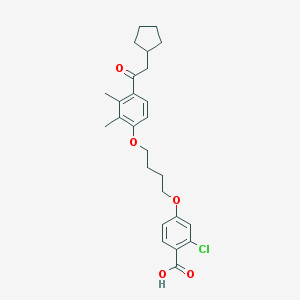 CHEMBL3287682 CHEMBL3287682 | C26H31ClO5 | 458.979 | 5 / 1 | 7.0 | No |
| 536459 | 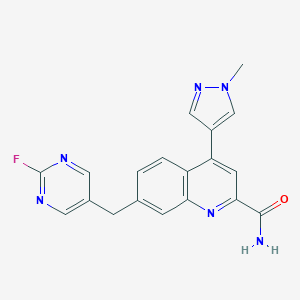 CHEMBL3897362 CHEMBL3897362 | C19H15FN6O | 362.368 | 6 / 1 | 2.0 | Yes |
| 557847 | 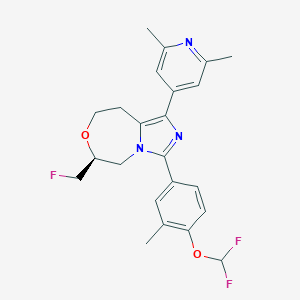 SCHEMBL15969145 SCHEMBL15969145 | C23H24F3N3O2 | 431.459 | 7 / 0 | 4.6 | Yes |
| 548106 | 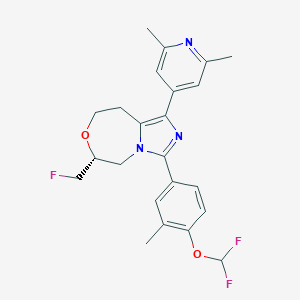 CHEMBL3955717 CHEMBL3955717 | C23H24F3N3O2 | 431.459 | 7 / 0 | 4.6 | Yes |
| 557861 | 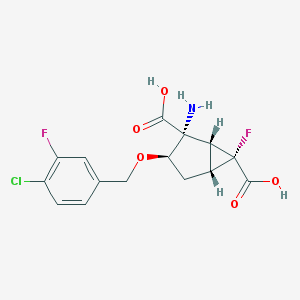 CHEMBL362325 CHEMBL362325 | C15H14ClF2NO5 | 361.726 | 8 / 3 | -1.2 | Yes |
| 557865 | 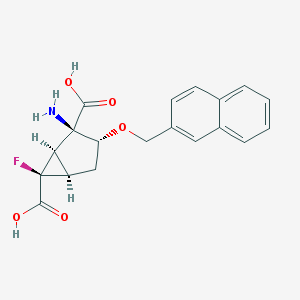 CHEMBL185210 CHEMBL185210 | C19H18FNO5 | 359.353 | 7 / 3 | -0.7 | Yes |
| 19476 | 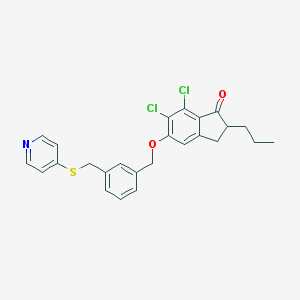 CHEMBL181364 CHEMBL181364 | C25H23Cl2NO2S | 472.424 | 4 / 0 | 6.8 | No |
| 19615 | 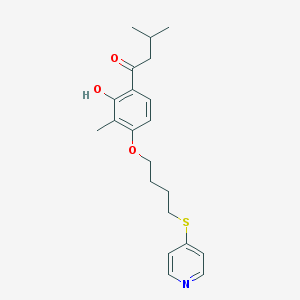 CHEMBL183319 CHEMBL183319 | C21H27NO3S | 373.511 | 5 / 1 | 5.3 | No |
| 19636 | 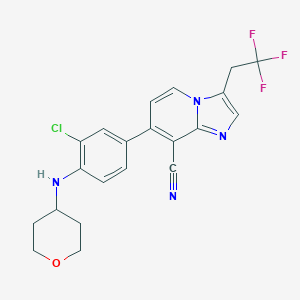 CHEMBL2152108 CHEMBL2152108 | C21H18ClF3N4O | 434.847 | 7 / 1 | 5.5 | No |
| 19689 | 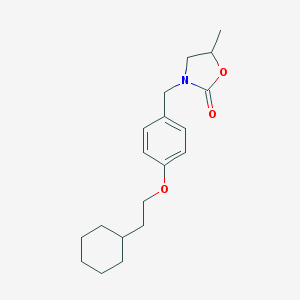 CHEMBL444778 CHEMBL444778 | C19H27NO3 | 317.429 | 3 / 0 | 4.9 | Yes |
| 548117 | 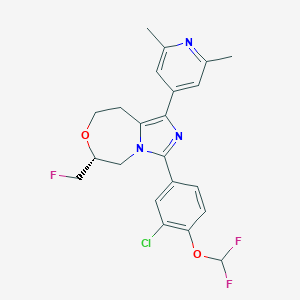 CHEMBL3899252 CHEMBL3899252 | C22H21ClF3N3O2 | 451.874 | 7 / 0 | 4.9 | Yes |
| 557889 | 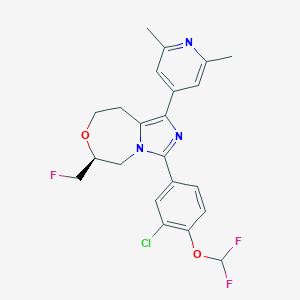 SCHEMBL18104073 SCHEMBL18104073 | C22H21ClF3N3O2 | 451.874 | 7 / 0 | 4.9 | Yes |
| 536486 | 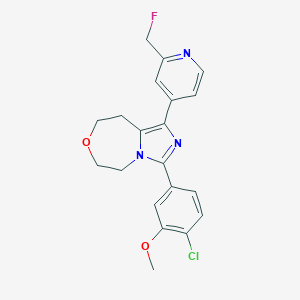 CHEMBL3972441 CHEMBL3972441 | C20H19ClFN3O2 | 387.839 | 5 / 0 | 2.9 | Yes |
| 465376 | 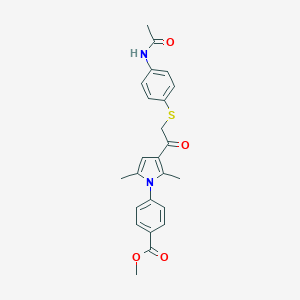 AC1N2AWG AC1N2AWG | C24H24N2O4S | 436.526 | 5 / 1 | 4.1 | Yes |
| 536514 | 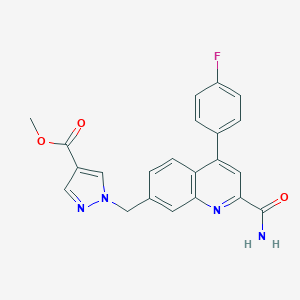 CHEMBL3969686 CHEMBL3969686 | C22H17FN4O3 | 404.401 | 6 / 1 | 2.9 | Yes |
| 548128 | 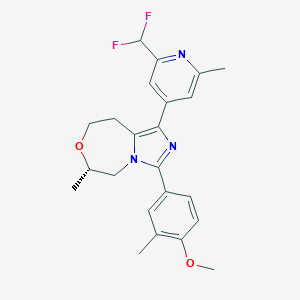 CHEMBL3947078 CHEMBL3947078 | C23H25F2N3O2 | 413.469 | 6 / 0 | 3.9 | Yes |
| 557946 | 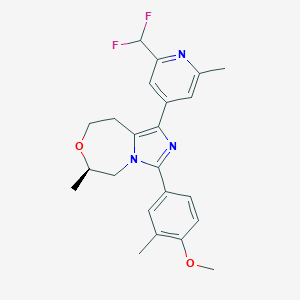 BDBM252781 BDBM252781 | C23H25F2N3O2 | 413.469 | 6 / 0 | 3.9 | Yes |
| 557954 | 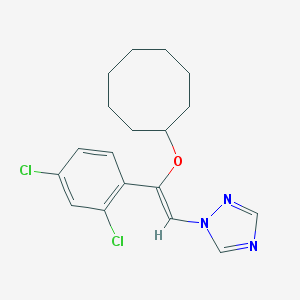 CHEMBL68739 CHEMBL68739 | C18H21Cl2N3O | 366.286 | 3 / 0 | 6.0 | No |
| 557959 | 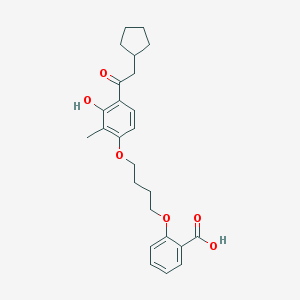 CHEMBL3287680 CHEMBL3287680 | C25H30O6 | 426.509 | 6 / 2 | 6.2 | No |
| 22046 | 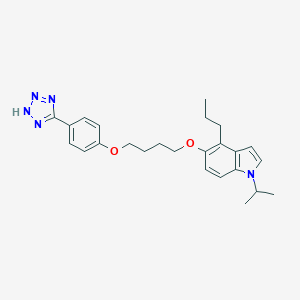 CHEMBL191391 CHEMBL191391 | C25H31N5O2 | 433.556 | 5 / 1 | 5.6 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218