

You can:
| Name | P2Y purinoceptor 14 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | P2RY14 |
| Synonym | G-protein coupled receptor 105 GPR105 G protein-coupled receptor 105 P2Y purinoceptor 14 P2Y14 [ Show all ] |
| Disease | N/A |
| Length | 338 |
| Amino acid sequence | MINSTSTQPPDESCSQNLLITQQIIPVLYCMVFIAGILLNGVSGWIFFYVPSSKSFIIYLKNIVIADFVMSLTFPFKILGDSGLGPWQLNVFVCRVSAVLFYVNMYVSIVFFGLISFDRYYKIVKPLWTSFIQSVSYSKLLSVIVWMLMLLLAVPNIILTNQSVREVTQIKCIELKSELGRKWHKASNYIFVAIFWIVFLLLIVFYTAITKKIFKSHLKSSRNSTSVKKKSSRNIFSIVFVFFVCFVPYHIARIPYTKSQTEAHYSCQSKEILRYMKEFTLLLSAANVCLDPIIYFFLCQPFREILCKKLHIPLKAQNDLDISRIKRGNTTLESTDTL |
| UniProt | Q15391 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q15391 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q15391. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL4518 |
| IUPHAR | 330 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 4854 | 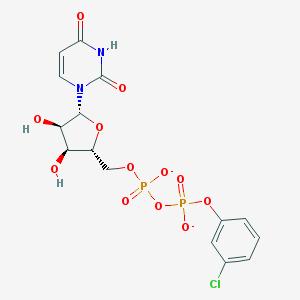 BDBM50303350 BDBM50303350 | C15H15ClN2O12P2-2 | 512.685 | 12 / 3 | -1.9 | No |
| 5497 | 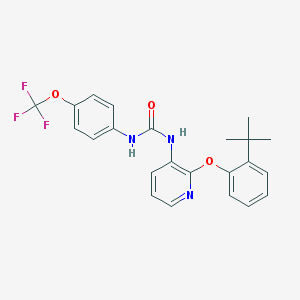 BPTU BPTU | C23H22F3N3O3 | 445.442 | 7 / 2 | 6.1 | No |
| 5640 | 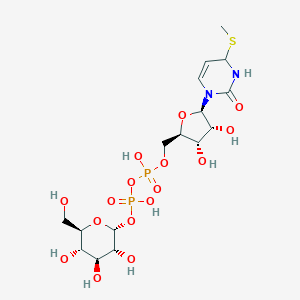 CHEMBL439378 CHEMBL439378 | C16H28N2O16P2S | 598.406 | 17 / 9 | -5.5 | No |
| 10835 | 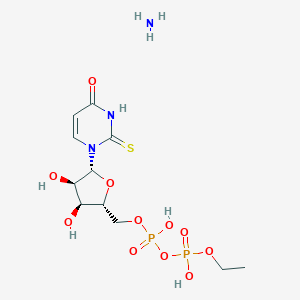 CHEMBL1162563 CHEMBL1162563 | C11H21N3O11P2S | 465.307 | 13 / 6 | N/A | No |
| 15819 | 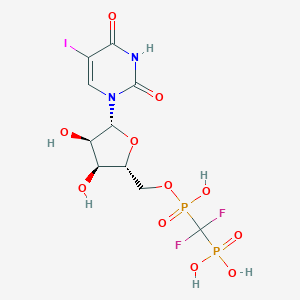 CHEMBL566925 CHEMBL566925 | C10H13F2IN2O11P2 | 564.066 | 13 / 6 | -3.8 | No |
| 24857 | 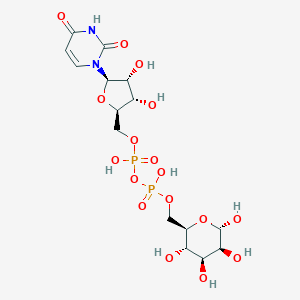 CHEMBL596145 CHEMBL596145 | C15H24N2O17P2 | 566.302 | 17 / 9 | -6.9 | No |
| 24858 | 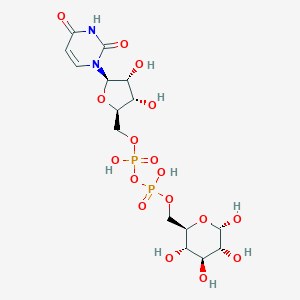 CHEMBL594065 CHEMBL594065 | C15H24N2O17P2 | 566.302 | 17 / 9 | -6.9 | No |
| 27554 | 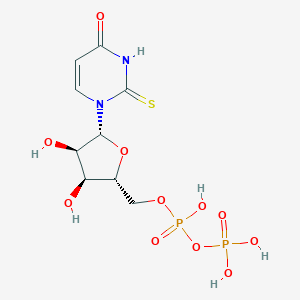 2-thio-UDP 2-thio-UDP | C9H14N2O11P2S | 420.222 | 12 / 6 | -4.1 | No |
| 31386 | 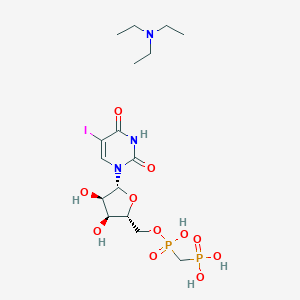 CHEMBL1085561 CHEMBL1085561 | C16H30IN3O11P2 | 629.278 | 12 / 6 | N/A | No |
| 34046 | 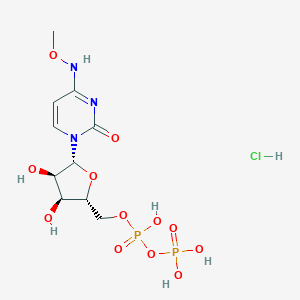 CHEMBL1204010 CHEMBL1204010 | C10H18ClN3O12P2 | 469.661 | 12 / 7 | N/A | No |
| 37983 | 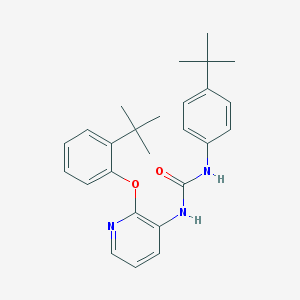 CHEMBL2333771 CHEMBL2333771 | C26H31N3O2 | 417.553 | 3 / 2 | 6.6 | No |
| 47052 | 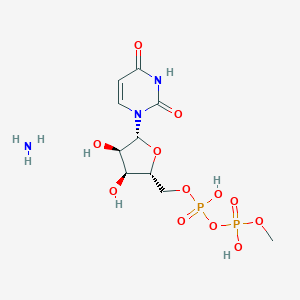 CHEMBL1162554 CHEMBL1162554 | C10H19N3O12P2 | 435.219 | 13 / 6 | N/A | No |
| 459672 | 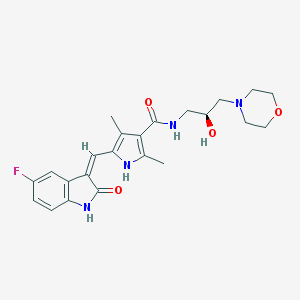 452105-23-6 452105-23-6 | C23H27FN4O4 | 442.491 | 6 / 4 | 0.9 | Yes |
| 66858 | 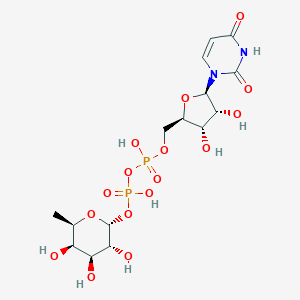 CHEMBL595940 CHEMBL595940 | C15H24N2O16P2 | 550.303 | 16 / 8 | -6.1 | No |
| 69329 | 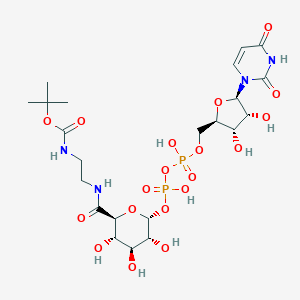 CHEMBL594703 CHEMBL594703 | C22H36N4O19P2 | 722.487 | 19 / 10 | -5.9 | No |
| 553627 | 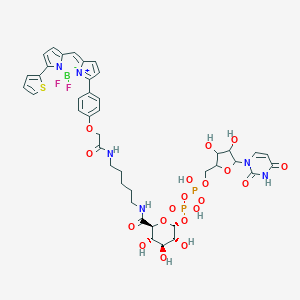 GTPL9471 GTPL9471 | C41H47BF2N6O19P2S | 1070.66 | 23 / 10 | N/A | No |
| 73200 | 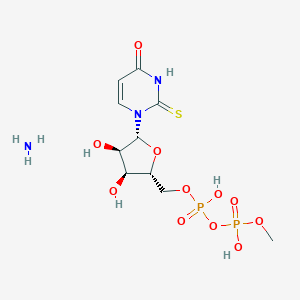 CHEMBL1162557 CHEMBL1162557 | C10H19N3O11P2S | 451.28 | 13 / 6 | N/A | No |
| 74633 | 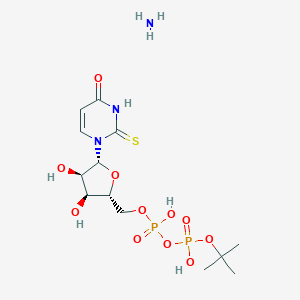 CHEMBL1162556 CHEMBL1162556 | C13H25N3O11P2S | 493.361 | 13 / 6 | N/A | No |
| 81349 | 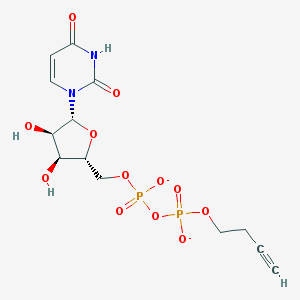 BDBM50303345 BDBM50303345 | C13H16N2O12P2-2 | 454.221 | 12 / 3 | -3.8 | No |
| 81558 | 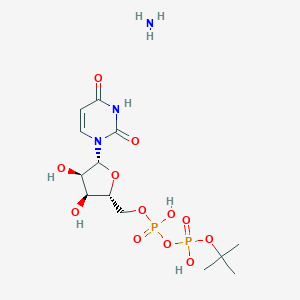 CHEMBL1162552 CHEMBL1162552 | C13H25N3O12P2 | 477.3 | 13 / 6 | N/A | No |
| 553675 | 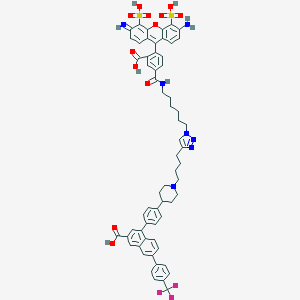 CID 124221651 CID 124221651 | C62H58F3N7O12S2 | 1214.3 | 20 / 7 | 6.5 | No |
| 553682 | 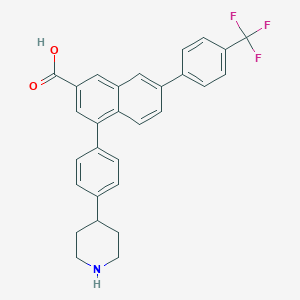 PPTN PPTN | C29H24F3NO2 | 475.511 | 6 / 2 | 4.7 | Yes |
| 89252 | 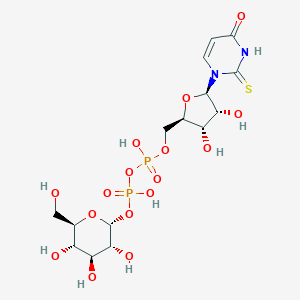 CHEMBL228111 CHEMBL228111 | C15H24N2O16P2S | 582.363 | 17 / 9 | -6.0 | No |
| 460047 | 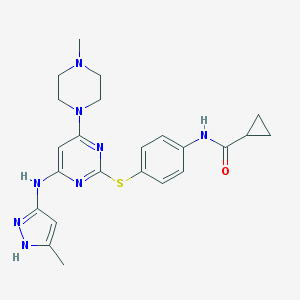 Tozasertib Tozasertib | C23H28N8OS | 464.592 | 8 / 3 | 3.4 | Yes |
| 102461 | 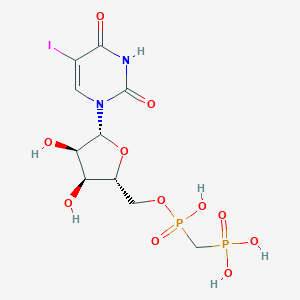 CHEMBL1085561 CHEMBL1085561 | C10H15IN2O11P2 | 528.085 | 11 / 6 | -4.5 | No |
| 102532 | 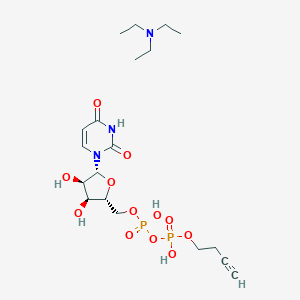 CHEMBL612064 CHEMBL612064 | C19H33N3O12P2 | 557.43 | 13 / 5 | N/A | No |
| 104005 | 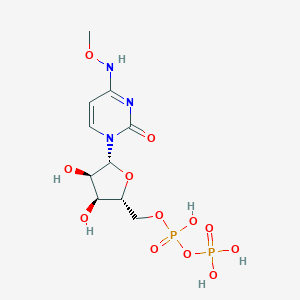 CHEMBL1084020 CHEMBL1084020 | C10H17N3O12P2 | 433.203 | 12 / 6 | -4.1 | No |
| 111093 | 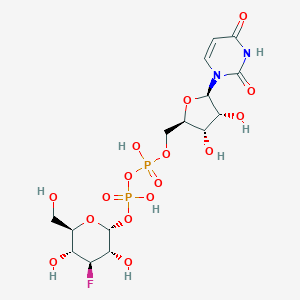 CHEMBL594064 CHEMBL594064 | C15H23FN2O16P2 | 568.293 | 17 / 8 | -5.3 | No |
| 112446 | 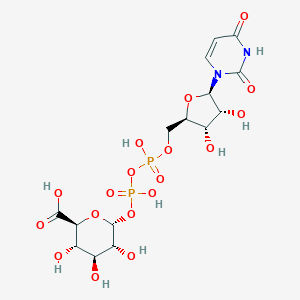 Udp-glucuronic acid Udp-glucuronic acid | C15H22N2O18P2 | 580.285 | 18 / 9 | -6.4 | No |
| 112449 | 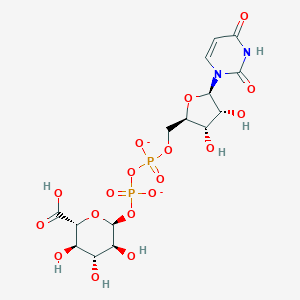 BDBM50197975 BDBM50197975 | C15H20N2O18P2-2 | 578.269 | 18 / 7 | -6.6 | No |
| 112450 | 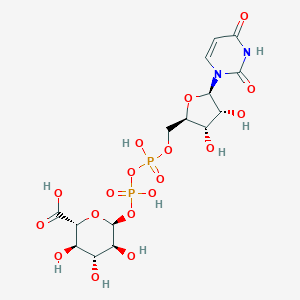 CHEMBL1162071 CHEMBL1162071 | C15H22N2O18P2 | 580.285 | 18 / 9 | -6.4 | No |
| 553827 | 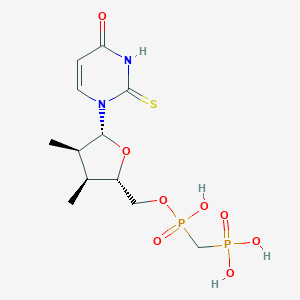 alpha,beta-methylene-UDP alpha,beta-methylene-UDP | C12H20N2O8P2S | 414.306 | 9 / 4 | -1.8 | Yes |
| 115568 | 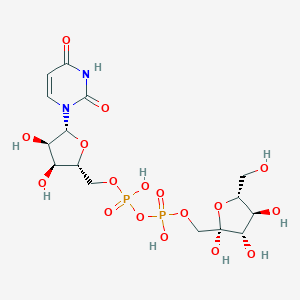 CHEMBL226713 CHEMBL226713 | C15H24N2O17P2 | 566.302 | 17 / 9 | -6.9 | No |
| 460272 | 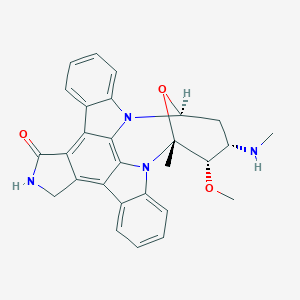 staurosporine staurosporine | C28H26N4O3 | 466.541 | 4 / 2 | 3.2 | Yes |
| 122637 | 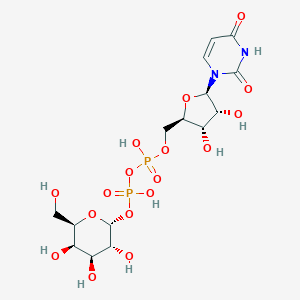 Udp galactose Udp galactose | C15H24N2O17P2 | 566.302 | 17 / 9 | -6.3 | No |
| 122641 | 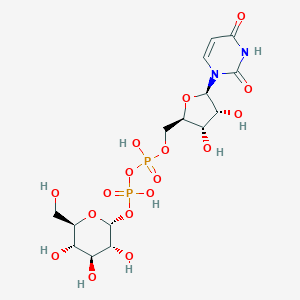 UDP-glucose UDP-glucose | C15H24N2O17P2 | 566.302 | 17 / 9 | -6.3 | No |
| 122647 | 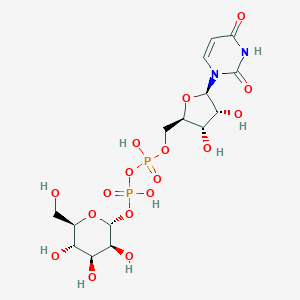 URIDINE-5'-DIPHOSPHATE-MANNOSE URIDINE-5'-DIPHOSPHATE-MANNOSE | C15H24N2O17P2 | 566.302 | 17 / 9 | -6.3 | No |
| 122648 | 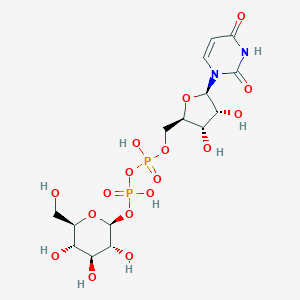 CHEMBL595216 CHEMBL595216 | C15H24N2O17P2 | 566.302 | 17 / 9 | -6.3 | No |
| 128670 | 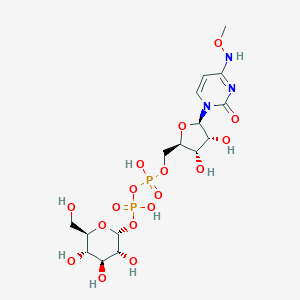 CHEMBL594086 CHEMBL594086 | C16H27N3O17P2 | 595.344 | 17 / 9 | -6.5 | No |
| 140510 | 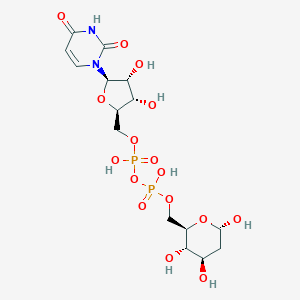 CHEMBL604700 CHEMBL604700 | C15H24N2O16P2 | 550.303 | 16 / 8 | -5.9 | No |
| 149898 | 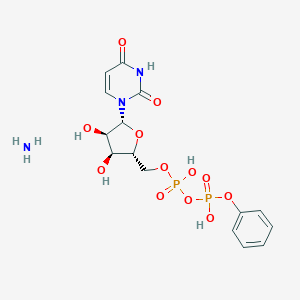 CHEMBL1162559 CHEMBL1162559 | C15H21N3O12P2 | 497.29 | 13 / 6 | N/A | No |
| 153078 | 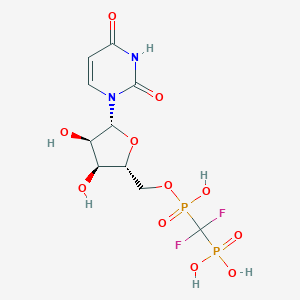 CHEMBL488133 CHEMBL488133 | C10H14F2N2O11P2 | 438.169 | 13 / 6 | -4.1 | No |
| 554029 | 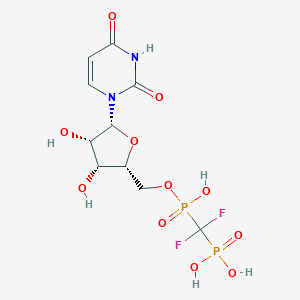 MRS2802 MRS2802 | C10H14F2N2O11P2 | 438.169 | 13 / 6 | -4.1 | No |
| 160480 | 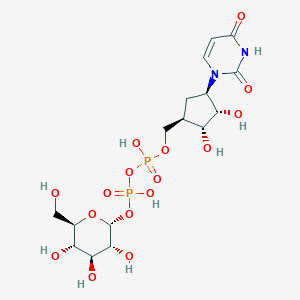 CHEMBL604267 CHEMBL604267 | C16H26N2O16P2 | 564.33 | 16 / 9 | -6.0 | No |
| 164815 | 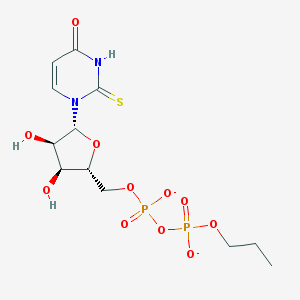 BDBM50303344 BDBM50303344 | C12H18N2O11P2S-2 | 460.287 | 12 / 3 | -2.9 | No |
| 178360 | 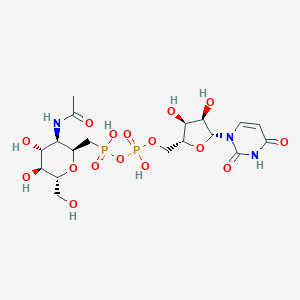 UDM UDM | C18H29N3O16P2 | 605.383 | 16 / 9 | -6.8 | No |
| 554207 | 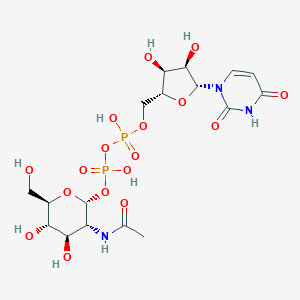 UDP-N-acetylglucosamine UDP-N-acetylglucosamine | C17H27N3O17P2 | 607.355 | 17 / 9 | -6.6 | No |
| 186298 | 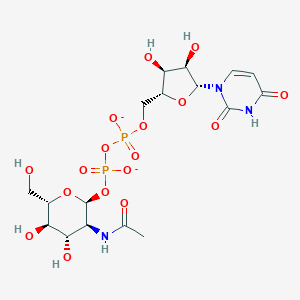 UDP-N-acetyl-glucosamine UDP-N-acetyl-glucosamine | C17H25N3O17P2-2 | 605.339 | 17 / 7 | -6.5 | No |
| 186299 | 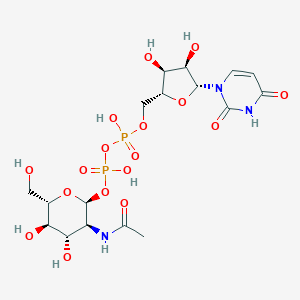 CHEMBL1162072 CHEMBL1162072 | C17H27N3O17P2 | 607.355 | 17 / 9 | -6.6 | No |
| 187629 | 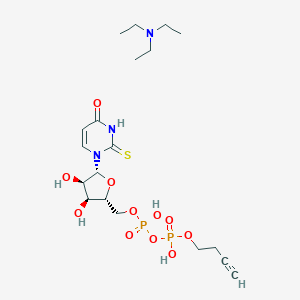 CHEMBL611792 CHEMBL611792 | C19H33N3O11P2S | 573.491 | 13 / 5 | N/A | No |
| 191279 | 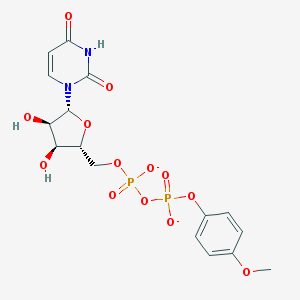 BDBM50303337 BDBM50303337 | C16H18N2O13P2-2 | 508.269 | 13 / 3 | -2.6 | No |
| 196758 | 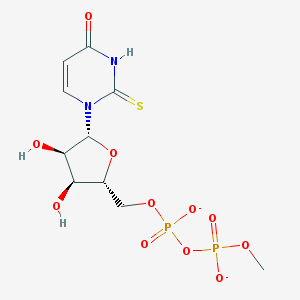 BDBM50303342 BDBM50303342 | C10H14N2O11P2S-2 | 432.233 | 12 / 3 | -3.7 | No |
| 461065 | 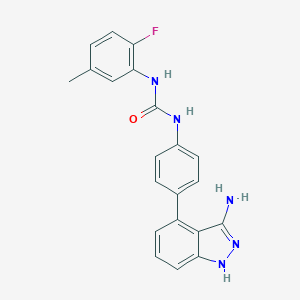 Linifanib Linifanib | C21H18FN5O | 375.407 | 4 / 4 | 3.9 | Yes |
| 214497 | 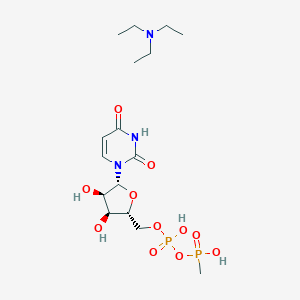 CHEMBL612063 CHEMBL612063 | C16H31N3O11P2 | 503.382 | 12 / 5 | N/A | No |
| 218951 | 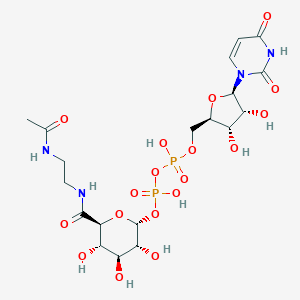 CHEMBL595076 CHEMBL595076 | C19H30N4O18P2 | 664.407 | 18 / 10 | -7.3 | No |
| 222686 | 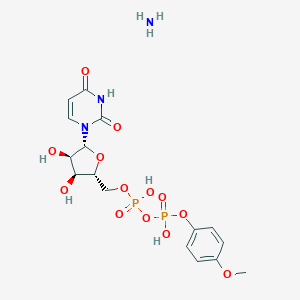 CHEMBL1162562 CHEMBL1162562 | C16H23N3O13P2 | 527.316 | 14 / 6 | N/A | No |
| 222907 | 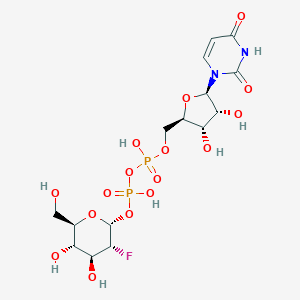 CHEMBL593830 CHEMBL593830 | C15H23FN2O16P2 | 568.293 | 17 / 8 | -5.6 | No |
| 226809 | 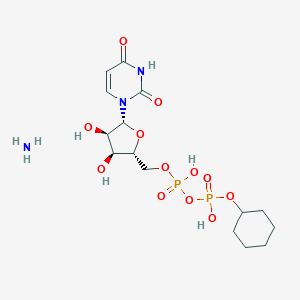 CHEMBL1162558 CHEMBL1162558 | C15H27N3O12P2 | 503.338 | 13 / 6 | N/A | No |
| 231965 | 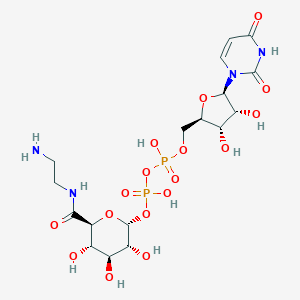 CHEMBL605512 CHEMBL605512 | C17H28N4O17P2 | 622.37 | 18 / 10 | -9.9 | No |
| 232024 | 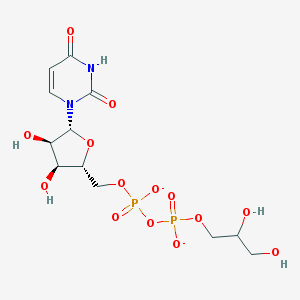 BDBM50303334 BDBM50303334 | C12H18N2O14P2-2 | 476.224 | 14 / 5 | -5.7 | No |
| 236550 | 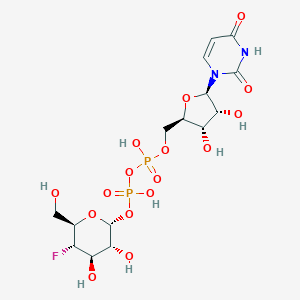 CHEMBL593125 CHEMBL593125 | C15H23FN2O16P2 | 568.293 | 17 / 8 | -6.2 | No |
| 461270 | 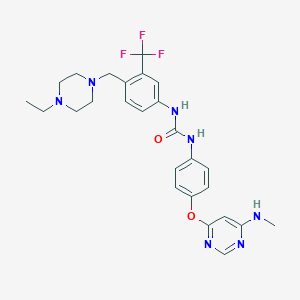 AST 487 AST 487 | C26H30F3N7O2 | 529.568 | 10 / 3 | 3.8 | No |
| 240973 | 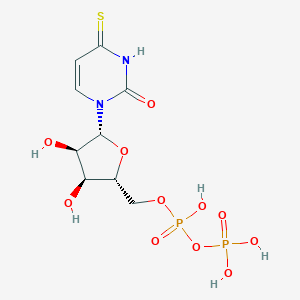 4-Thio-UDP 4-Thio-UDP | C9H14N2O11P2S | 420.222 | 12 / 6 | -4.1 | No |
| 245473 | 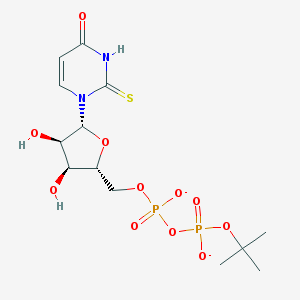 BDBM50303347 BDBM50303347 | C13H20N2O11P2S-2 | 474.314 | 12 / 3 | -2.8 | No |
| 248594 | 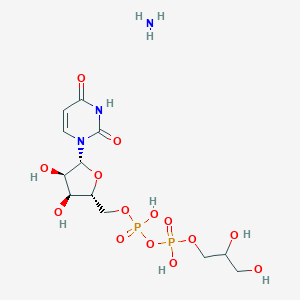 CHEMBL1162560 CHEMBL1162560 | C12H23N3O14P2 | 495.271 | 15 / 8 | N/A | No |
| 252318 | 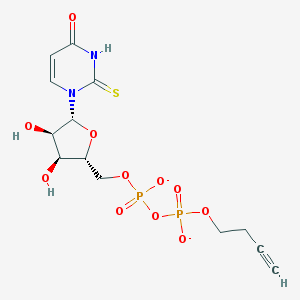 BDBM50303333 BDBM50303333 | C13H16N2O11P2S-2 | 470.282 | 12 / 3 | -3.2 | No |
| 256854 | 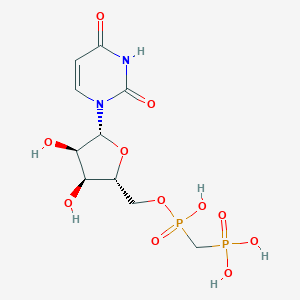 CHEMBL507060 CHEMBL507060 | C10H16N2O11P2 | 402.189 | 11 / 6 | -4.9 | No |
| 260832 | 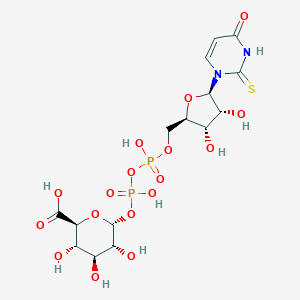 CHEMBL387520 CHEMBL387520 | C15H22N2O17P2S | 596.346 | 18 / 9 | -5.8 | No |
| 267062 | 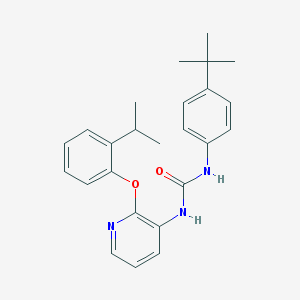 CHEMBL2333772 CHEMBL2333772 | C25H29N3O2 | 403.526 | 3 / 2 | 6.1 | No |
| 267371 | 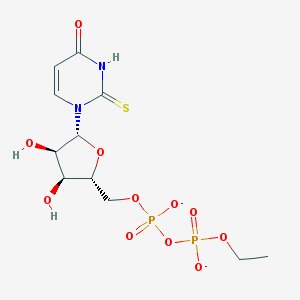 BDBM50303343 BDBM50303343 | C11H16N2O11P2S-2 | 446.26 | 12 / 3 | -3.4 | No |
| 276597 | 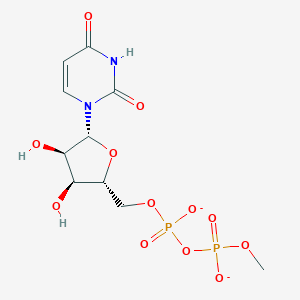 P1-uridyl-P2-methyl diphosphate P1-uridyl-P2-methyl diphosphate | C10H14N2O12P2-2 | 416.172 | 12 / 3 | -4.3 | No |
| 285698 | 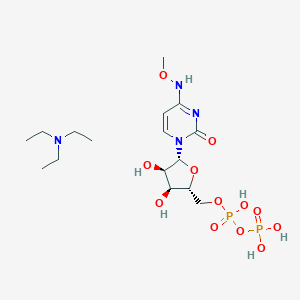 CHEMBL1084020 CHEMBL1084020 | C16H32N4O12P2 | 534.396 | 13 / 6 | N/A | No |
| 554725 | 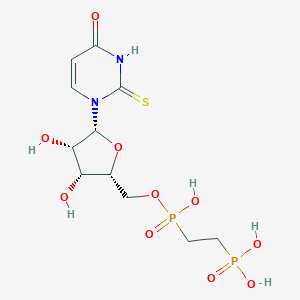 MRS2905 MRS2905 | C11H18N2O10P2S | 432.277 | 11 / 6 | -4.4 | No |
| 293168 | 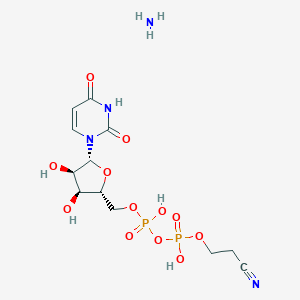 CHEMBL1162555 CHEMBL1162555 | C12H20N4O12P2 | 474.256 | 14 / 6 | N/A | No |
| 303242 | 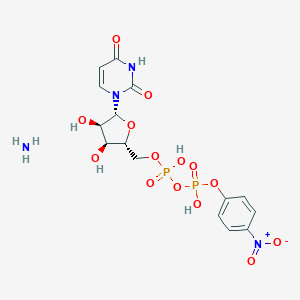 CHEMBL612119 CHEMBL612119 | C15H20N4O14P2 | 542.287 | 15 / 6 | N/A | No |
| 316932 | 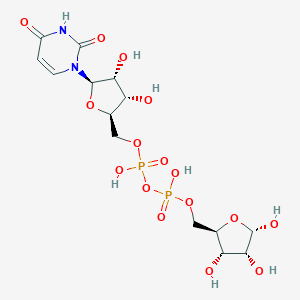 CHEMBL595467 CHEMBL595467 | C14H22N2O16P2 | 536.276 | 16 / 8 | -6.2 | No |
| 316933 | 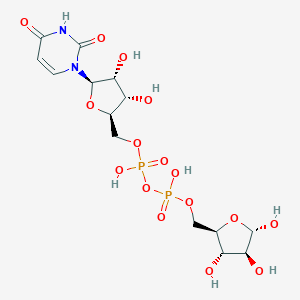 CHEMBL593842 CHEMBL593842 | C14H22N2O16P2 | 536.276 | 16 / 8 | -6.2 | No |
| 320744 | 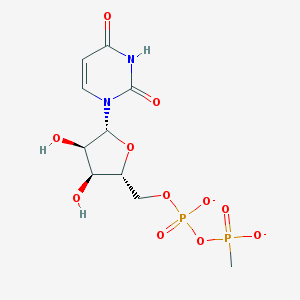 BDBM50303351 BDBM50303351 | C10H14N2O11P2-2 | 400.173 | 11 / 3 | -4.4 | No |
| 554923 | 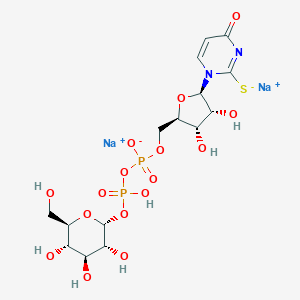 MRS 2690 MRS 2690 | C15H22N2Na2O16P2S | 626.326 | 17 / 7 | N/A | No |
| 344144 | 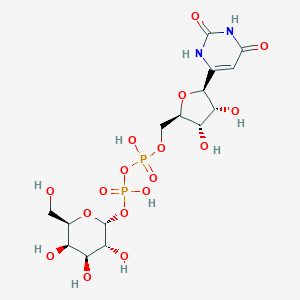 CHEMBL1743884 CHEMBL1743884 | C15H24N2O17P2 | 566.302 | 17 / 10 | -7.1 | No |
| 349200 | 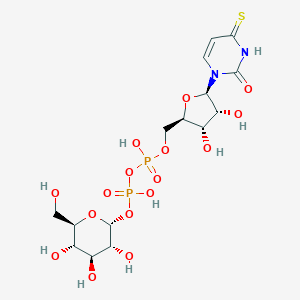 CHEMBL374384 CHEMBL374384 | C15H24N2O16P2S | 582.363 | 17 / 9 | -6.0 | No |
| 349829 | 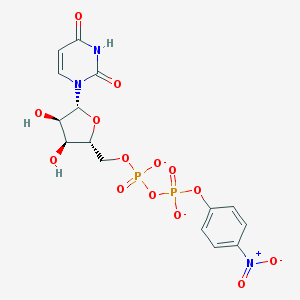 BDBM50303349 BDBM50303349 | C15H15N3O14P2-2 | 523.24 | 14 / 3 | -2.7 | No |
| 350577 | 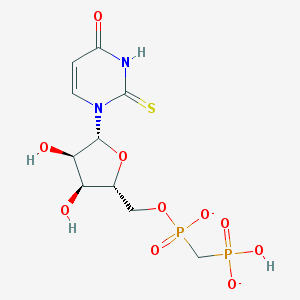 BDBM50303338 BDBM50303338 | C10H14N2O10P2S-2 | 416.234 | 11 / 4 | -4.4 | No |
| 368335 | 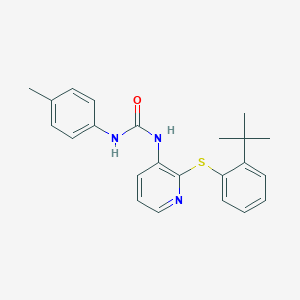 CHEMBL2333767 CHEMBL2333767 | C23H25N3OS | 391.533 | 3 / 2 | 5.9 | No |
| 370725 | 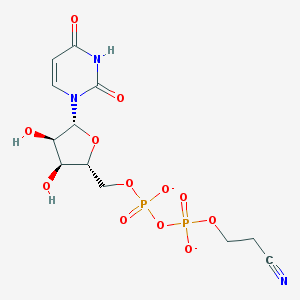 BDBM50303332 BDBM50303332 | C12H15N3O12P2-2 | 455.209 | 13 / 3 | -4.7 | No |
| 462424 | 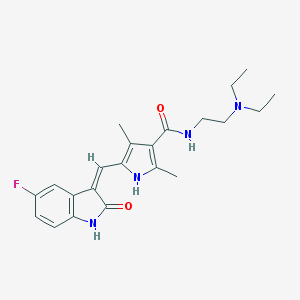 Sunitinib Sunitinib | C22H27FN4O2 | 398.482 | 4 / 3 | 2.6 | Yes |
| 375501 | 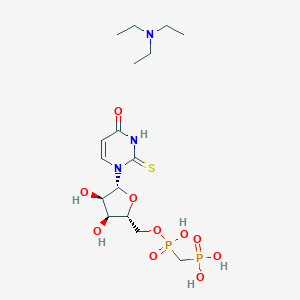 CHEMBL611791 CHEMBL611791 | C16H31N3O10P2S | 519.443 | 12 / 6 | N/A | No |
| 377569 | 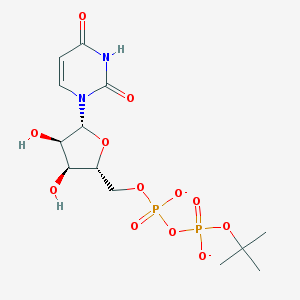 BDBM50303346 BDBM50303346 | C13H20N2O12P2-2 | 458.253 | 12 / 3 | -3.4 | No |
| 381921 | 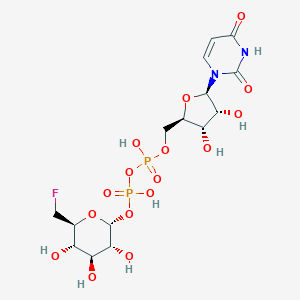 CHEMBL593126 CHEMBL593126 | C15H23FN2O16P2 | 568.293 | 17 / 8 | -6.2 | No |
| 382531 | 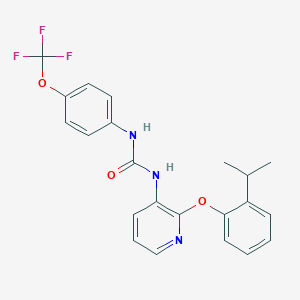 CHEMBL2333773 CHEMBL2333773 | C22H20F3N3O3 | 431.415 | 7 / 2 | 5.6 | No |
| 388143 | 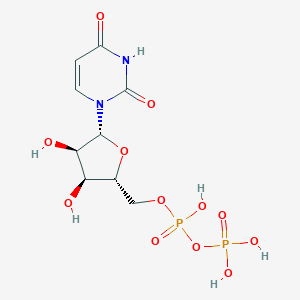 Uridine 5'-diphosphate Uridine 5'-diphosphate | C9H14N2O12P2 | 404.161 | 12 / 6 | -4.7 | No |
| 392887 | 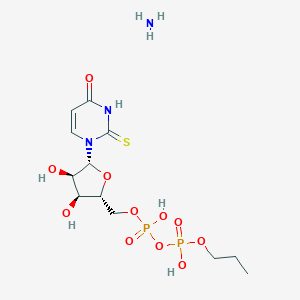 CHEMBL1162553 CHEMBL1162553 | C12H23N3O11P2S | 479.334 | 13 / 6 | N/A | No |
| 402627 | 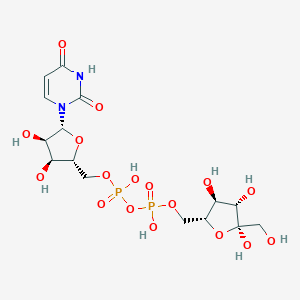 CHEMBL604676 CHEMBL604676 | C15H24N2O17P2 | 566.302 | 17 / 9 | -6.9 | No |
| 421417 | 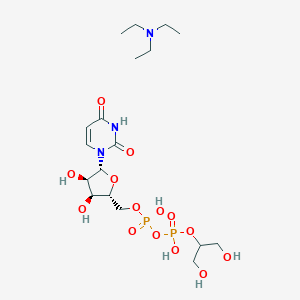 CHEMBL612065 CHEMBL612065 | C18H35N3O14P2 | 579.433 | 15 / 7 | N/A | No |
| 428561 | 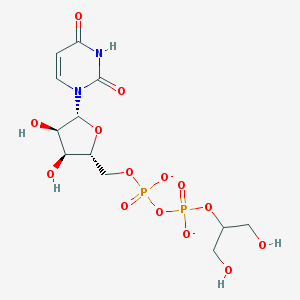 BDBM50303335 BDBM50303335 | C12H18N2O14P2-2 | 476.224 | 14 / 5 | -5.7 | No |
| 428882 | 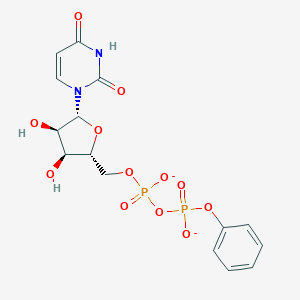 P1-uridyl-P2-phenyl diphosphate P1-uridyl-P2-phenyl diphosphate | C15H16N2O12P2-2 | 478.243 | 12 / 3 | -2.5 | No |
| 430401 | 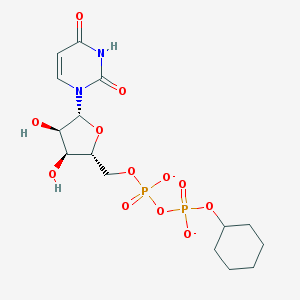 BDBM50303336 BDBM50303336 | C15H22N2O12P2-2 | 484.291 | 12 / 3 | -2.5 | No |
| 515957 | 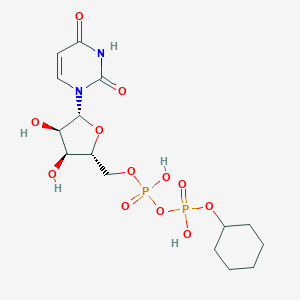 CHEMBL1199733 CHEMBL1199733 | C15H24N2O12P2 | 486.307 | 12 / 5 | -2.4 | No |
| 431448 | 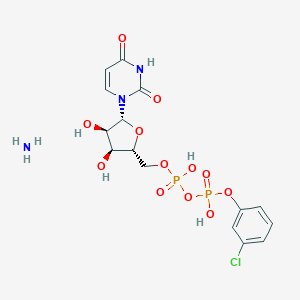 CHEMBL1162561 CHEMBL1162561 | C15H20ClN3O12P2 | 531.732 | 13 / 6 | N/A | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218