

You can:
| Name | Leukotriene B4 receptor 1 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | LTB4R |
| Synonym | BLT1 receptor BLTR Chemoattractant receptor-like 1 G-protein coupled receptor 16 GPR16 [ Show all ] |
| Disease | Inflammatory disease Inflammatory bowel disease Human immunodeficiency virus infection Pancreatic cancer Psoriasis [ Show all ] |
| Length | 352 |
| Amino acid sequence | MNTTSSAAPPSLGVEFISLLAIILLSVALAVGLPGNSFVVWSILKRMQKRSVTALMVLNLALADLAVLLTAPFFLHFLAQGTWSFGLAGCRLCHYVCGVSMYASVLLITAMSLDRSLAVARPFVSQKLRTKAMARRVLAGIWVLSFLLATPVLAYRTVVPWKTNMSLCFPRYPSEGHRAFHLIFEAVTGFLLPFLAVVASYSDIGRRLQARRFRRSRRTGRLVVLIILTFAAFWLPYHVVNLAEAGRALAGQAAGLGLVGKRLSLARNVLIALAFLSSSVNPVLYACAGGGLLRSAGVGFVAKLLEGTGSEASSTRRGGSLGQTARSGPAALEPGPSESLTASSPLKLNELN |
| UniProt | Q15722 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q15722 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q15722. |
| BioLiP | N/A |
| Therapeutic Target Database | T59626 |
| ChEMBL | CHEMBL3911 |
| IUPHAR | 267 |
| DrugBank | BE0003490 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 251 | 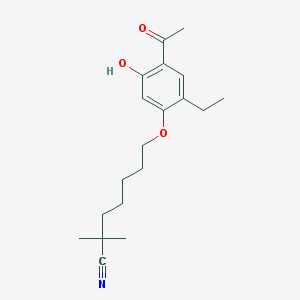 CHEMBL299404 CHEMBL299404 | C19H27NO3 | 317.429 | 4 / 1 | 4.8 | Yes |
| 1643 | 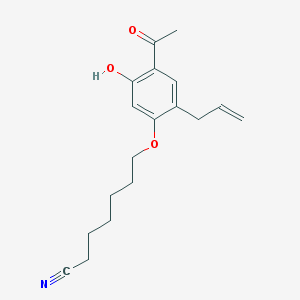 CHEMBL298953 CHEMBL298953 | C18H23NO3 | 301.386 | 4 / 1 | 4.1 | Yes |
| 1771 | 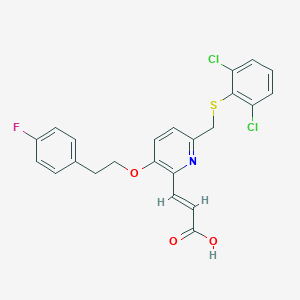 CHEMBL123722 CHEMBL123722 | C23H18Cl2FNO3S | 478.359 | 6 / 1 | 6.2 | No |
| 2197 | 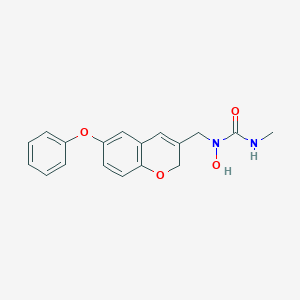 CHEMBL118929 CHEMBL118929 | C18H18N2O4 | 326.352 | 4 / 2 | 2.0 | Yes |
| 3209 | 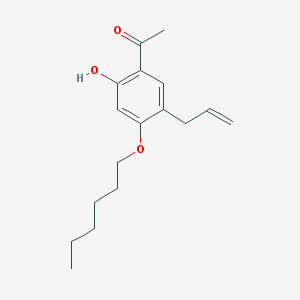 117690-47-8 117690-47-8 | C17H24O3 | 276.376 | 3 / 1 | 5.2 | No |
| 3818 | 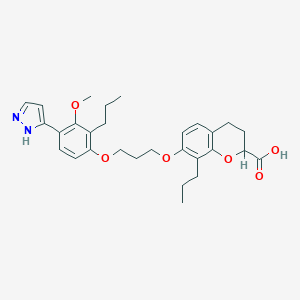 CHEMBL169683 CHEMBL169683 | C29H36N2O6 | 508.615 | 7 / 2 | 6.5 | No |
| 3920 | 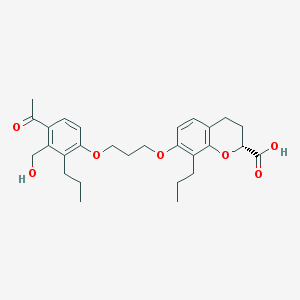 CHEMBL262530 CHEMBL262530 | C28H36O7 | 484.589 | 7 / 2 | 5.4 | No |
| 4887 | 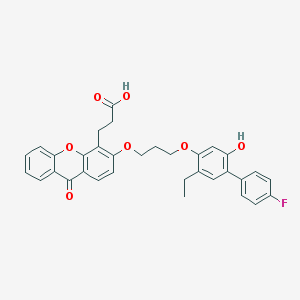 CHEMBL129463 CHEMBL129463 | C33H29FO7 | 556.586 | 8 / 2 | 6.9 | No |
| 5169 | 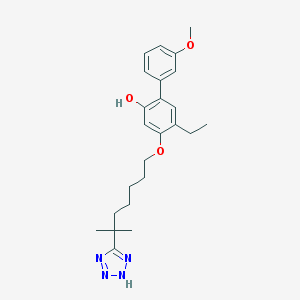 CHEMBL167923 CHEMBL167923 | C24H32N4O3 | 424.545 | 6 / 2 | 6.0 | No |
| 6183 | 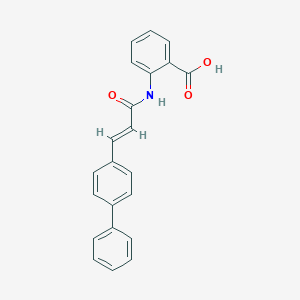 CHEMBL176974 CHEMBL176974 | C22H17NO3 | 343.382 | 3 / 2 | 5.6 | No |
| 555507 | 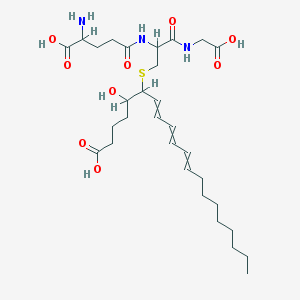 Leukotriene C3 Leukotriene C3 | C30H49N3O9S | 627.794 | 11 / 7 | 1.3 | No |
| 8985 | 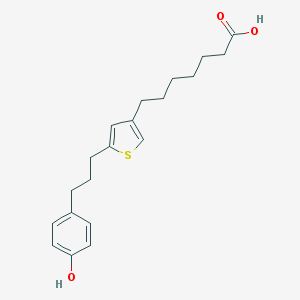 CHEMBL107781 CHEMBL107781 | C20H26O3S | 346.485 | 4 / 2 | 5.6 | No |
| 11040 | 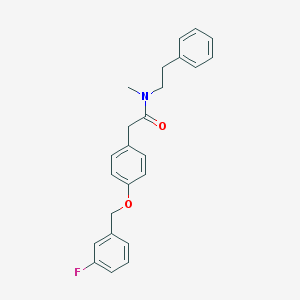 CHEMBL124685 CHEMBL124685 | C24H24FNO2 | 377.459 | 3 / 0 | 4.8 | Yes |
| 14291 | 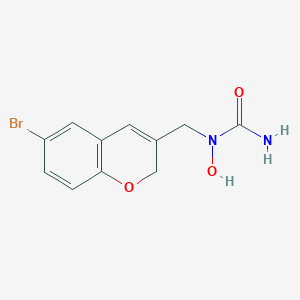 CHEMBL331748 CHEMBL331748 | C11H11BrN2O3 | 299.124 | 3 / 2 | 0.7 | Yes |
| 14443 | 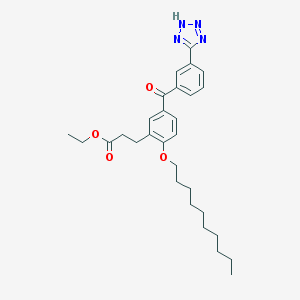 CHEMBL94442 CHEMBL94442 | C29H38N4O4 | 506.647 | 7 / 1 | 7.4 | No |
| 16587 | 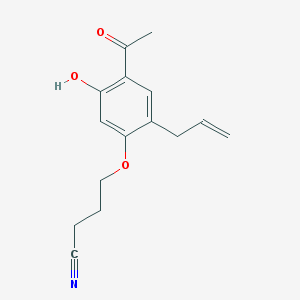 CHEMBL51908 CHEMBL51908 | C15H17NO3 | 259.305 | 4 / 1 | 2.9 | Yes |
| 17694 | 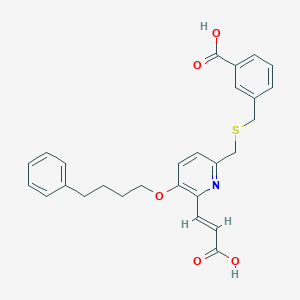 CHEMBL115658 CHEMBL115658 | C27H27NO5S | 477.575 | 7 / 2 | 5.0 | Yes |
| 18073 | 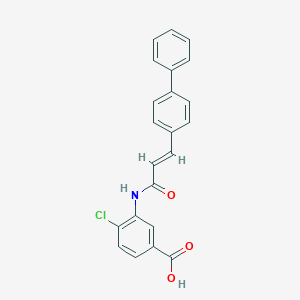 CHEMBL176903 CHEMBL176903 | C22H16ClNO3 | 377.824 | 3 / 2 | 5.0 | Yes |
| 18294 | 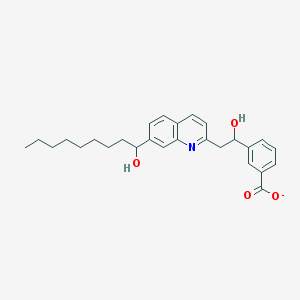 CHEMBL326397 CHEMBL326397 | C27H32NO4- | 434.556 | 5 / 2 | 6.6 | No |
| 18553 | 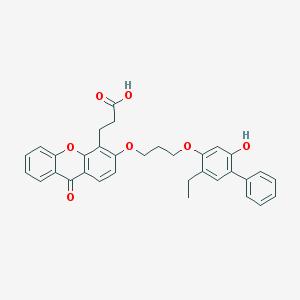 CHEMBL129272 CHEMBL129272 | C33H30O7 | 538.596 | 7 / 2 | 6.8 | No |
| 18920 | 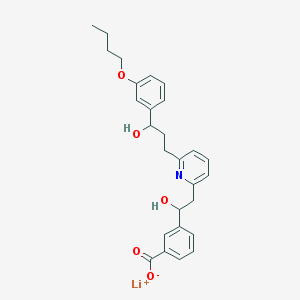 CHEMBL114454 CHEMBL114454 | C27H30LiNO5 | 455.479 | 6 / 2 | N/A | N/A |
| 19221 | 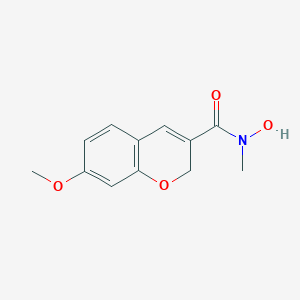 CHEMBL118377 CHEMBL118377 | C12H13NO4 | 235.239 | 4 / 1 | 0.9 | Yes |
| 21701 | 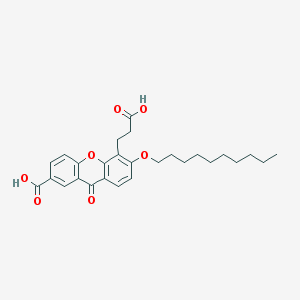 CHEMBL288758 CHEMBL288758 | C27H32O7 | 468.546 | 7 / 2 | 6.7 | No |
| 22574 | 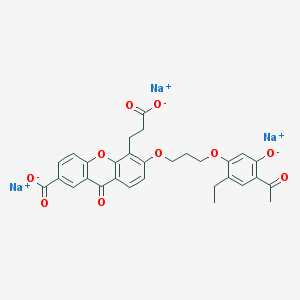 CHEMBL292768 CHEMBL292768 | C30H25Na3O10 | 614.489 | 10 / 0 | N/A | No |
| 22891 | 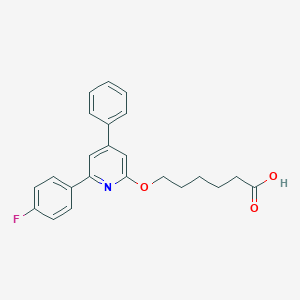 CHEMBL137801 CHEMBL137801 | C23H22FNO3 | 379.431 | 5 / 1 | 5.1 | No |
| 24214 | 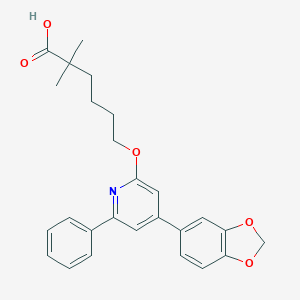 CHEMBL137330 CHEMBL137330 | C26H27NO5 | 433.504 | 6 / 1 | 5.8 | No |
| 24579 | 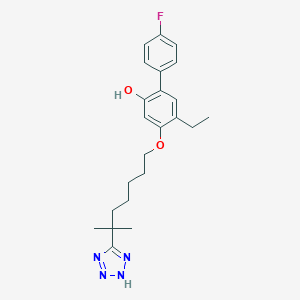 CHEMBL167520 CHEMBL167520 | C23H29FN4O2 | 412.509 | 6 / 2 | 6.1 | No |
| 24690 | 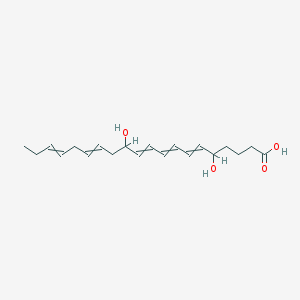 5,12-dihydroxyicosa-6,8,10,14,17-pentaenoic acid 5,12-dihydroxyicosa-6,8,10,14,17-pentaenoic acid | C20H30O4 | 334.456 | 4 / 3 | 3.4 | Yes |
| 25843 | 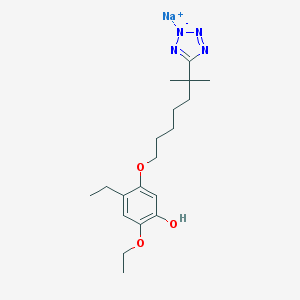 CHEMBL48667 CHEMBL48667 | C19H29N4NaO3 | 384.456 | 7 / 1 | N/A | N/A |
| 26660 | 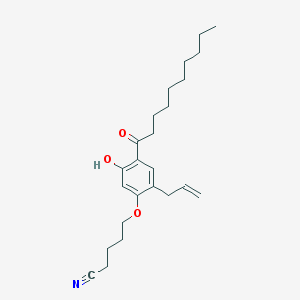 CHEMBL54873 CHEMBL54873 | C24H35NO3 | 385.548 | 4 / 1 | 7.3 | No |
| 26712 | 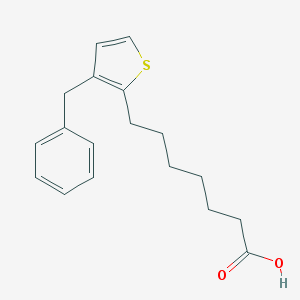 2-Thiopheneheptanoic acid, 3-(phenylmethyl)- 2-Thiopheneheptanoic acid, 3-(phenylmethyl)- | C18H22O2S | 302.432 | 3 / 1 | 5.1 | No |
| 27729 | 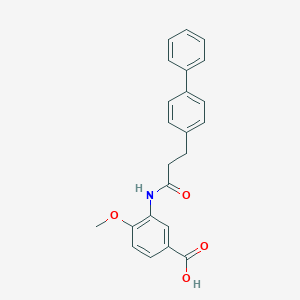 CHEMBL172091 CHEMBL172091 | C23H21NO4 | 375.424 | 4 / 2 | 4.1 | Yes |
| 31882 | 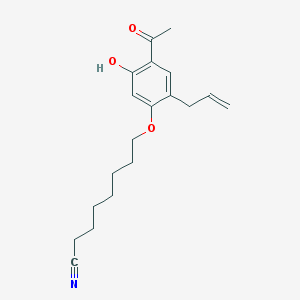 CHEMBL53774 CHEMBL53774 | C19H25NO3 | 315.413 | 4 / 1 | 4.7 | Yes |
| 34067 | 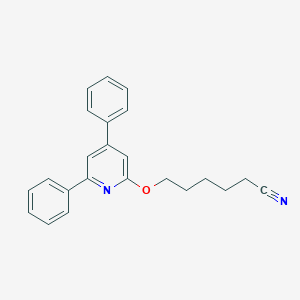 CHEMBL137307 CHEMBL137307 | C23H22N2O | 342.442 | 3 / 0 | 5.2 | No |
| 34706 | 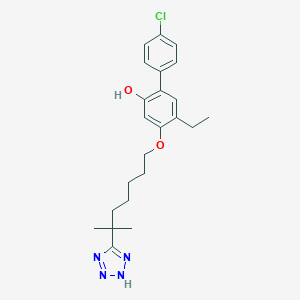 CHEMBL167850 CHEMBL167850 | C23H29ClN4O2 | 428.961 | 5 / 2 | 6.6 | No |
| 34844 | 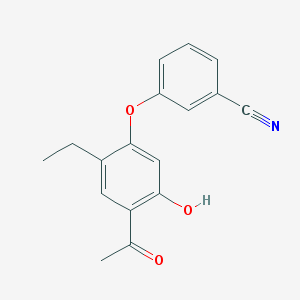 3-(4-Acetyl-2-ethyl-5-hydroxyphenoxy)benzonitrile 3-(4-Acetyl-2-ethyl-5-hydroxyphenoxy)benzonitrile | C17H15NO3 | 281.311 | 4 / 1 | 3.8 | Yes |
| 34914 | 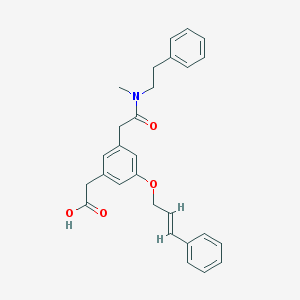 CHEMBL125447 CHEMBL125447 | C28H29NO4 | 443.543 | 4 / 1 | 4.8 | Yes |
| 35172 | 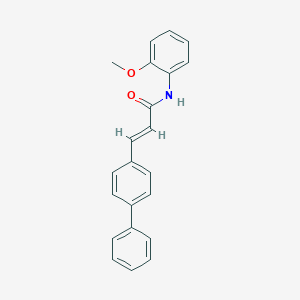 CHEMBL171853 CHEMBL171853 | C22H19NO2 | 329.399 | 2 / 1 | 4.8 | Yes |
| 35689 | 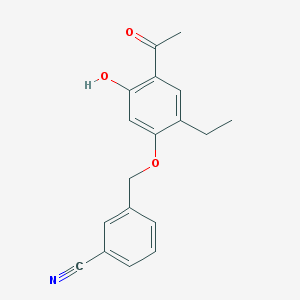 CHEMBL53934 CHEMBL53934 | C18H17NO3 | 295.338 | 4 / 1 | 3.8 | Yes |
| 37568 | 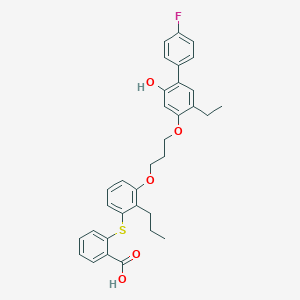 CHEMBL139254 CHEMBL139254 | C33H33FO5S | 560.68 | 7 / 2 | 9.0 | No |
| 38390 | 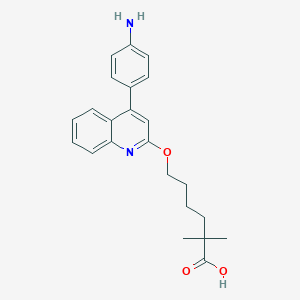 CHEMBL134697 CHEMBL134697 | C23H26N2O3 | 378.472 | 5 / 2 | 4.9 | Yes |
| 39323 | 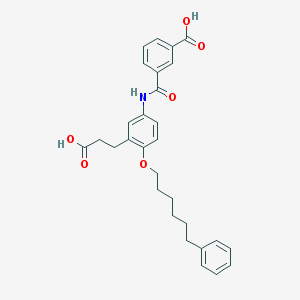 CHEMBL93764 CHEMBL93764 | C29H31NO6 | 489.568 | 6 / 3 | 5.8 | No |
| 39619 | 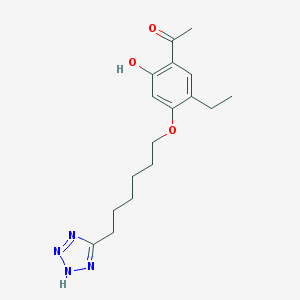 CHEMBL52021 CHEMBL52021 | C17H24N4O3 | 332.404 | 6 / 2 | 3.7 | Yes |
| 39703 | 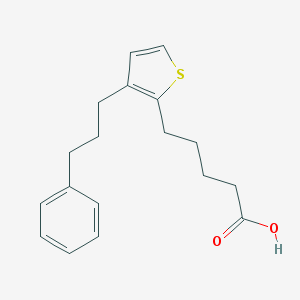 CHEMBL108094 CHEMBL108094 | C18H22O2S | 302.432 | 3 / 1 | 4.9 | Yes |
| 41888 | 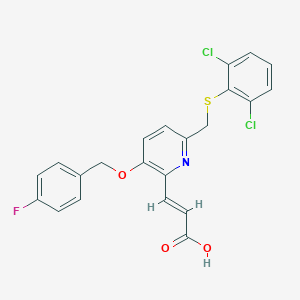 CHEMBL127341 CHEMBL127341 | C22H16Cl2FNO3S | 464.332 | 6 / 1 | 5.7 | No |
| 42301 | 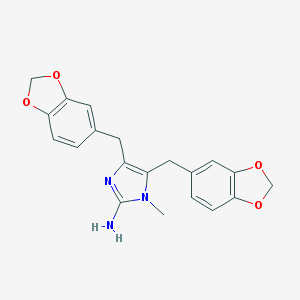 Leucettamine A Leucettamine A | C20H19N3O4 | 365.389 | 6 / 1 | 3.2 | Yes |
| 42729 | 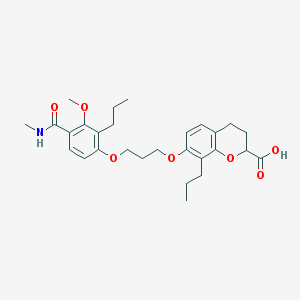 SC-50073 SC-50073 | C28H37NO7 | 499.604 | 7 / 2 | 5.8 | No |
| 43043 | 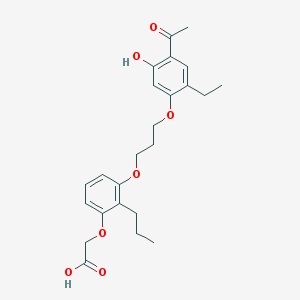 CHEMBL15648 CHEMBL15648 | C24H30O7 | 430.497 | 7 / 2 | 5.5 | No |
| 43378 | 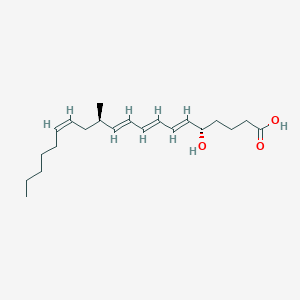 CHEMBL53967 CHEMBL53967 | C21H34O3 | 334.5 | 3 / 2 | 5.6 | No |
| 44631 | 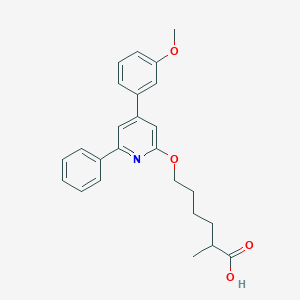 CHEMBL137336 CHEMBL137336 | C25H27NO4 | 405.494 | 5 / 1 | 5.6 | No |
| 45508 | 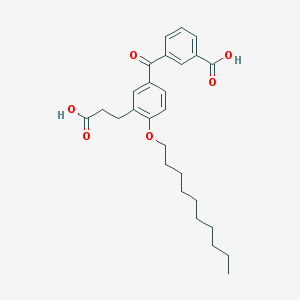 LY 213024 LY 213024 | C27H34O6 | 454.563 | 6 / 2 | 7.0 | No |
| 45929 | 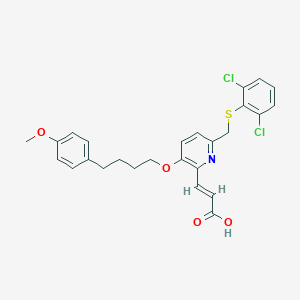 CHEMBL337995 CHEMBL337995 | C26H25Cl2NO4S | 518.449 | 6 / 1 | 6.7 | No |
| 46927 | 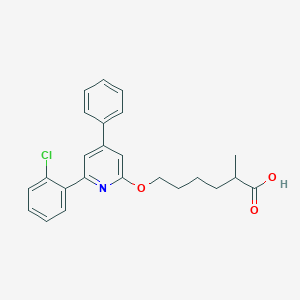 CHEMBL342416 CHEMBL342416 | C24H24ClNO3 | 409.91 | 4 / 1 | 6.2 | No |
| 47692 | 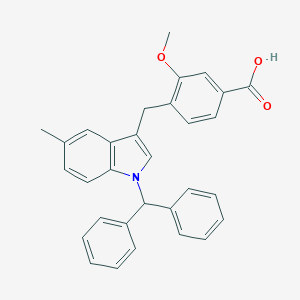 CHEMBL337790 CHEMBL337790 | C31H27NO3 | 461.561 | 3 / 1 | 7.1 | No |
| 48517 | 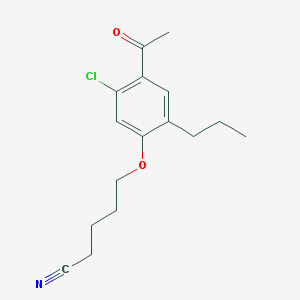 CHEMBL53326 CHEMBL53326 | C16H20ClNO2 | 293.791 | 3 / 0 | 3.9 | Yes |
| 49148 | 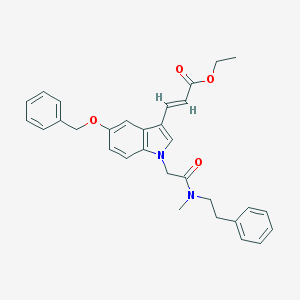 CHEMBL332422 CHEMBL332422 | C31H32N2O4 | 496.607 | 4 / 0 | 5.7 | No |
| 49183 | 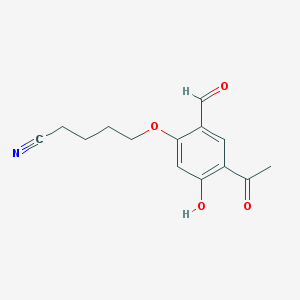 CHEMBL53013 CHEMBL53013 | C14H15NO4 | 261.277 | 5 / 1 | 1.6 | Yes |
| 49649 | 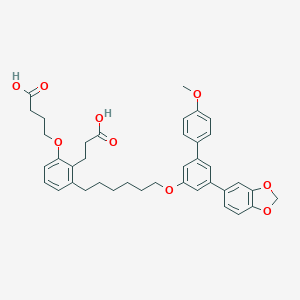 CHEMBL1099338 CHEMBL1099338 | C39H42O9 | 654.756 | 9 / 2 | 8.3 | No |
| 50443 | 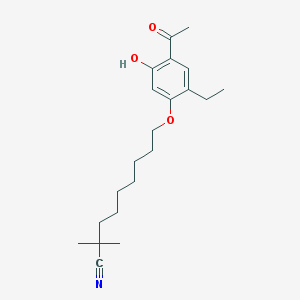 CHEMBL52595 CHEMBL52595 | C21H31NO3 | 345.483 | 4 / 1 | 5.9 | No |
| 52193 | 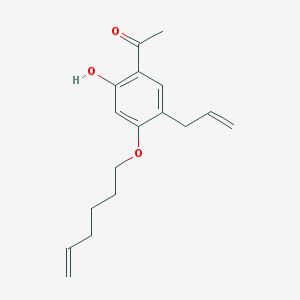 CHEMBL51042 CHEMBL51042 | C17H22O3 | 274.36 | 3 / 1 | 4.7 | Yes |
| 52233 | 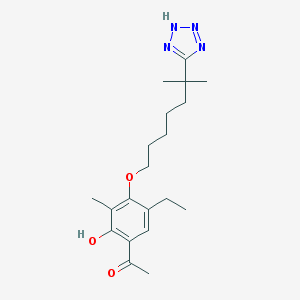 CHEMBL53059 CHEMBL53059 | C20H30N4O3 | 374.485 | 6 / 2 | 5.0 | Yes |
| 52682 | 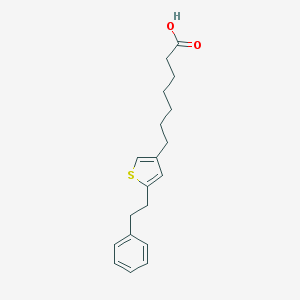 CHEMBL106055 CHEMBL106055 | C19H24O2S | 316.459 | 3 / 1 | 5.6 | No |
| 54199 | 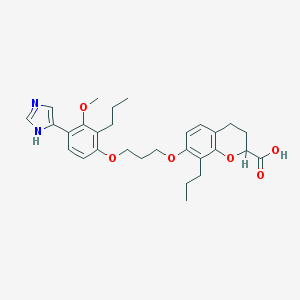 CHEMBL352863 CHEMBL352863 | C29H36N2O6 | 508.615 | 7 / 2 | 6.4 | No |
| 55197 | 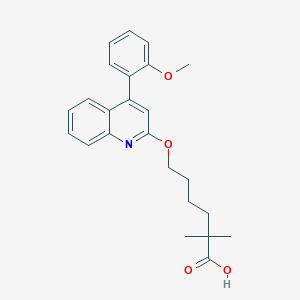 CHEMBL137203 CHEMBL137203 | C24H27NO4 | 393.483 | 5 / 1 | 5.6 | No |
| 55337 | 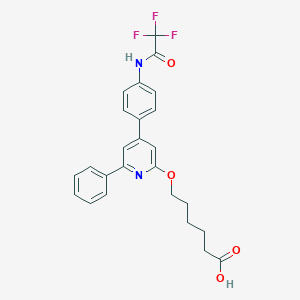 CHEMBL336303 CHEMBL336303 | C25H23F3N2O4 | 472.464 | 8 / 2 | 5.3 | No |
| 55787 | 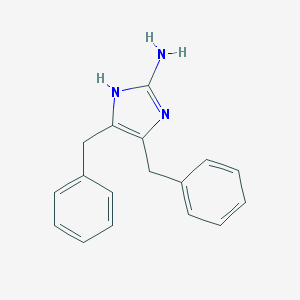 CHEMBL116278 CHEMBL116278 | C17H17N3 | 263.344 | 2 / 2 | 3.6 | Yes |
| 55798 | 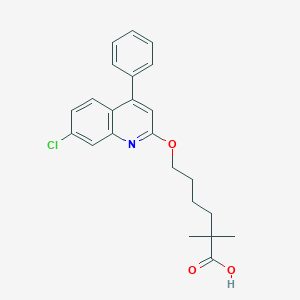 CHEMBL335142 CHEMBL335142 | C23H24ClNO3 | 397.899 | 4 / 1 | 6.2 | No |
| 55992 | 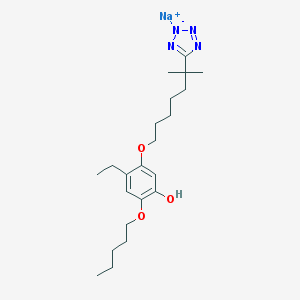 CHEMBL416259 CHEMBL416259 | C22H35N4NaO3 | 426.537 | 7 / 1 | N/A | N/A |
| 56718 | 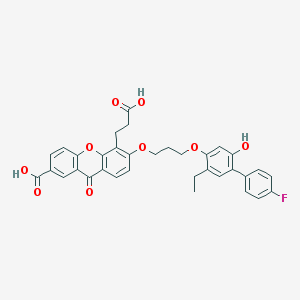 LY-292728 LY-292728 | C34H29FO9 | 600.595 | 10 / 3 | 6.4 | No |
| 57122 | 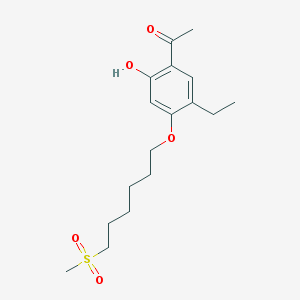 CHEMBL52425 CHEMBL52425 | C17H26O5S | 342.45 | 5 / 1 | 3.4 | Yes |
| 58342 | 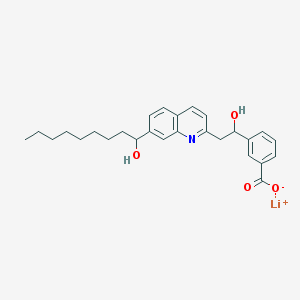 CHEMBL326397 CHEMBL326397 | C27H32LiNO4 | 441.496 | 5 / 2 | N/A | N/A |
| 58982 | 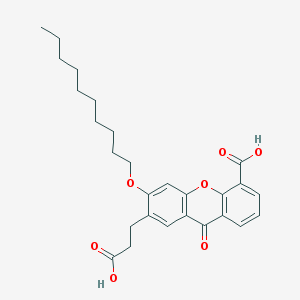 CHEMBL297501 CHEMBL297501 | C27H32O7 | 468.546 | 7 / 2 | 6.7 | No |
| 59318 | 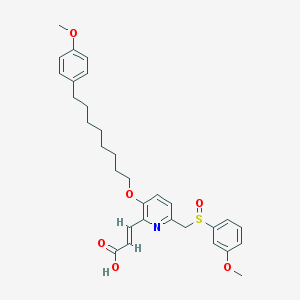 CHEMBL113167 CHEMBL113167 | C31H37NO6S | 551.698 | 8 / 1 | 6.3 | No |
| 59776 | 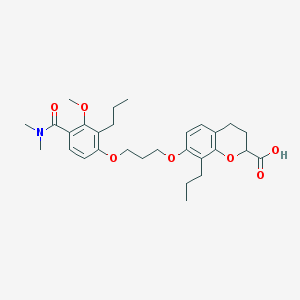 SC-52569 SC-52569 | C29H39NO7 | 513.631 | 7 / 1 | 6.0 | No |
| 60724 | 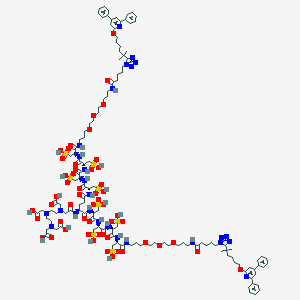 DTPA Conjugate DTPA Conjugate | C123H180N26O53S8 | 3127.4 | 64 / 25 | -11.7 | No |
| 62171 | 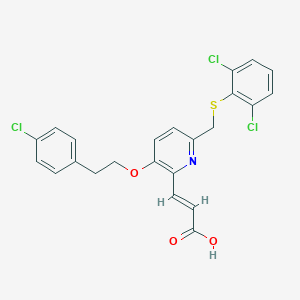 CHEMBL127376 CHEMBL127376 | C23H18Cl3NO3S | 494.811 | 5 / 1 | 6.7 | No |
| 444164 | 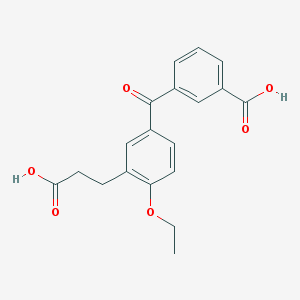 CHEMBL92771 CHEMBL92771 | C19H18O6 | 342.347 | 6 / 2 | 2.9 | Yes |
| 62612 | 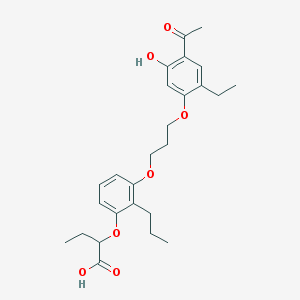 CHEMBL15295 CHEMBL15295 | C26H34O7 | 458.551 | 7 / 2 | 6.4 | No |
| 64238 | 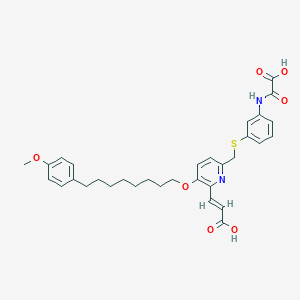 CHEMBL334267 CHEMBL334267 | C32H36N2O7S | 592.707 | 9 / 3 | 6.8 | No |
| 64864 | 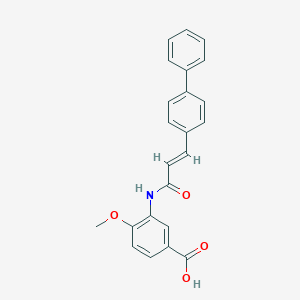 CHEMBL367943 CHEMBL367943 | C23H19NO4 | 373.408 | 4 / 2 | 4.3 | Yes |
| 68307 | 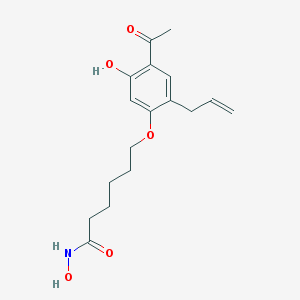 CHEMBL298512 CHEMBL298512 | C17H23NO5 | 321.373 | 5 / 3 | 2.7 | Yes |
| 559459 | 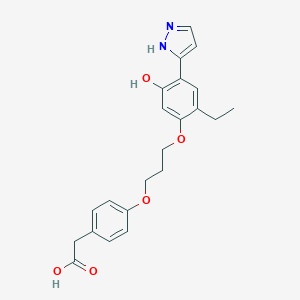 CHEMBL81817 CHEMBL81817 | C22H24N2O5 | 396.443 | 6 / 3 | 3.8 | Yes |
| 69132 | 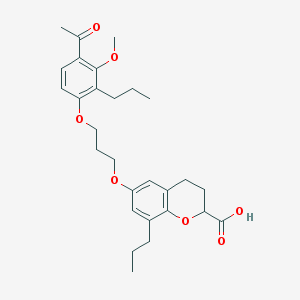 CHEMBL15817 CHEMBL15817 | C28H36O7 | 484.589 | 7 / 1 | 6.2 | No |
| 69800 | 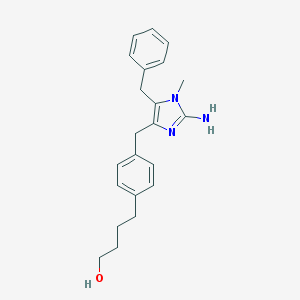 CHEMBL112309 CHEMBL112309 | C22H27N3O | 349.478 | 3 / 2 | 3.9 | Yes |
| 70482 | 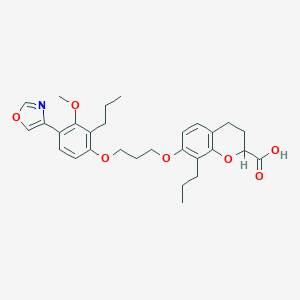 CHEMBL168658 CHEMBL168658 | C29H35NO7 | 509.599 | 8 / 1 | 6.6 | No |
| 70657 | 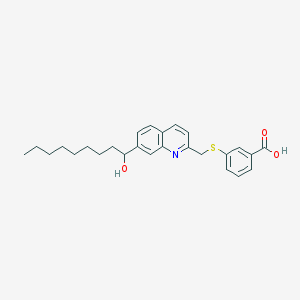 CHEMBL113792 CHEMBL113792 | C26H31NO3S | 437.598 | 5 / 2 | 6.9 | No |
| 71391 | 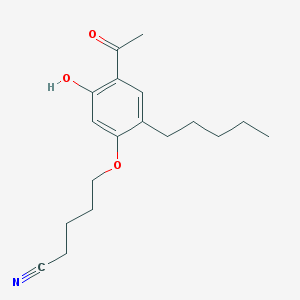 CHEMBL52630 CHEMBL52630 | C18H25NO3 | 303.402 | 4 / 1 | 4.6 | Yes |
| 72848 | 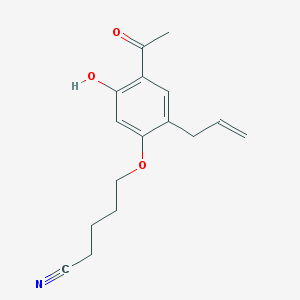 5-(4-acetyl-5-hydroxy-2-allylphenoxy)pentanenitrile 5-(4-acetyl-5-hydroxy-2-allylphenoxy)pentanenitrile | C16H19NO3 | 273.332 | 4 / 1 | 3.2 | Yes |
| 74111 | 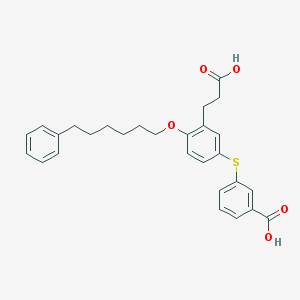 CHEMBL316674 CHEMBL316674 | C28H30O5S | 478.603 | 6 / 2 | 7.0 | No |
| 74547 | 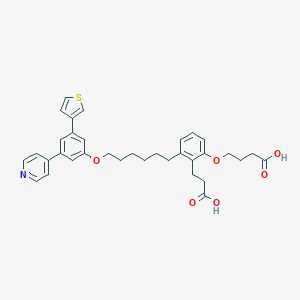 CHEMBL1099332 CHEMBL1099332 | C34H37NO6S | 587.731 | 8 / 2 | 7.1 | No |
| 74768 | 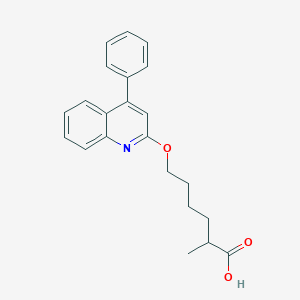 CHEMBL137570 CHEMBL137570 | C22H23NO3 | 349.43 | 4 / 1 | 5.3 | No |
| 75563 | 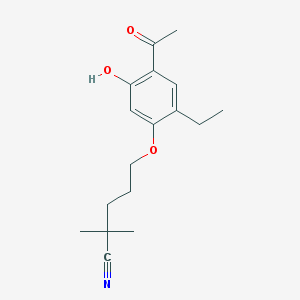 CHEMBL52352 CHEMBL52352 | C17H23NO3 | 289.375 | 4 / 1 | 3.9 | Yes |
| 76111 | 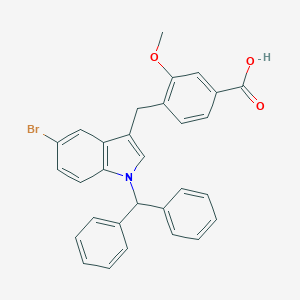 CHEMBL140539 CHEMBL140539 | C30H24BrNO3 | 526.43 | 3 / 1 | 7.4 | No |
| 76523 | 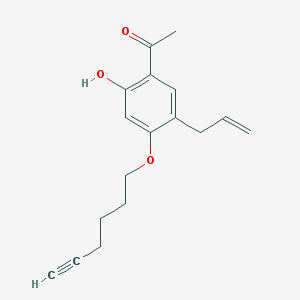 CHEMBL51906 CHEMBL51906 | C17H20O3 | 272.344 | 3 / 1 | 4.1 | Yes |
| 76854 | 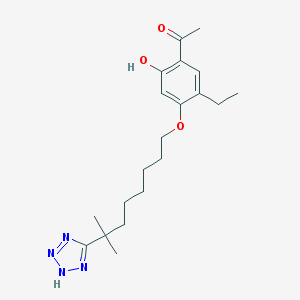 CHEMBL51558 CHEMBL51558 | C20H30N4O3 | 374.485 | 6 / 2 | 5.1 | No |
| 79004 | 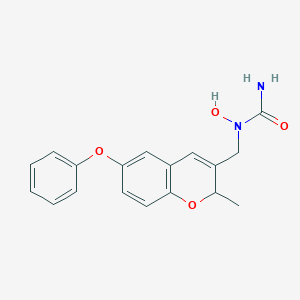 CHEMBL332092 CHEMBL332092 | C18H18N2O4 | 326.352 | 4 / 2 | 2.0 | Yes |
| 79365 | 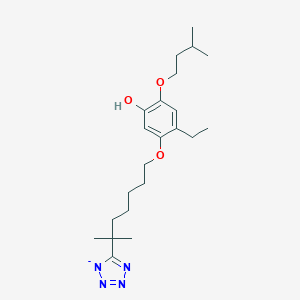 CHEMBL295799 CHEMBL295799 | C22H35N4O3- | 403.547 | 7 / 1 | 5.4 | No |
| 79858 | 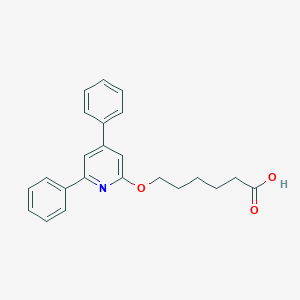 CHEMBL137504 CHEMBL137504 | C23H23NO3 | 361.441 | 4 / 1 | 5.0 | Yes |
| 80257 | 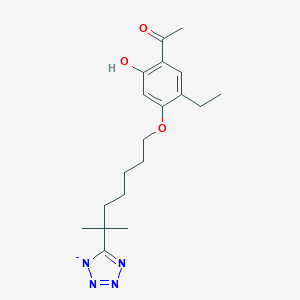 CHEMBL45460 CHEMBL45460 | C19H27N4O3- | 359.45 | 7 / 1 | 4.0 | Yes |
| 80616 | 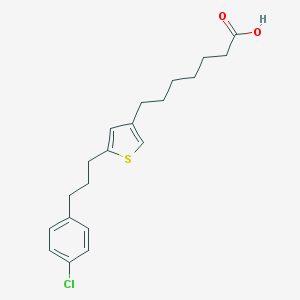 CHEMBL109015 CHEMBL109015 | C20H25ClO2S | 364.928 | 3 / 1 | 6.6 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218