

You can:
| Name | Free fatty acid receptor 4 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | FFAR4 |
| Synonym | PGR4 Omega-3 fatty acid receptor 1 O3FAR1 GT01 GPR129 [ Show all ] |
| Disease | N/A |
| Length | 377 |
| Amino acid sequence | MSPECARAAGDAPLRSLEQANRTRFPFFSDVKGDHRLVLAAVETTVLVLIFAVSLLGNVCALVLVARRRRRGATACLVLNLFCADLLFISAIPLVLAVRWTEAWLLGPVACHLLFYVMTLSGSVTILTLAAVSLERMVCIVHLQRGVRGPGRRARAVLLALIWGYSAVAALPLCVFFRVVPQRLPGADQEISICTLIWPTIPGEISWDVSFVTLNFLVPGLVIVISYSKILQTSEHLLDARAVVTHSEITKASRKRLTVSLAYSESHQIRVSQQDFRLFRTLFLLMVSFFIMWSPIIITILLILIQNFKQDLVIWPSLFFWVVAFTFANSALNPILYNMTLCRNEWKKIFCCFWFPEKGAILTDTSVKRNDLSIISG |
| UniProt | Q5NUL3 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q5NUL3 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q5NUL3. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL5339 |
| IUPHAR | 127 |
| DrugBank | BE0003399 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 557375 | 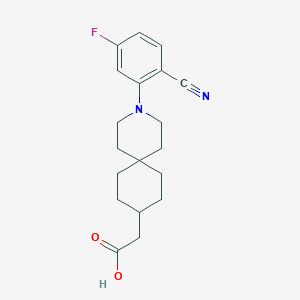 SCHEMBL16483386 SCHEMBL16483386 | C19H23FN2O2 | 330.403 | 5 / 1 | 4.2 | Yes |
| 547935 | 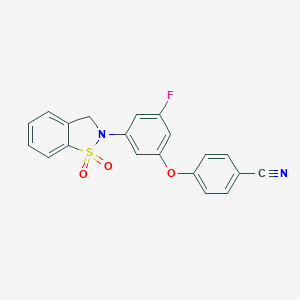 CHEMBL3979588 CHEMBL3979588 | C20H13FN2O3S | 380.393 | 6 / 0 | 3.5 | Yes |
| 547938 | 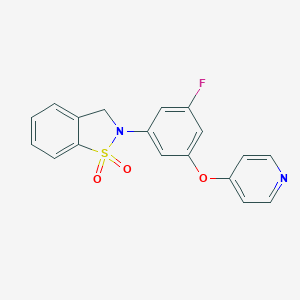 CHEMBL3971461 CHEMBL3971461 | C18H13FN2O3S | 356.371 | 6 / 0 | 2.8 | Yes |
| 547955 | 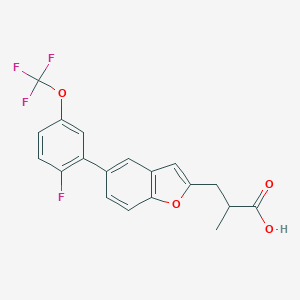 CHEMBL3921062 CHEMBL3921062 | C19H14F4O4 | 382.311 | 8 / 1 | 5.6 | No |
| 547963 | 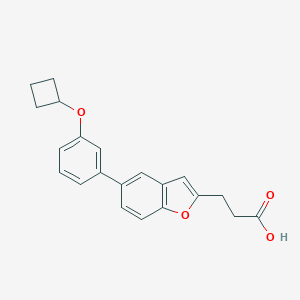 CHEMBL3968537 CHEMBL3968537 | C21H20O4 | 336.387 | 4 / 1 | 4.6 | Yes |
| 7828 | 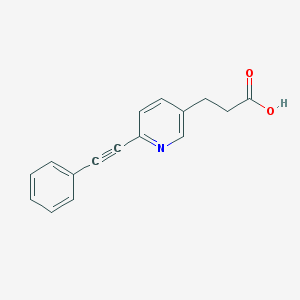 CHEMBL2386357 CHEMBL2386357 | C16H13NO2 | 251.285 | 3 / 1 | 2.7 | Yes |
| 8965 | 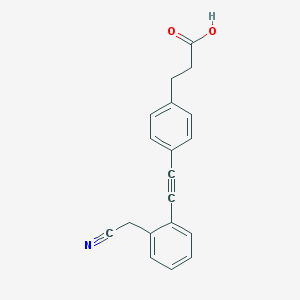 CHEMBL2315528 CHEMBL2315528 | C19H15NO2 | 289.334 | 3 / 1 | 3.4 | Yes |
| 442091 | 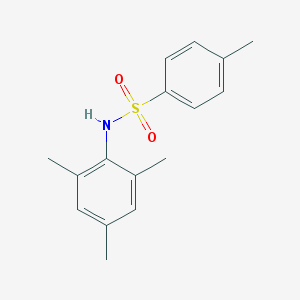 CHEMBL3311314 CHEMBL3311314 | C16H19NO2S | 289.393 | 3 / 1 | 3.9 | Yes |
| 548019 | 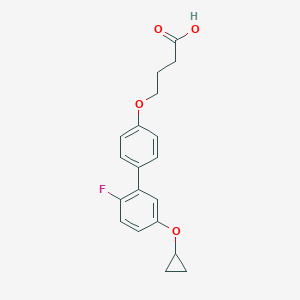 CHEMBL3944507 CHEMBL3944507 | C19H19FO4 | 330.355 | 5 / 1 | 4.0 | Yes |
| 553322 | 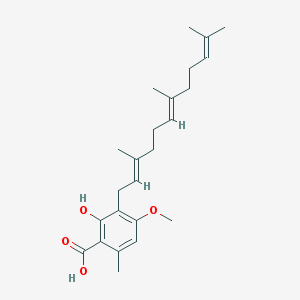 2-hydroxy-4-methoxy-6-methyl-3-[(2E,6E)-3,7,11-trimethyldodeca-2,6,10-trienyl]benzoic Acid 2-hydroxy-4-methoxy-6-methyl-3-[(2E,6E)-3,7,11-trimethyldodeca-2,6,10-trienyl]benzoic Acid | C24H34O4 | 386.532 | 4 / 2 | 7.6 | No |
| 548043 | 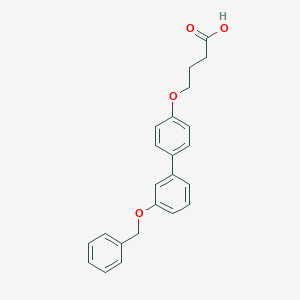 CHEMBL3980036 CHEMBL3980036 | C23H22O4 | 362.425 | 4 / 1 | 4.8 | Yes |
| 557695 | 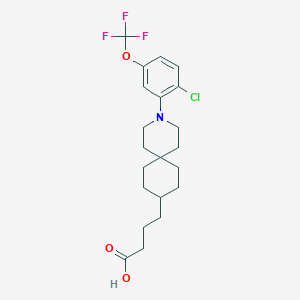 SCHEMBL16482978 SCHEMBL16482978 | C21H27ClF3NO3 | 433.896 | 7 / 1 | 7.1 | No |
| 15877 | 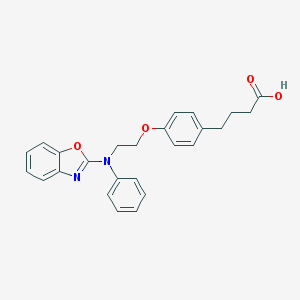 CHEMBL508388 CHEMBL508388 | C25H24N2O4 | 416.477 | 6 / 1 | 5.6 | No |
| 19612 | 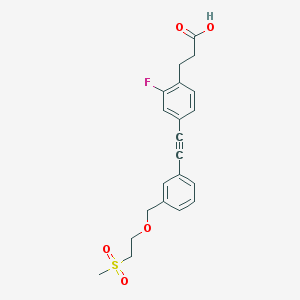 CHEMBL2386369 CHEMBL2386369 | C21H21FO5S | 404.452 | 6 / 1 | 2.8 | Yes |
| 557956 | 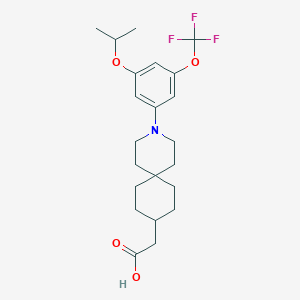 SCHEMBL16483002 SCHEMBL16483002 | C22H30F3NO4 | 429.48 | 8 / 1 | 6.3 | No |
| 558164 | 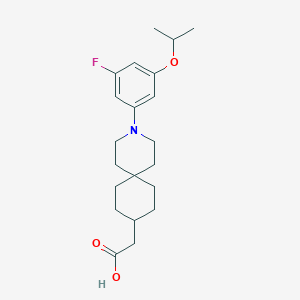 SCHEMBL16483003 SCHEMBL16483003 | C21H30FNO3 | 363.473 | 5 / 1 | 5.2 | No |
| 548230 | 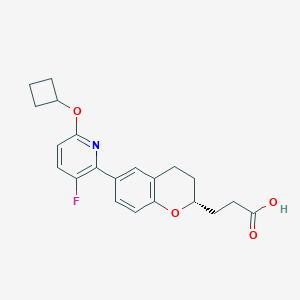 CHEMBL3946820 CHEMBL3946820 | C21H22FNO4 | 371.408 | 6 / 1 | 4.1 | Yes |
| 522505 | 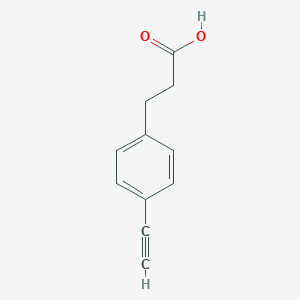 3-(4-ethynylphenyl)propanoic acid 3-(4-ethynylphenyl)propanoic acid | C11H10O2 | 174.199 | 2 / 1 | 2.1 | Yes |
| 548270 | 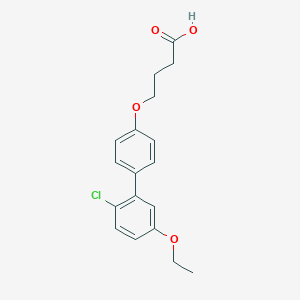 CHEMBL3924759 CHEMBL3924759 | C18H19ClO4 | 334.796 | 4 / 1 | 4.7 | Yes |
| 558278 | 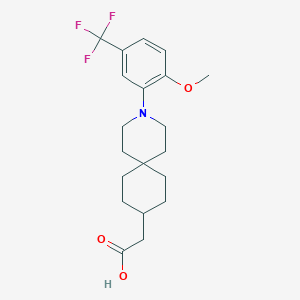 SCHEMBL16483468 SCHEMBL16483468 | C20H26F3NO3 | 385.427 | 7 / 1 | 5.2 | No |
| 467071 | 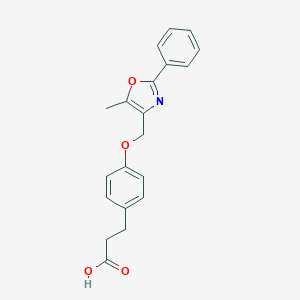 CHEMBL3600993 CHEMBL3600993 | C20H19NO4 | 337.375 | 5 / 1 | 3.7 | Yes |
| 34523 | 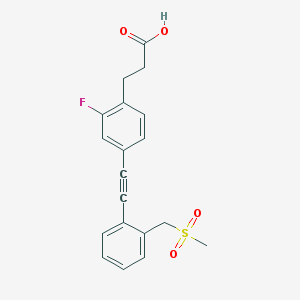 CHEMBL2386366 CHEMBL2386366 | C19H17FO4S | 360.399 | 5 / 1 | 3.0 | Yes |
| 34587 | 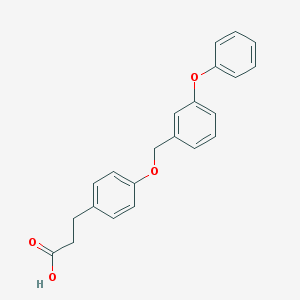 CHEMBL1688458 CHEMBL1688458 | C22H20O4 | 348.398 | 4 / 1 | 4.7 | Yes |
| 36310 | 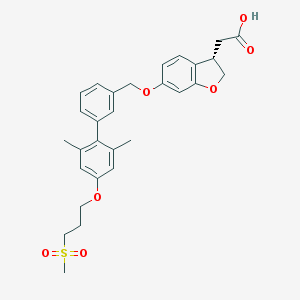 TAK-875 TAK-875 | C29H32O7S | 524.628 | 7 / 1 | 4.7 | No |
| 443305 | 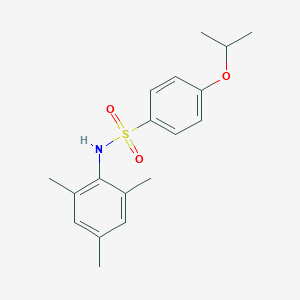 CHEMBL3311322 CHEMBL3311322 | C18H23NO3S | 333.446 | 4 / 1 | 4.3 | Yes |
| 443309 | 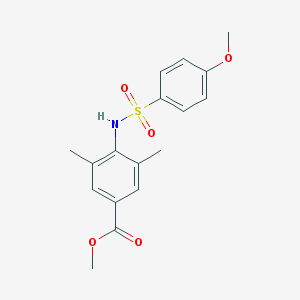 CHEMBL3311201 CHEMBL3311201 | C17H19NO5S | 349.401 | 6 / 1 | 3.0 | Yes |
| 41620 | 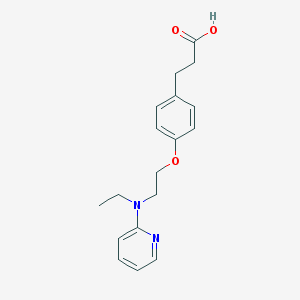 CHEMBL181264 CHEMBL181264 | C18H22N2O3 | 314.385 | 5 / 1 | 3.1 | Yes |
| 537156 | 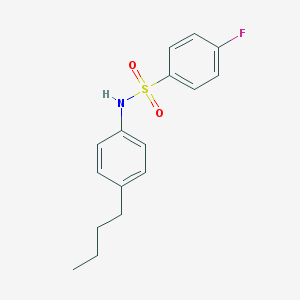 DC260126 DC260126 | C16H18FNO2S | 307.383 | 4 / 1 | 4.4 | Yes |
| 558720 | 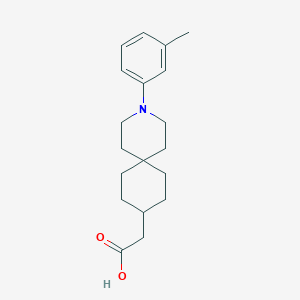 SCHEMBL16482992 SCHEMBL16482992 | C19H27NO2 | 301.43 | 3 / 1 | 4.7 | Yes |
| 48805 | 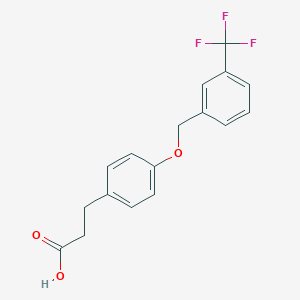 CHEMBL1773257 CHEMBL1773257 | C17H15F3O3 | 324.299 | 6 / 1 | 4.0 | Yes |
| 548481 | 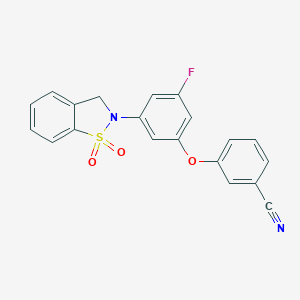 CHEMBL3923955 CHEMBL3923955 | C20H13FN2O3S | 380.393 | 6 / 0 | 3.5 | Yes |
| 53290 | 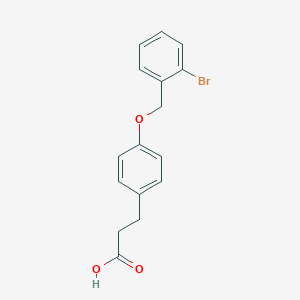 CHEMBL1773253 CHEMBL1773253 | C16H15BrO3 | 335.197 | 3 / 1 | 3.8 | Yes |
| 558971 | 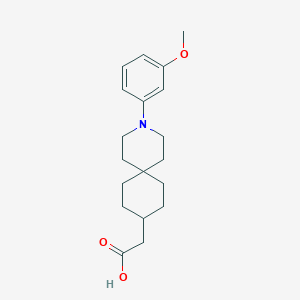 SCHEMBL16482993 SCHEMBL16482993 | C19H27NO3 | 317.429 | 4 / 1 | 4.3 | Yes |
| 523131 | 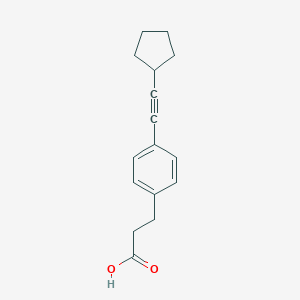 CHEMBL3787110 CHEMBL3787110 | C16H18O2 | 242.318 | 2 / 1 | 4.1 | Yes |
| 558997 | 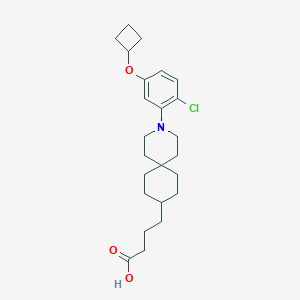 SCHEMBL16483416 SCHEMBL16483416 | C24H34ClNO3 | 419.99 | 4 / 1 | 6.8 | No |
| 548523 | 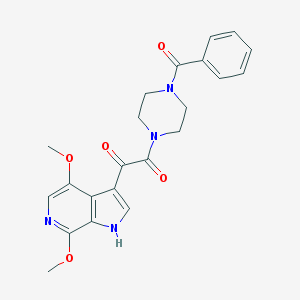 BMS-488043 BMS-488043 | C22H22N4O5 | 422.441 | 6 / 1 | 1.5 | Yes |
| 58864 | 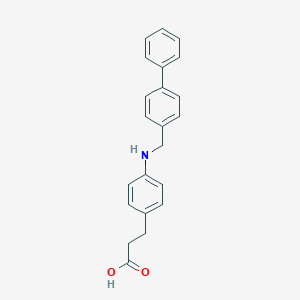 carboxylic acid agonist, 19 carboxylic acid agonist, 19 | C22H21NO2 | 331.415 | 3 / 2 | 4.8 | Yes |
| 59169 | 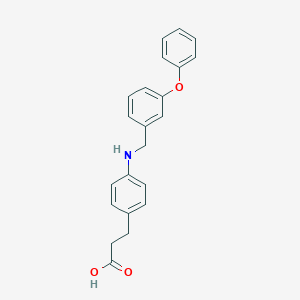 GW9508 GW9508 | C22H21NO3 | 347.414 | 4 / 2 | 4.7 | Yes |
| 537470 | 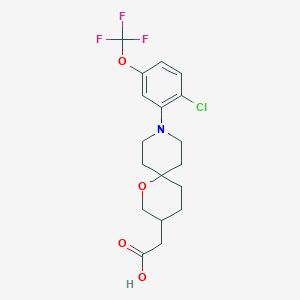 CHEMBL3986483 CHEMBL3986483 | C18H21ClF3NO4 | 407.814 | 8 / 1 | 4.3 | Yes |
| 444185 | 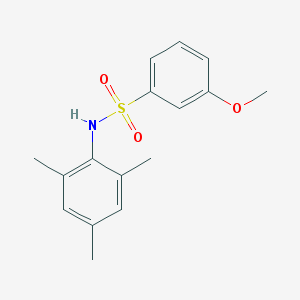 CHEMBL3311312 CHEMBL3311312 | C16H19NO3S | 305.392 | 4 / 1 | 3.5 | Yes |
| 64084 | 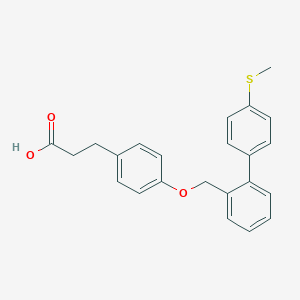 CHEMBL2058528 CHEMBL2058528 | C23H22O3S | 378.486 | 4 / 1 | 5.3 | No |
| 559334 | 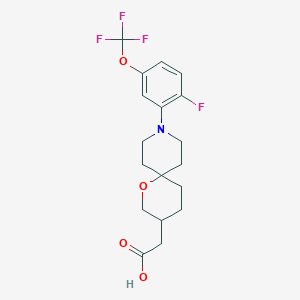 SCHEMBL16466351 SCHEMBL16466351 | C18H21F4NO4 | 391.363 | 9 / 1 | 3.8 | Yes |
| 559388 | 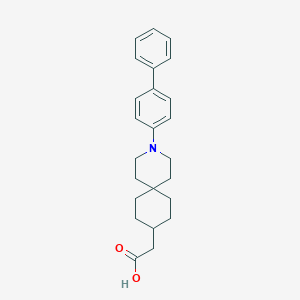 SCHEMBL16483023 SCHEMBL16483023 | C24H29NO2 | 363.501 | 3 / 1 | 6.0 | No |
| 68526 | 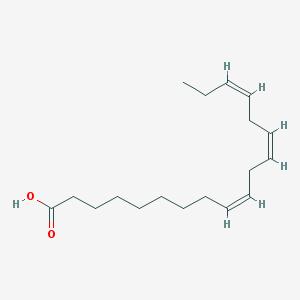 linolenic acid linolenic acid | C18H30O2 | 278.436 | 2 / 1 | 5.9 | No |
| 559493 | 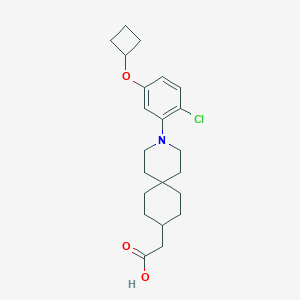 SCHEMBL16482968 SCHEMBL16482968 | C22H30ClNO3 | 391.936 | 4 / 1 | 5.9 | No |
| 537732 | 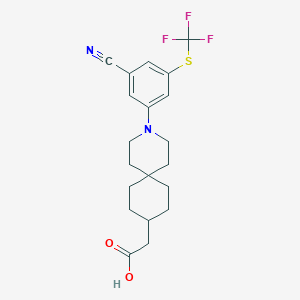 CHEMBL3953289 CHEMBL3953289 | C20H23F3N2O2S | 412.471 | 8 / 1 | 5.8 | No |
| 72277 | 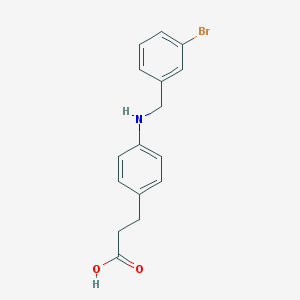 carboxylic acid agonist, 27 carboxylic acid agonist, 27 | C16H16BrNO2 | 334.213 | 3 / 2 | 3.8 | Yes |
| 523729 | 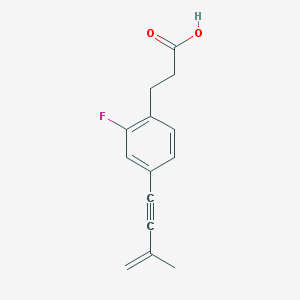 CHEMBL3787463 CHEMBL3787463 | C14H13FO2 | 232.254 | 3 / 1 | 3.6 | Yes |
| 559662 | 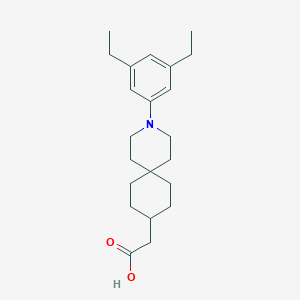 SCHEMBL16482969 SCHEMBL16482969 | C22H33NO2 | 343.511 | 3 / 1 | 5.9 | No |
| 444727 | 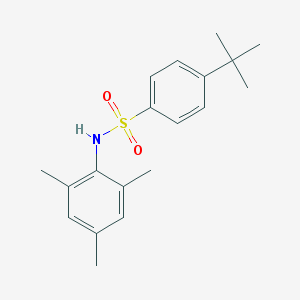 CHEMBL3311320 CHEMBL3311320 | C19H25NO2S | 331.474 | 3 / 1 | 5.2 | No |
| 548797 | 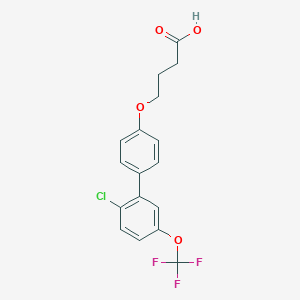 CHEMBL3968947 CHEMBL3968947 | C17H14ClF3O4 | 374.74 | 7 / 1 | 5.5 | No |
| 548850 | 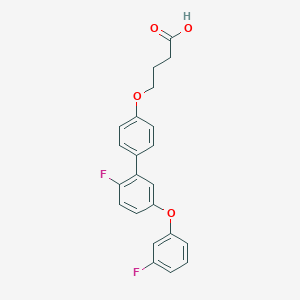 CHEMBL3953203 CHEMBL3953203 | C22H18F2O4 | 384.379 | 6 / 1 | 5.1 | No |
| 83174 | 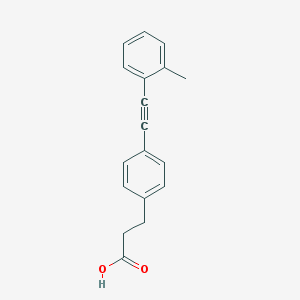 TUG-424 TUG-424 | C18H16O2 | 264.324 | 2 / 1 | 4.1 | Yes |
| 83246 | 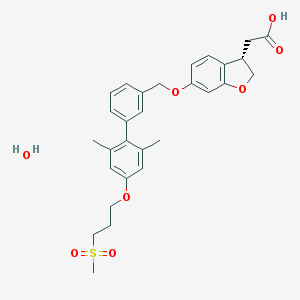 CHEMBL2047159 CHEMBL2047159 | C29H34O8S | 542.643 | 8 / 2 | N/A | No |
| 445023 | 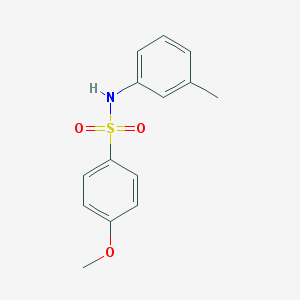 CHEMBL3311304 CHEMBL3311304 | C14H15NO3S | 277.338 | 4 / 1 | 2.6 | Yes |
| 559892 | 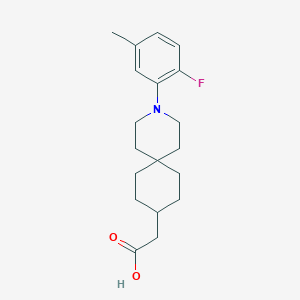 SCHEMBL16482991 SCHEMBL16482991 | C19H26FNO2 | 319.42 | 4 / 1 | 4.8 | Yes |
| 445096 | 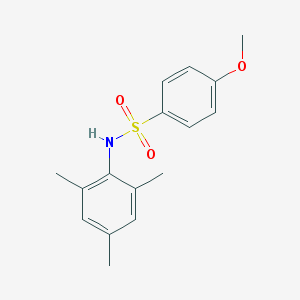 GSK137647A GSK137647A | C16H19NO3S | 305.392 | 4 / 1 | 3.5 | Yes |
| 85591 | 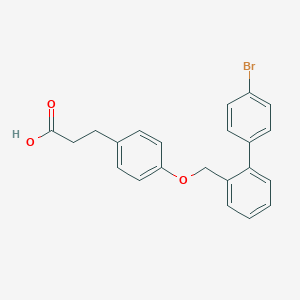 CHEMBL2058523 CHEMBL2058523 | C22H19BrO3 | 411.295 | 3 / 1 | 5.5 | No |
| 559943 | 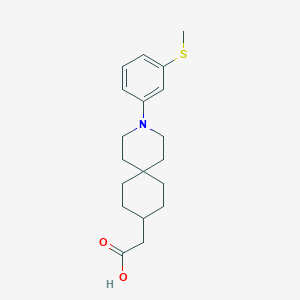 SCHEMBL16482940 SCHEMBL16482940 | C19H27NO2S | 333.49 | 4 / 1 | 4.9 | Yes |
| 87505 | 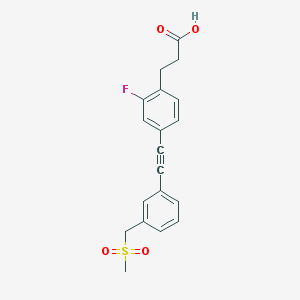 CHEMBL2386367 CHEMBL2386367 | C19H17FO4S | 360.399 | 5 / 1 | 3.0 | Yes |
| 548929 | 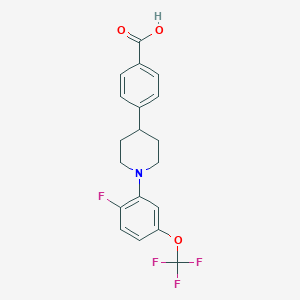 CHEMBL3964985 CHEMBL3964985 | C19H17F4NO3 | 383.343 | 8 / 1 | 5.1 | No |
| 445335 | 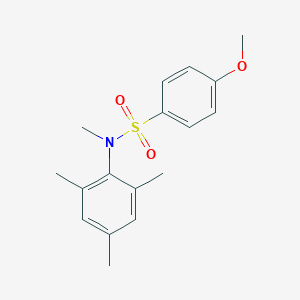 CHEMBL3311309 CHEMBL3311309 | C17H21NO3S | 319.419 | 4 / 0 | 3.7 | Yes |
| 91542 | 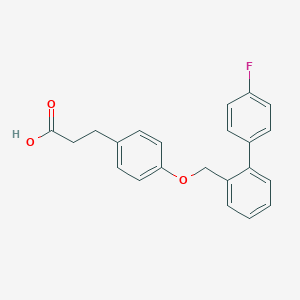 CHEMBL2058521 CHEMBL2058521 | C22H19FO3 | 350.389 | 4 / 1 | 4.9 | Yes |
| 538301 | 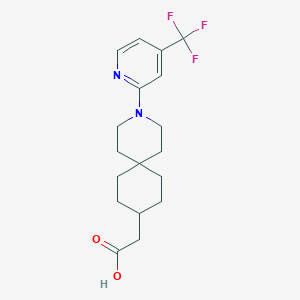 CHEMBL3954140 CHEMBL3954140 | C18H23F3N2O2 | 356.389 | 7 / 1 | 4.5 | Yes |
| 445411 | 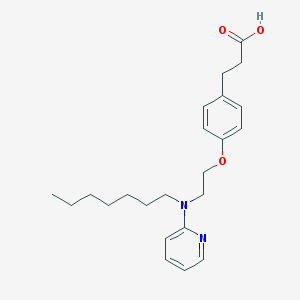 CHEMBL180558 CHEMBL180558 | C23H32N2O3 | 384.52 | 5 / 1 | 5.6 | No |
| 93276 | 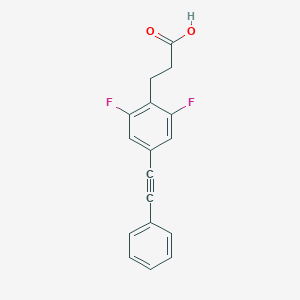 CHEMBL2386360 CHEMBL2386360 | C17H12F2O2 | 286.278 | 4 / 1 | 3.9 | Yes |
| 445619 | 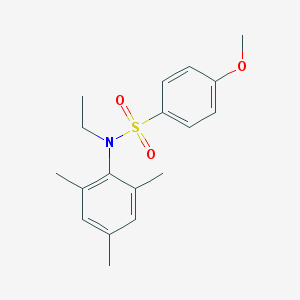 CHEMBL3311310 CHEMBL3311310 | C18H23NO3S | 333.446 | 4 / 0 | 4.1 | Yes |
| 101614 | 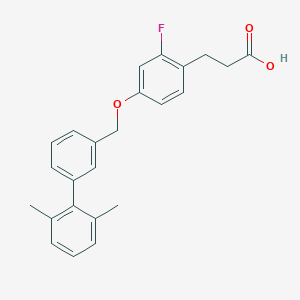 CHEMBL1688473 CHEMBL1688473 | C24H23FO3 | 378.443 | 4 / 1 | 5.6 | No |
| 445776 | 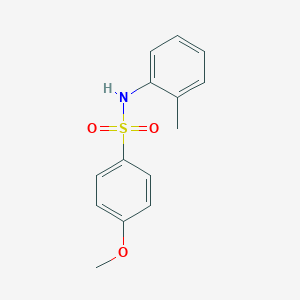 4-methoxy-N-(2-methylphenyl)benzenesulfonamide 4-methoxy-N-(2-methylphenyl)benzenesulfonamide | C14H15NO3S | 277.338 | 4 / 1 | 2.7 | Yes |
| 538565 | 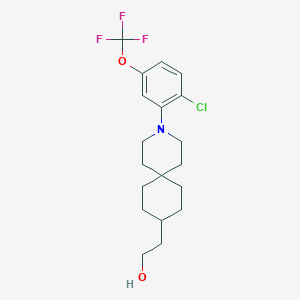 CHEMBL3967504 CHEMBL3967504 | C19H25ClF3NO2 | 391.859 | 6 / 1 | 6.3 | No |
| 538571 | 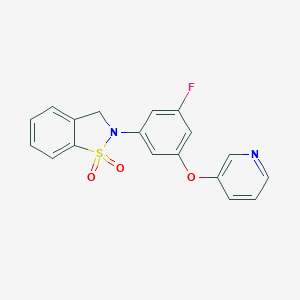 CHEMBL3908362 CHEMBL3908362 | C18H13FN2O3S | 356.371 | 6 / 0 | 2.8 | Yes |
| 538603 | 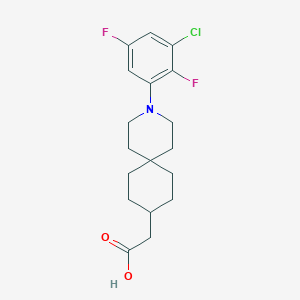 CHEMBL3901994 CHEMBL3901994 | C18H22ClF2NO2 | 357.826 | 5 / 1 | 5.2 | No |
| 560674 | 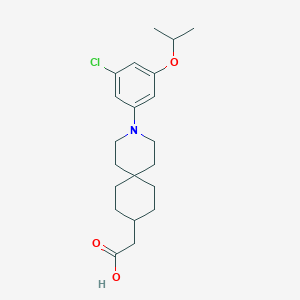 SCHEMBL16482967 SCHEMBL16482967 | C21H30ClNO3 | 379.925 | 4 / 1 | 5.7 | No |
| 549242 | 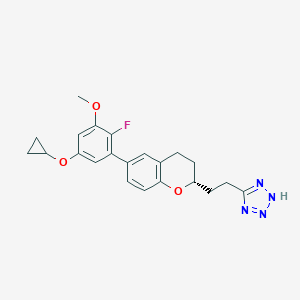 CHEMBL3932664 CHEMBL3932664 | C22H23FN4O3 | 410.449 | 7 / 1 | 4.5 | Yes |
| 112757 | 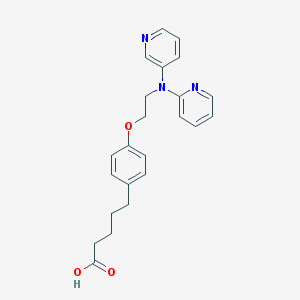 CHEMBL514894 CHEMBL514894 | C23H25N3O3 | 391.471 | 6 / 1 | 4.1 | Yes |
| 549270 | 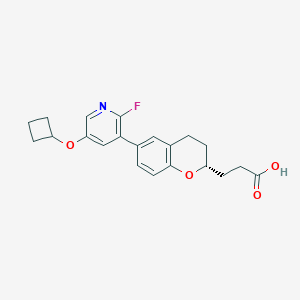 CHEMBL3900705 CHEMBL3900705 | C21H22FNO4 | 371.408 | 6 / 1 | 4.1 | Yes |
| 560832 | 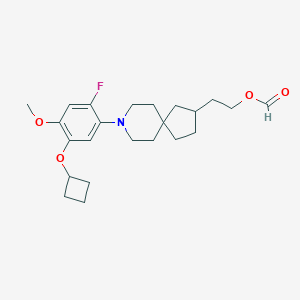 SCHEMBL17125285 SCHEMBL17125285 | C23H32FNO4 | 405.51 | 6 / 0 | 5.5 | No |
| 549283 | 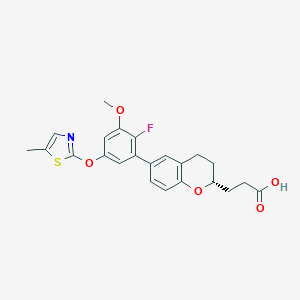 CHEMBL3943315 CHEMBL3943315 | C23H22FNO5S | 443.489 | 8 / 1 | 5.2 | No |
| 560869 | 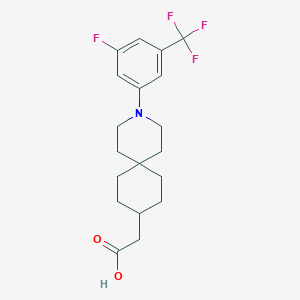 SCHEMBL16482946 SCHEMBL16482946 | C19H23F4NO2 | 373.392 | 7 / 1 | 5.3 | No |
| 560942 | 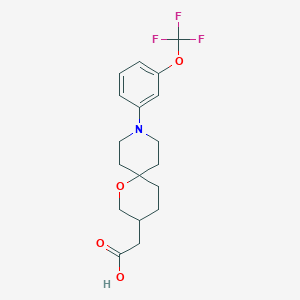 SCHEMBL16483013 SCHEMBL16483013 | C18H22F3NO4 | 373.372 | 8 / 1 | 3.7 | Yes |
| 560979 | 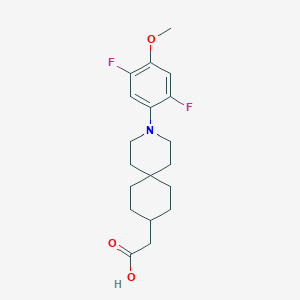 SCHEMBL16482930 SCHEMBL16482930 | C19H25F2NO3 | 353.41 | 6 / 1 | 4.5 | Yes |
| 560997 | 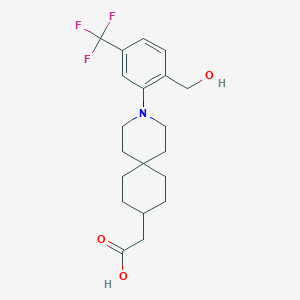 SCHEMBL16484240 SCHEMBL16484240 | C20H26F3NO3 | 385.427 | 7 / 2 | 4.3 | Yes |
| 561072 | 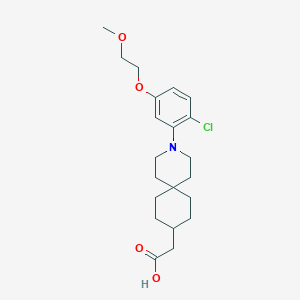 SCHEMBL16482939 SCHEMBL16482939 | C21H30ClNO4 | 395.924 | 5 / 1 | 4.8 | Yes |
| 549399 | 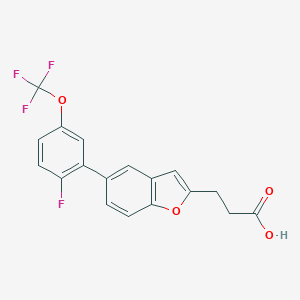 CHEMBL3896657 CHEMBL3896657 | C18H12F4O4 | 368.284 | 8 / 1 | 5.0 | Yes |
| 549416 | 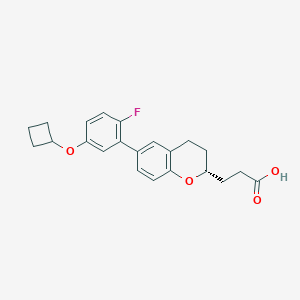 CHEMBL3970238 CHEMBL3970238 | C22H23FO4 | 370.42 | 5 / 1 | 4.8 | Yes |
| 561183 | 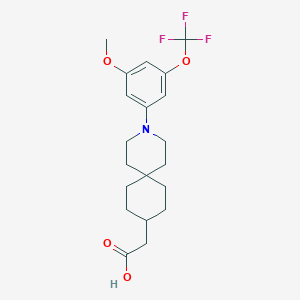 SCHEMBL16482965 SCHEMBL16482965 | C20H26F3NO4 | 401.426 | 8 / 1 | 5.5 | No |
| 539151 | 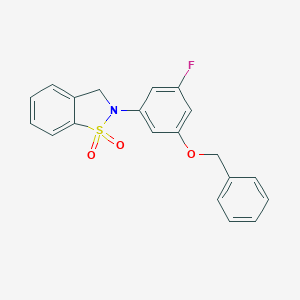 CHEMBL3958418 CHEMBL3958418 | C20H16FNO3S | 369.41 | 5 / 0 | 3.8 | Yes |
| 446738 | 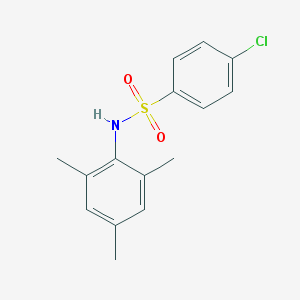 4-chloro-N-mesitylbenzenesulfonamide 4-chloro-N-mesitylbenzenesulfonamide | C15H16ClNO2S | 309.808 | 3 / 1 | 4.2 | Yes |
| 561286 | 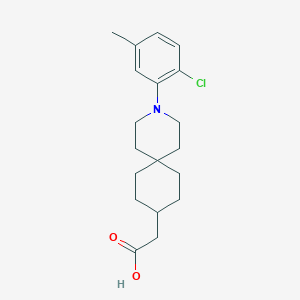 SCHEMBL16482934 SCHEMBL16482934 | C19H26ClNO2 | 335.872 | 3 / 1 | 5.3 | No |
| 132542 | 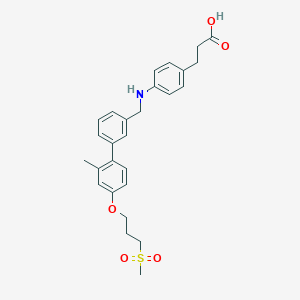 CHEMBL2163924 CHEMBL2163924 | C27H31NO5S | 481.607 | 6 / 2 | 4.9 | Yes |
| 549594 | 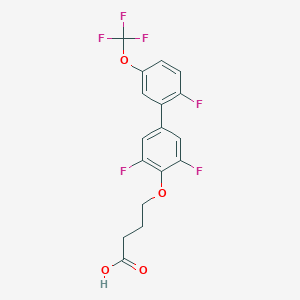 CHEMBL3973101 CHEMBL3973101 | C17H12F6O4 | 394.269 | 10 / 1 | 4.9 | Yes |
| 561494 | 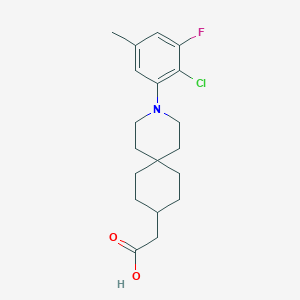 SCHEMBL16483225 SCHEMBL16483225 | C19H25ClFNO2 | 353.862 | 4 / 1 | 5.4 | No |
| 135444 | 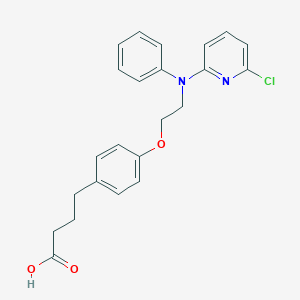 CHEMBL459168 CHEMBL459168 | C23H23ClN2O3 | 410.898 | 5 / 1 | 5.6 | No |
| 525479 | 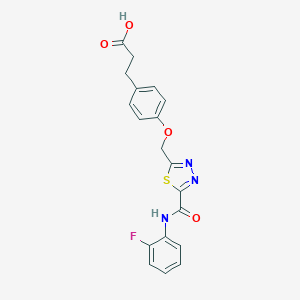 CHEMBL3805888 CHEMBL3805888 | C19H16FN3O4S | 401.412 | 8 / 2 | 2.9 | Yes |
| 561602 | 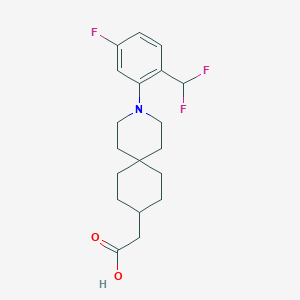 SCHEMBL16482936 SCHEMBL16482936 | C19H24F3NO2 | 355.401 | 6 / 1 | 5.1 | No |
| 141177 | 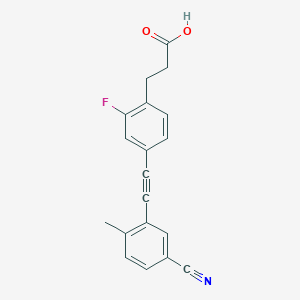 CHEMBL2386363 CHEMBL2386363 | C19H14FNO2 | 307.324 | 4 / 1 | 3.9 | Yes |
| 549687 | 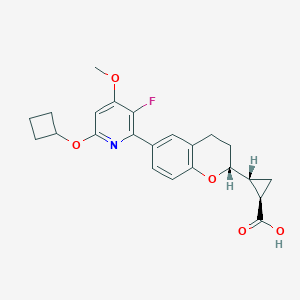 CHEMBL3898962 CHEMBL3898962 | C23H24FNO5 | 413.445 | 7 / 1 | 4.2 | Yes |
| 549697 | 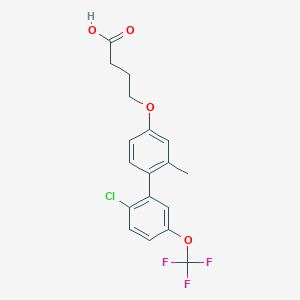 CHEMBL3961009 CHEMBL3961009 | C18H16ClF3O4 | 388.767 | 7 / 1 | 5.5 | No |
| 561778 | 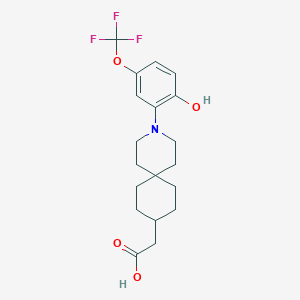 SCHEMBL16483118 SCHEMBL16483118 | C19H24F3NO4 | 387.399 | 8 / 2 | 5.2 | No |
| 561779 | 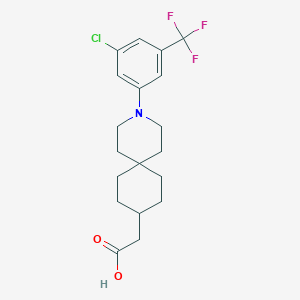 SCHEMBL16483326 SCHEMBL16483326 | C19H23ClF3NO2 | 389.843 | 6 / 1 | 5.9 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218