

You can:
| Name | Calcitonin gene-related peptide type 1 receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | CALCRL |
| Synonym | CRLR CLR (unofficial abbreviation in common use) CGRP type 1 receptor Calcitonin receptor-like receptor |
| Disease | Migraine; Cluster headaches Migraine |
| Length | 461 |
| Amino acid sequence | MEKKCTLNFLVLLPFFMILVTAELEESPEDSIQLGVTRNKIMTAQYECYQKIMQDPIQQAEGVYCNRTWDGWLCWNDVAAGTESMQLCPDYFQDFDPSEKVTKICDQDGNWFRHPASNRTWTNYTQCNVNTHEKVKTALNLFYLTIIGHGLSIASLLISLGIFFYFKSLSCQRITLHKNLFFSFVCNSVVTIIHLTAVANNQALVATNPVSCKVSQFIHLYLMGCNYFWMLCEGIYLHTLIVVAVFAEKQHLMWYYFLGWGFPLIPACIHAIARSLYYNDNCWISSDTHLLYIIHGPICAALLVNLFFLLNIVRVLITKLKVTHQAESNLYMKAVRATLILVPLLGIEFVLIPWRPEGKIAEEVYDYIMHILMHFQGLLVSTIFCFFNGEVQAILRRNWNQYKIQFGNSFSNSEALRSASYTVSTISDGPGYSHDCPSEHLNGKSIHDIENVLLKPENLYN |
| UniProt | Q16602 |
| Protein Data Bank | 3n7r, 6e3y, 6d1u, 5v6y, 4rwg, 4rwf, 3n7s |
| GPCR-HGmod model | Q16602 |
| 3D structure model | This structure is from PDB ID 3n7r. |
| BioLiP | BL0314647, BL0314642, BL0314643,BL0314645,BL0314648, BL0314644,BL0314646,BL0314649, BL0403127,BL0403130,BL0403133,, BL0403128,BL0403131,BL0403134,, BL0403129,BL0403132,BL0403135,, BL0425957,BL0425959,BL0425961, BL0425958,BL0425960,BL0425962, BL0427207,BL0427208, BL0314641, BL0183084, BL0183083, BL0183080, BL0183079 |
| Therapeutic Target Database | T32262 |
| ChEMBL | CHEMBL3798 |
| IUPHAR | N/A |
| DrugBank | BE0009009 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 774 | 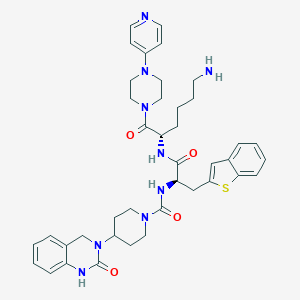 CHEMBL3288289 CHEMBL3288289 | C40H49N9O4S | 751.951 | 8 / 4 | 3.2 | No |
| 1500 | 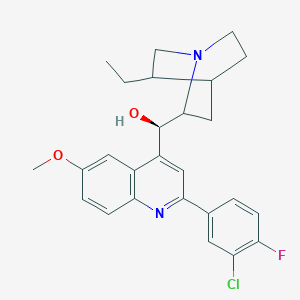 CHEMBL313797 CHEMBL313797 | C26H28ClFN2O2 | 454.97 | 5 / 1 | 5.5 | No |
| 1828 | 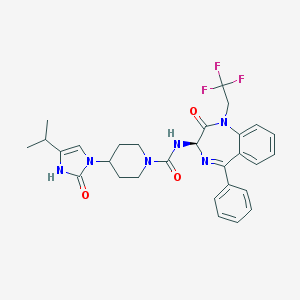 CHEMBL385399 CHEMBL385399 | C29H31F3N6O3 | 568.601 | 7 / 2 | 4.0 | No |
| 1973 | 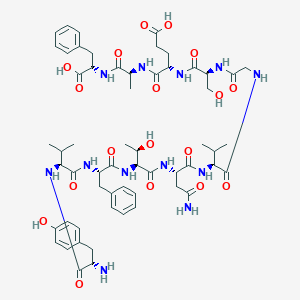 CHEMBL262961 CHEMBL262961 | C58H80N12O18 | 1233.34 | 19 / 17 | -3.3 | No |
| 2410 | 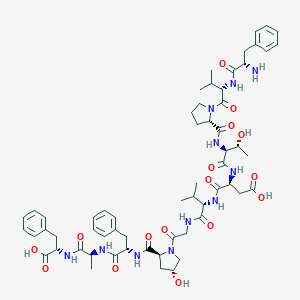 CHEMBL265906 CHEMBL265906 | C60H81N11O16 | 1212.37 | 17 / 13 | -1.1 | No |
| 2416 | 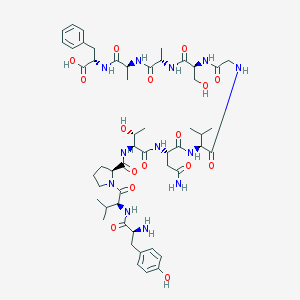 CHEMBL402699 CHEMBL402699 | C52H76N12O16 | 1125.25 | 17 / 15 | -4.1 | No |
| 3368 | 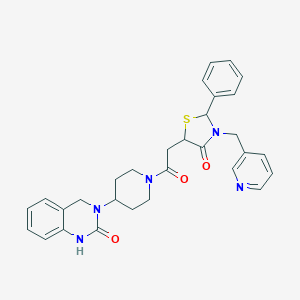 CHEMBL3114468 CHEMBL3114468 | C30H31N5O3S | 541.67 | 5 / 1 | 2.7 | No |
| 3604 | 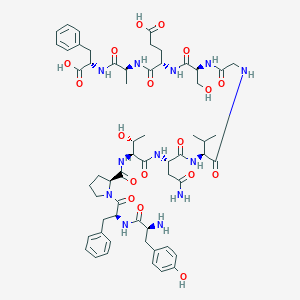 CHEMBL376040 CHEMBL376040 | C58H78N12O18 | 1231.33 | 19 / 16 | -4.0 | No |
| 5091 | 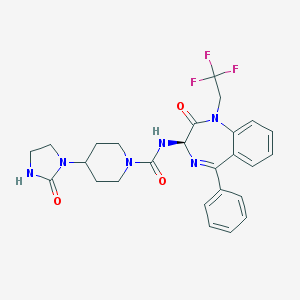 CHEMBL425807 CHEMBL425807 | C26H27F3N6O3 | 528.536 | 7 / 2 | 2.8 | No |
| 6946 | 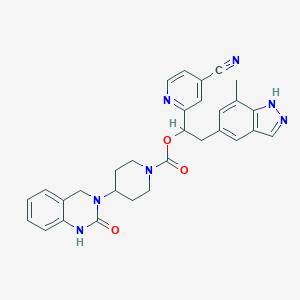 CHEMBL2022597 CHEMBL2022597 | C30H29N7O3 | 535.608 | 6 / 2 | 3.3 | No |
| 7205 | 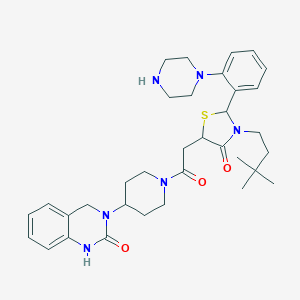 CHEMBL3114474 CHEMBL3114474 | C34H46N6O3S | 618.841 | 6 / 2 | 3.8 | No |
| 7409 | 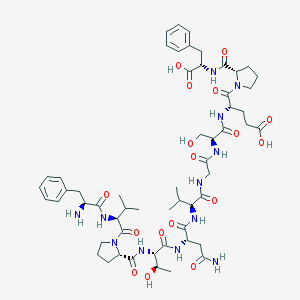 CHEMBL384703 CHEMBL384703 | C56H80N12O17 | 1193.32 | 18 / 14 | -3.9 | No |
| 7477 | 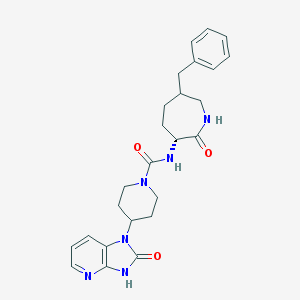 CHEMBL241117 CHEMBL241117 | C25H30N6O3 | 462.554 | 4 / 3 | 2.0 | Yes |
| 9319 | 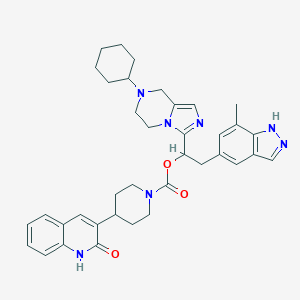 CHEMBL2430166 CHEMBL2430166 | C37H43N7O3 | 633.797 | 6 / 2 | 4.8 | No |
| 9346 | 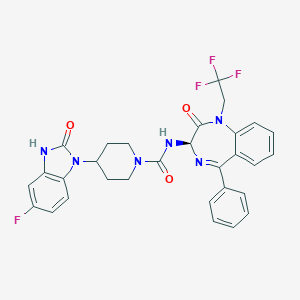 CHEMBL386706 CHEMBL386706 | C30H26F4N6O3 | 594.571 | 8 / 2 | 4.3 | No |
| 9390 | 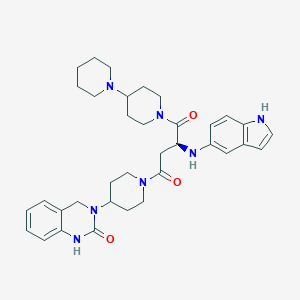 CHEMBL2018498 CHEMBL2018498 | C35H45N7O3 | 611.791 | 5 / 3 | 4.0 | No |
| 9558 | 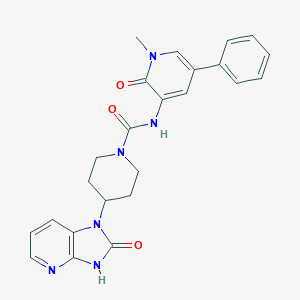 CHEMBL258070 CHEMBL258070 | C24H24N6O3 | 444.495 | 4 / 2 | 1.3 | Yes |
| 10103 | 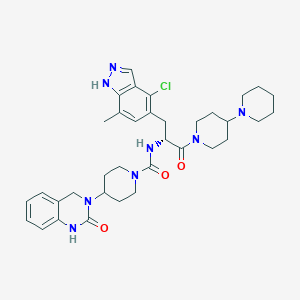 CHEMBL2336418 CHEMBL2336418 | C35H45ClN8O3 | 661.248 | 5 / 3 | 4.1 | No |
| 10574 | 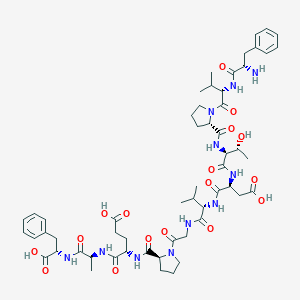 CHEMBL274851 CHEMBL274851 | C56H79N11O17 | 1178.31 | 18 / 13 | -2.2 | No |
| 10881 | 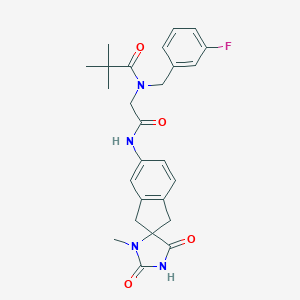 CHEMBL571580 CHEMBL571580 | C26H29FN4O4 | 480.54 | 5 / 2 | 2.7 | Yes |
| 14984 | 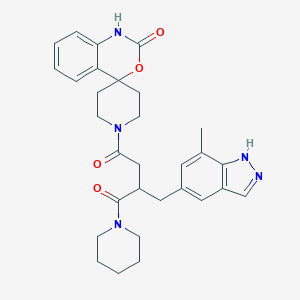 CHEMBL2018512 CHEMBL2018512 | C30H35N5O4 | 529.641 | 5 / 2 | 3.1 | No |
| 15082 | 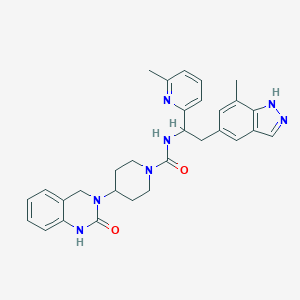 CHEMBL2024601 CHEMBL2024601 | C30H33N7O2 | 523.641 | 4 / 3 | 3.4 | No |
| 15152 | 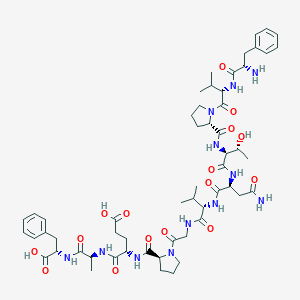 CHEMBL407854 CHEMBL407854 | C56H80N12O16 | 1177.32 | 17 / 13 | -2.9 | No |
| 15426 | 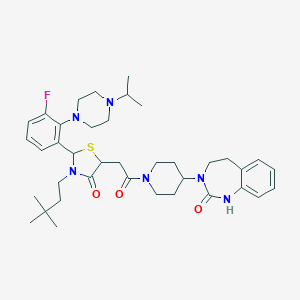 CHEMBL3114495 CHEMBL3114495 | C38H53FN6O3S | 692.939 | 7 / 1 | 5.7 | No |
| 536396 | 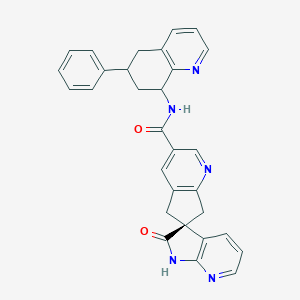 CHEMBL3943580 CHEMBL3943580 | C30H25N5O2 | 487.563 | 5 / 2 | 2.8 | Yes |
| 557733 | 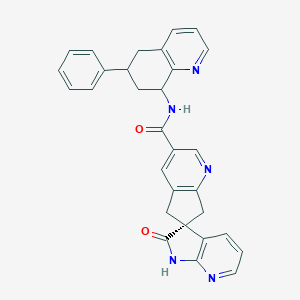 US9227973, 5 isomer B US9227973, 5 isomer B | C30H25N5O2 | 487.563 | 5 / 2 | 2.8 | Yes |
| 17061 | 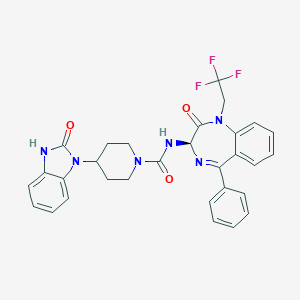 CHEMBL381798 CHEMBL381798 | C30H27F3N6O3 | 576.58 | 7 / 2 | 4.2 | No |
| 17549 | 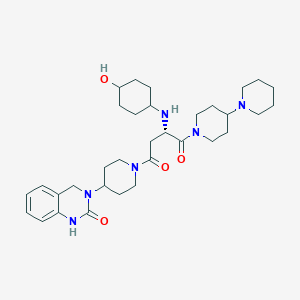 CHEMBL2018357 CHEMBL2018357 | C33H50N6O4 | 594.801 | 6 / 3 | 2.4 | No |
| 17935 | 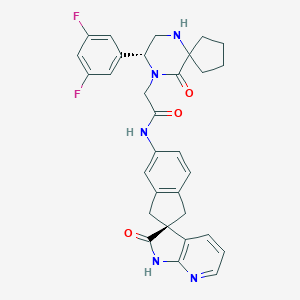 MK-3207 MK-3207 | C31H29F2N5O3 | 557.602 | 7 / 3 | 2.8 | No |
| 18657 | 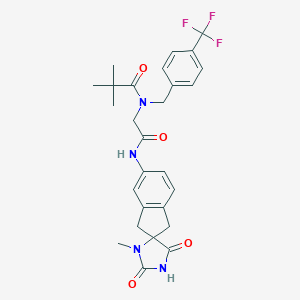 CHEMBL577304 CHEMBL577304 | C27H29F3N4O4 | 530.548 | 7 / 2 | 3.5 | No |
| 18894 | 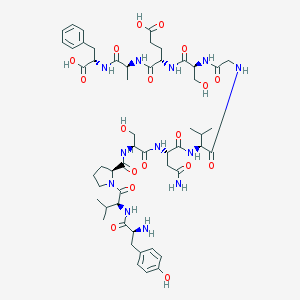 CHEMBL263335 CHEMBL263335 | C53H76N12O18 | 1169.26 | 19 / 16 | -5.0 | No |
| 19315 | 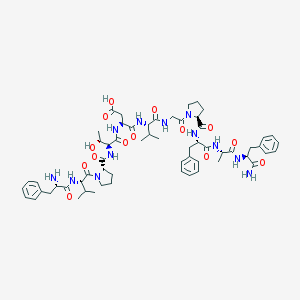 CHEMBL376719 CHEMBL376719 | C60H82N12O14 | 1195.39 | 15 / 12 | -0.8 | No |
| 19885 | 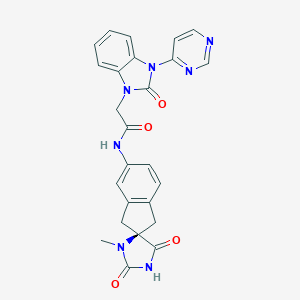 CHEMBL482178 CHEMBL482178 | C25H21N7O4 | 483.488 | 6 / 2 | 1.1 | Yes |
| 20004 | 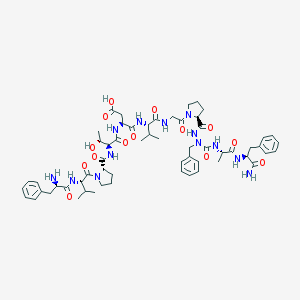 CHEMBL435149 CHEMBL435149 | C59H81N13O14 | 1196.37 | 15 / 12 | -1.1 | No |
| 20812 | 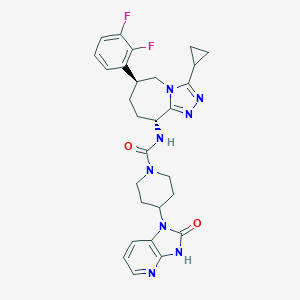 CHEMBL1770556 CHEMBL1770556 | C28H30F2N8O2 | 548.599 | 7 / 2 | 1.9 | No |
| 22063 | 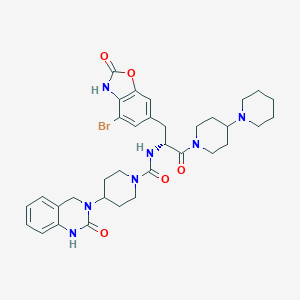 CHEMBL2336416 CHEMBL2336416 | C34H42BrN7O5 | 708.658 | 6 / 3 | 3.4 | No |
| 22614 | 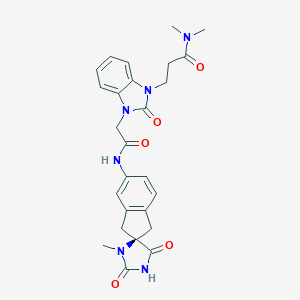 CHEMBL453865 CHEMBL453865 | C26H28N6O5 | 504.547 | 5 / 2 | 0.3 | No |
| 22755 | 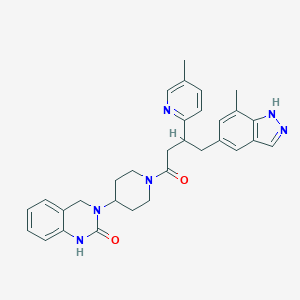 CHEMBL2024587 CHEMBL2024587 | C31H34N6O2 | 522.653 | 4 / 2 | 3.7 | No |
| 24803 | 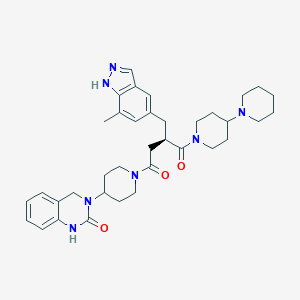 CHEMBL2018518 CHEMBL2018518 | C36H47N7O3 | 625.818 | 5 / 2 | 3.5 | No |
| 24804 | 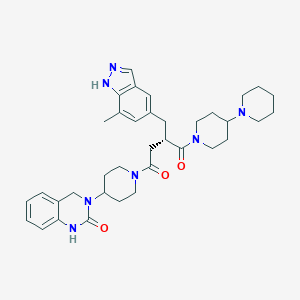 CHEMBL2018517 CHEMBL2018517 | C36H47N7O3 | 625.818 | 5 / 2 | 3.5 | No |
| 24805 | 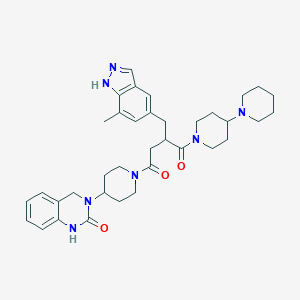 SCHEMBL3590844 SCHEMBL3590844 | C36H47N7O3 | 625.818 | 5 / 2 | 3.5 | No |
| 25268 | 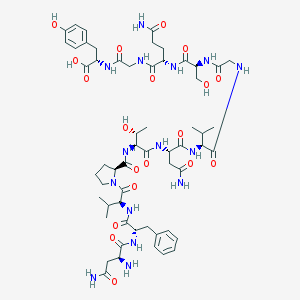 CHEMBL409205 CHEMBL409205 | C57H83N15O19 | 1282.38 | 20 / 18 | -7.4 | No |
| 25644 | 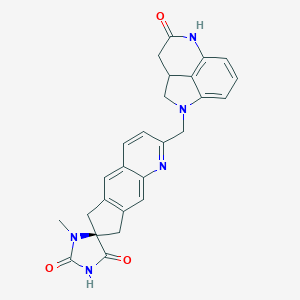 CHEMBL1090841 CHEMBL1090841 | C26H23N5O3 | 453.502 | 5 / 2 | 1.6 | Yes |
| 27204 | 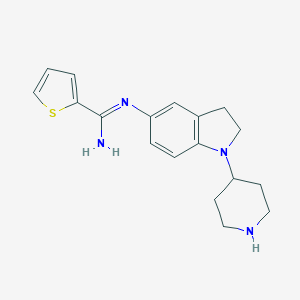 CHEMBL2203713 CHEMBL2203713 | C18H22N4S | 326.462 | 4 / 2 | 2.9 | Yes |
| 27292 | 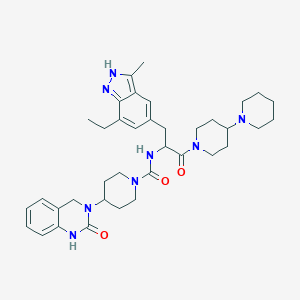 CHEMBL2336419 CHEMBL2336419 | C37H50N8O3 | 654.86 | 5 / 3 | 4.3 | No |
| 27419 | 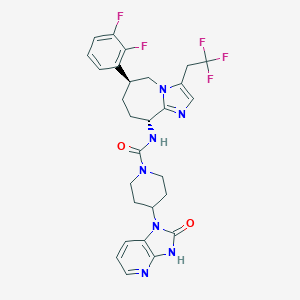 CHEMBL1770557 CHEMBL1770557 | C28H28F5N7O2 | 589.571 | 9 / 2 | 3.3 | No |
| 27775 | 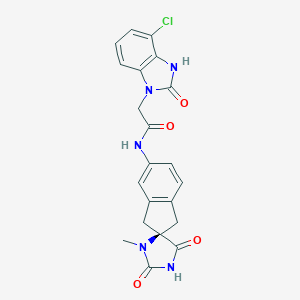 CHEMBL471530 CHEMBL471530 | C21H18ClN5O4 | 439.856 | 4 / 3 | 1.3 | Yes |
| 28514 | 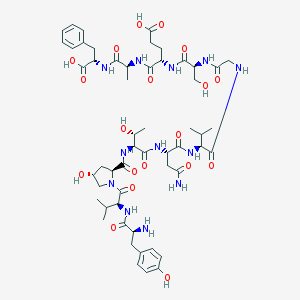 CHEMBL407697 CHEMBL407697 | C54H78N12O19 | 1199.28 | 20 / 17 | -5.6 | No |
| 28717 | 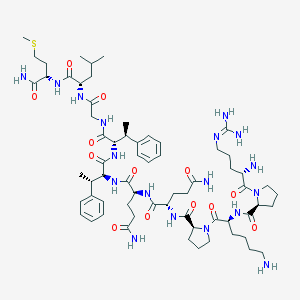 CHEMBL2371143 CHEMBL2371143 | C65H102N18O13S | 1375.7 | 17 / 15 | -1.7 | No |
| 28964 | 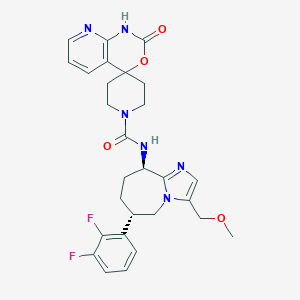 CHEMBL1770723 CHEMBL1770723 | C28H30F2N6O4 | 552.583 | 8 / 2 | 1.7 | No |
| 29167 | 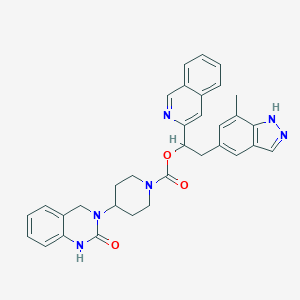 CHEMBL2024599 CHEMBL2024599 | C33H32N6O3 | 560.658 | 5 / 2 | 4.8 | No |
| 29516 | 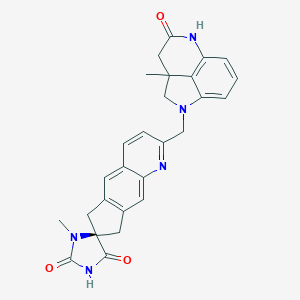 CHEMBL1092838 CHEMBL1092838 | C27H25N5O3 | 467.529 | 5 / 2 | 2.2 | Yes |
| 29567 | 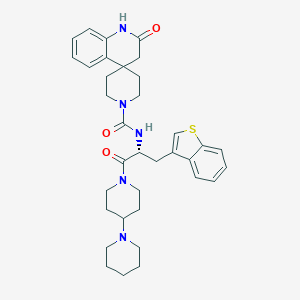 CHEMBL2059896 CHEMBL2059896 | C35H43N5O3S | 613.821 | 5 / 2 | 4.7 | No |
| 29928 | 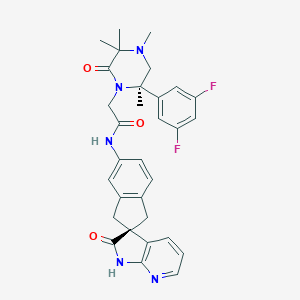 CHEMBL2431247 CHEMBL2431247 | C31H31F2N5O3 | 559.618 | 7 / 2 | 2.9 | No |
| 30106 |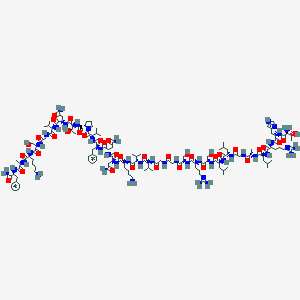  CHEMBL2369463 CHEMBL2369463 | C134H221N43O37 | 3026.51 | 43 / 44 | -10.8 | No |
| 30287 | 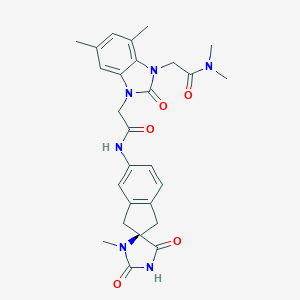 CHEMBL519421 CHEMBL519421 | C27H30N6O5 | 518.574 | 5 / 2 | 1.2 | No |
| 31738 | 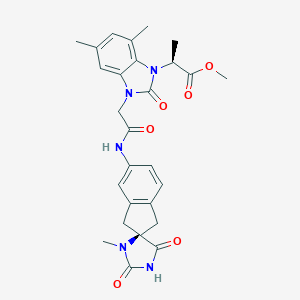 CHEMBL488255 CHEMBL488255 | C27H29N5O6 | 519.558 | 6 / 2 | 1.9 | No |
| 33568 | 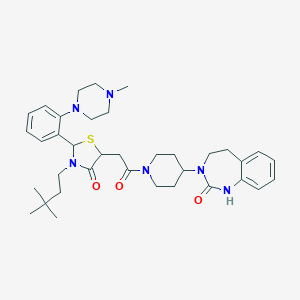 CHEMBL3114488 CHEMBL3114488 | C36H50N6O3S | 646.895 | 6 / 1 | 4.8 | No |
| 33774 | 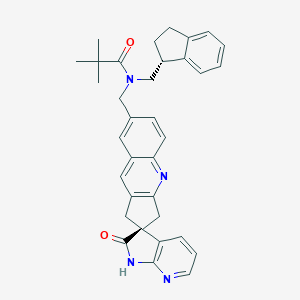 CHEMBL1269645 CHEMBL1269645 | C34H34N4O2 | 530.672 | 4 / 1 | 5.0 | No |
| 34895 | 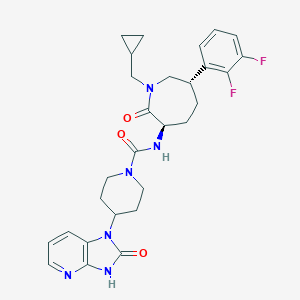 CHEMBL237663 CHEMBL237663 | C28H32F2N6O3 | 538.6 | 6 / 2 | 2.6 | No |
| 35185 | 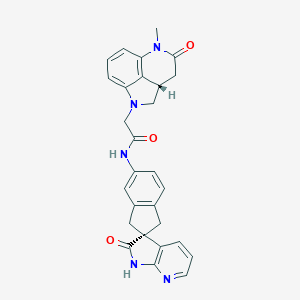 CHEMBL556703 CHEMBL556703 | C28H25N5O3 | 479.54 | 5 / 2 | 1.8 | Yes |
| 35290 | 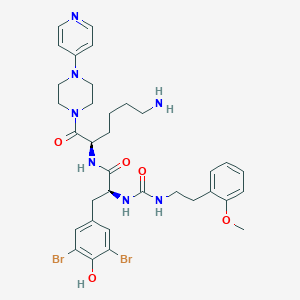 CHEMBL198105 CHEMBL198105 | C34H43Br2N7O5 | 789.57 | 8 / 5 | 3.9 | No |
| 35538 | 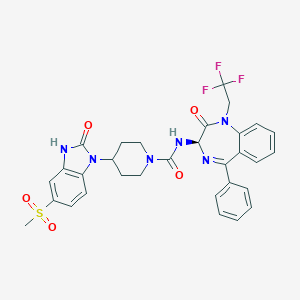 CHEMBL405544 CHEMBL405544 | C31H29F3N6O5S | 654.665 | 9 / 2 | 3.5 | No |
| 36407 | 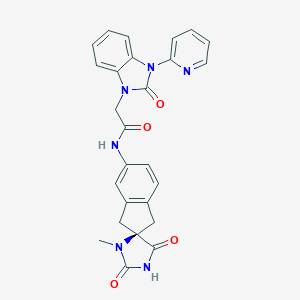 CHEMBL223156 CHEMBL223156 | C26H22N6O4 | 482.5 | 5 / 2 | 1.7 | Yes |
| 37239 | 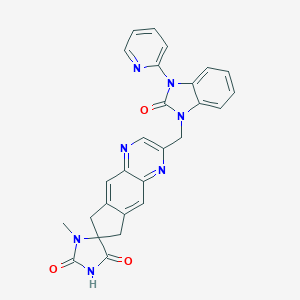 CHEMBL1089207 CHEMBL1089207 | C27H21N7O3 | 491.511 | 6 / 1 | 1.6 | Yes |
| 37298 | 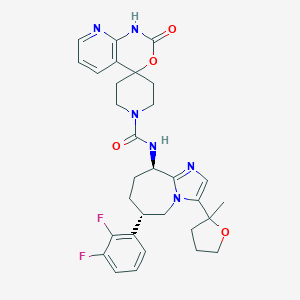 CHEMBL1770729 CHEMBL1770729 | C31H34F2N6O4 | 592.648 | 8 / 2 | 2.4 | No |
| 38342 | 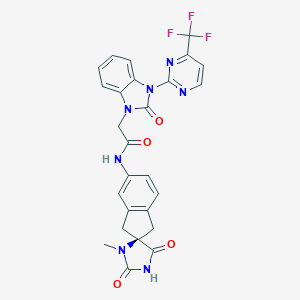 CHEMBL374251 CHEMBL374251 | C26H20F3N7O4 | 551.486 | 9 / 2 | 2.0 | No |
| 38460 | 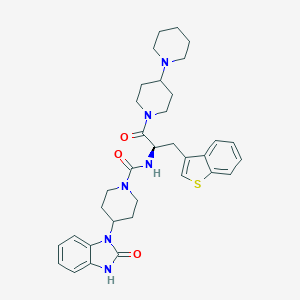 CHEMBL2059880 CHEMBL2059880 | C34H42N6O3S | 614.809 | 5 / 2 | 4.6 | No |
| 38978 | 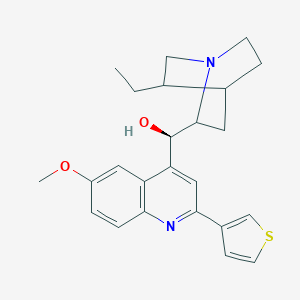 CHEMBL88196 CHEMBL88196 | C24H28N2O2S | 408.56 | 5 / 1 | 4.4 | Yes |
| 40898 | 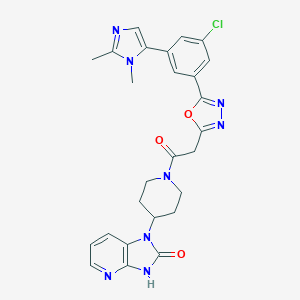 CHEMBL601857 CHEMBL601857 | C26H25ClN8O3 | 532.989 | 7 / 1 | 2.0 | No |
| 41231 | 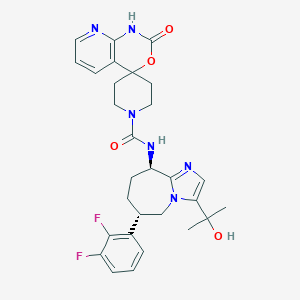 CHEMBL1770725 CHEMBL1770725 | C29H32F2N6O4 | 566.61 | 8 / 3 | 1.8 | No |
| 41233 | 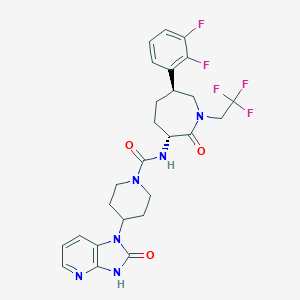 Telcagepant Telcagepant | C26H27F5N6O3 | 566.533 | 9 / 2 | 3.0 | No |
| 41645 | 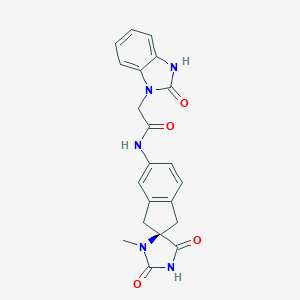 CHEMBL221009 CHEMBL221009 | C21H19N5O4 | 405.414 | 4 / 3 | 0.7 | Yes |
| 42152 | 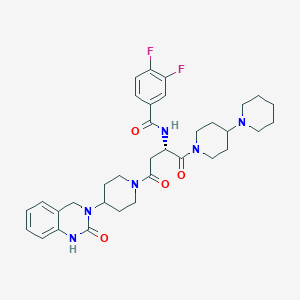 CHEMBL2018358 CHEMBL2018358 | C34H42F2N6O4 | 636.745 | 7 / 2 | 2.8 | No |
| 43882 | 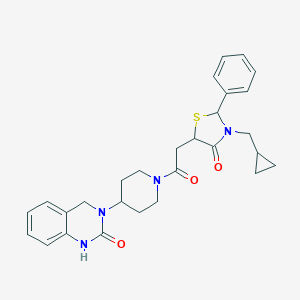 CHEMBL3114455 CHEMBL3114455 | C28H32N4O3S | 504.649 | 4 / 1 | 3.0 | No |
| 44412 | 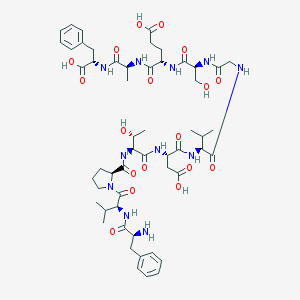 CHEMBL266506 CHEMBL266506 | C54H77N11O18 | 1168.27 | 19 / 15 | -3.6 | No |
| 44522 | 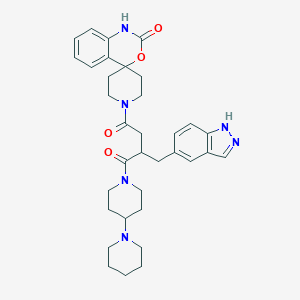 CHEMBL2018514 CHEMBL2018514 | C34H42N6O4 | 598.748 | 6 / 2 | 3.3 | No |
| 45953 | 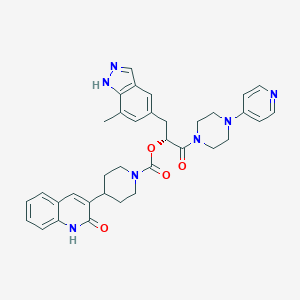 CHEMBL508349 CHEMBL508349 | C35H37N7O4 | 619.726 | 7 / 2 | 3.8 | No |
| 46309 | 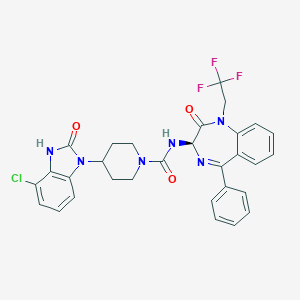 CHEMBL386204 CHEMBL386204 | C30H26ClF3N6O3 | 611.022 | 7 / 2 | 4.9 | No |
| 46632 | 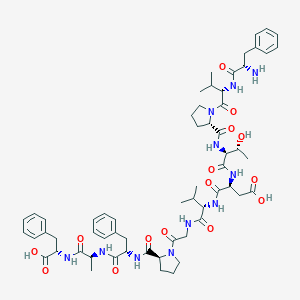 FVPTDVGPFAF FVPTDVGPFAF | C60H81N11O15 | 1196.37 | 16 / 12 | -0.1 | No |
| 47799 | 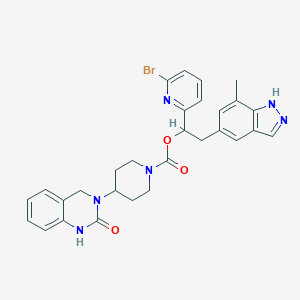 CHEMBL2024595 CHEMBL2024595 | C29H29BrN6O3 | 589.494 | 5 / 2 | 4.6 | No |
| 47936 | 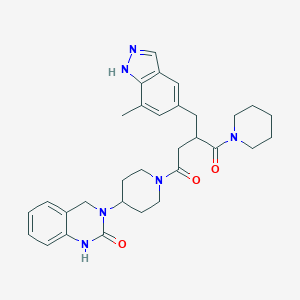 CHEMBL2018513 CHEMBL2018513 | C31H38N6O3 | 542.684 | 4 / 2 | 2.9 | No |
| 49226 | 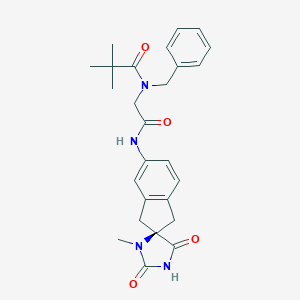 CHEMBL577737 CHEMBL577737 | C26H30N4O4 | 462.55 | 4 / 2 | 2.6 | Yes |
| 49227 | 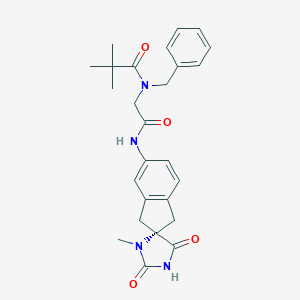 CHEMBL571823 CHEMBL571823 | C26H30N4O4 | 462.55 | 4 / 2 | 2.6 | Yes |
| 49314 | 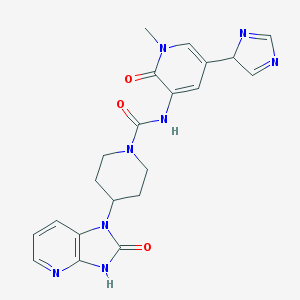 CHEMBL411896 CHEMBL411896 | C21H22N8O3 | 434.46 | 5 / 2 | -1.1 | Yes |
| 49850 | 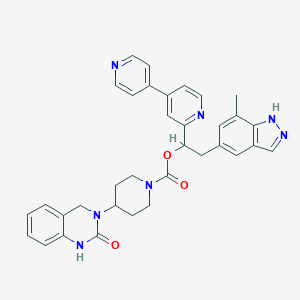 CHEMBL2022599 CHEMBL2022599 | C34H33N7O3 | 587.684 | 6 / 2 | 4.1 | No |
| 50090 | 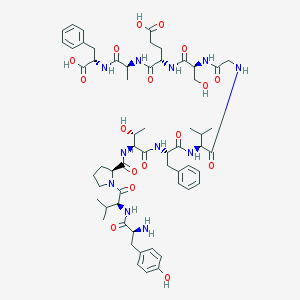 CHEMBL266666 CHEMBL266666 | C59H81N11O17 | 1216.36 | 18 / 15 | -1.5 | No |
| 50644 | 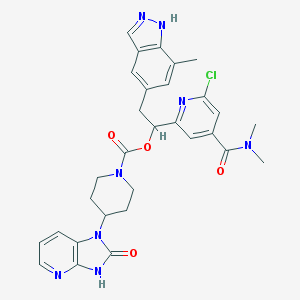 CHEMBL2023191 CHEMBL2023191 | C30H31ClN8O4 | 603.08 | 7 / 2 | 3.3 | No |
| 51292 | 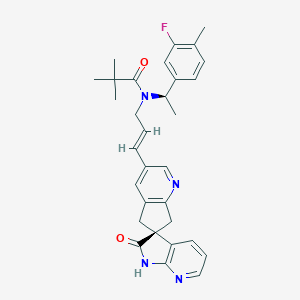 CHEMBL3099923 CHEMBL3099923 | C31H33FN4O2 | 512.629 | 5 / 1 | 4.5 | No |
| 51711 | 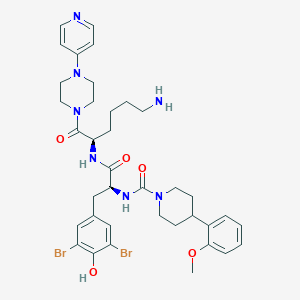 CHEMBL194632 CHEMBL194632 | C37H47Br2N7O5 | 829.635 | 8 / 4 | 4.3 | No |
| 52458 | 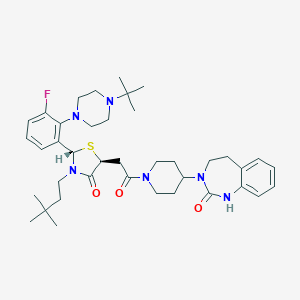 CHEMBL3114677 CHEMBL3114677 | C39H55FN6O3S | 706.966 | 7 / 1 | 5.8 | No |
| 52459 | 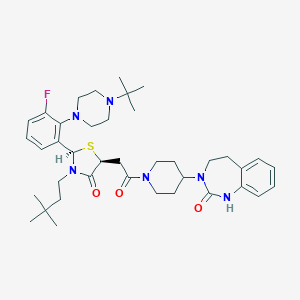 CHEMBL3114676 CHEMBL3114676 | C39H55FN6O3S | 706.966 | 7 / 1 | 5.8 | No |
| 52460 | 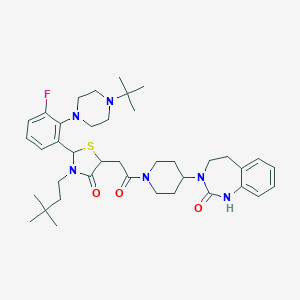 CHEMBL3114496 CHEMBL3114496 | C39H55FN6O3S | 706.966 | 7 / 1 | 5.8 | No |
| 53224 | 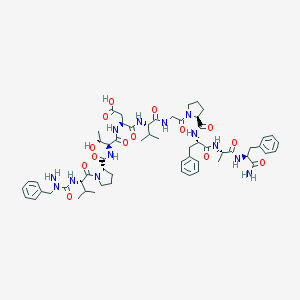 azaFVPTDVGPFAF-NH2 azaFVPTDVGPFAF-NH2 | C59H81N13O14 | 1196.37 | 15 / 12 | -1.1 | No |
| 53936 | 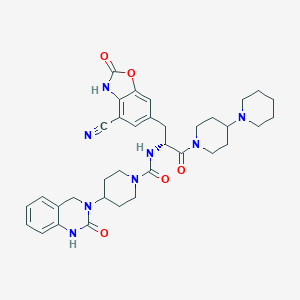 CHEMBL2336414 CHEMBL2336414 | C35H42N8O5 | 654.772 | 7 / 3 | 2.5 | No |
| 55526 | 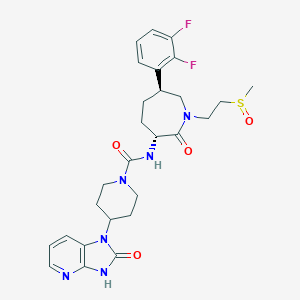 CHEMBL392730 CHEMBL392730 | C27H32F2N6O4S | 574.648 | 8 / 2 | 1.0 | No |
| 57714 | 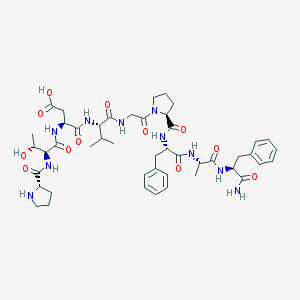 CHEMBL223642 CHEMBL223642 | C46H64N10O12 | 949.076 | 13 / 11 | -2.5 | No |
| 58796 | 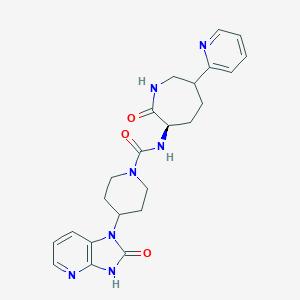 CHEMBL398779 CHEMBL398779 | C23H27N7O3 | 449.515 | 5 / 3 | 0.5 | Yes |
| 59365 | 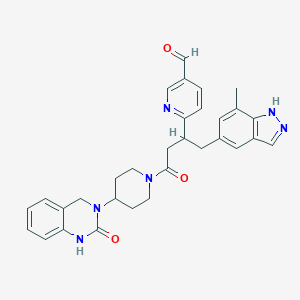 CHEMBL2024589 CHEMBL2024589 | C31H32N6O3 | 536.636 | 5 / 2 | 2.8 | No |
| 59400 | 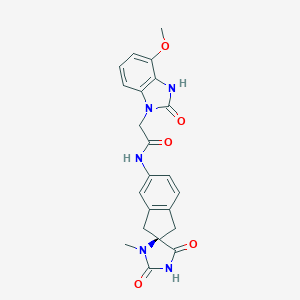 CHEMBL471704 CHEMBL471704 | C22H21N5O5 | 435.44 | 5 / 3 | 0.7 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218