

You can:
| Name | Glucagon receptor |
|---|---|
| Species | Mus musculus (Mouse) |
| Gene | Gcgr |
| Synonym | GGR GL-R glucagon receptor GR |
| Disease | N/A for non-human GPCRs |
| Length | 485 |
| Amino acid sequence | MPLTQLHCPHLLLLLLVLSCLPEAPSAQVMDFLFEKWKLYSDQCHHNLSLLPPPTELVCNRTFDKYSCWPDTPPNTTANISCPWYLPWYHKVQHRLVFKRCGPDGQWVRGPRGQPWRNASQCQLDDEEIEVQKGVAKMYSSQQVMYTVGYSLSLGALLLALVILLGLRKLHCTRNYIHGNLFASFVLKAGSVLVIDWLLKTRYSQKIGDDLSVSVWLSDGAMAGCRVATVIMQYGIIANYCWLLVEGVYLYSLLSLATFSERSFFSLYLGIGWGAPLLFVIPWVVVKCLFENVQCWTSNDNMGFWWILRIPVFLALLINFFIFVHIIHLLVAKLRAHQMHYADYKFRLARSTLTLIPLLGVHEVVFAFVTDEHAQGTLRSTKLFFDLFLSSFQGLLVAVLYCFLNKEVQAELMRRWRQWQEGKALQEERLASSHGSHMAPAGPCHGDPCEKLQLMSAGSSSGTGCVPSMETSLASSLPRLADSPT |
| UniProt | Q61606 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL4773 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 31167 | 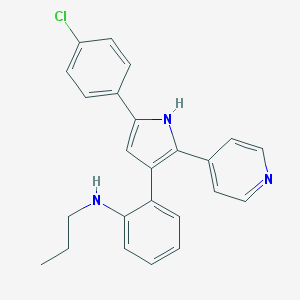 CHEMBL348555 CHEMBL348555 | C24H22ClN3 | 387.911 | 2 / 2 | 6.1 | No |
| 50145 | 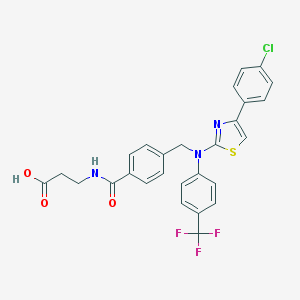 643011-22-7 643011-22-7 | C27H21ClF3N3O3S | 559.988 | 9 / 2 | 6.4 | No |
| 62741 | 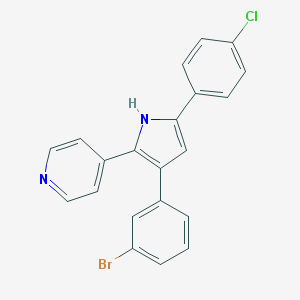 CHEMBL350255 CHEMBL350255 | C21H14BrClN2 | 409.711 | 1 / 1 | 5.9 | No |
| 64646 | 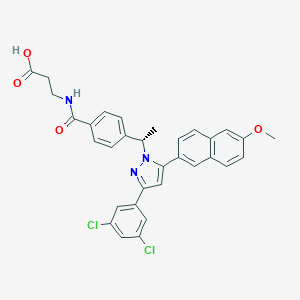 MK 0893 MK 0893 | C32H27Cl2N3O4 | 588.485 | 5 / 2 | 6.8 | No |
| 77349 | 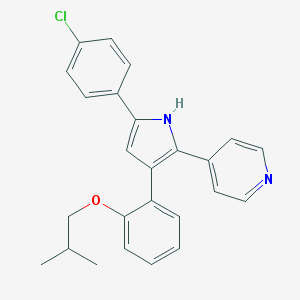 CHEMBL347663 CHEMBL347663 | C25H23ClN2O | 402.922 | 2 / 1 | 6.5 | No |
| 95077 | 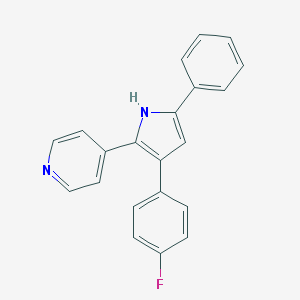 CHEMBL158671 CHEMBL158671 | C21H15FN2 | 314.363 | 2 / 1 | 4.7 | Yes |
| 114786 | 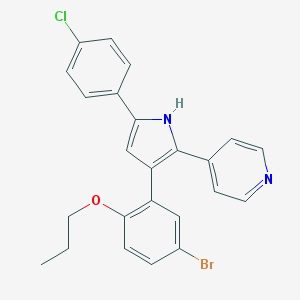 L-168,049 L-168,049 | C24H20BrClN2O | 467.791 | 2 / 1 | 6.8 | No |
| 115016 | 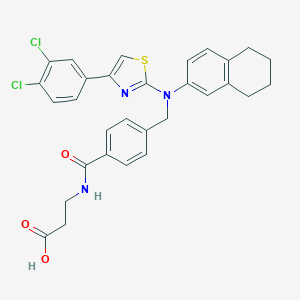 aminothiazole, 10 aminothiazole, 10 | C30H27Cl2N3O3S | 580.524 | 6 / 2 | 7.5 | No |
| 129012 | 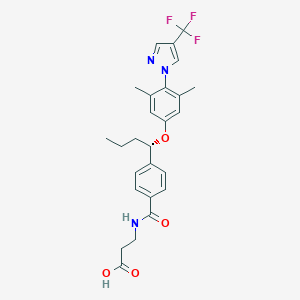 glucagon receptor antagonists-4 glucagon receptor antagonists-4 | C26H28F3N3O4 | 503.522 | 8 / 2 | 5.0 | No |
| 132424 | 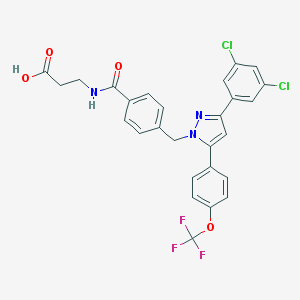 CHEMBL1644183 CHEMBL1644183 | C27H20Cl2F3N3O4 | 578.369 | 8 / 2 | 6.4 | No |
| 151872 | 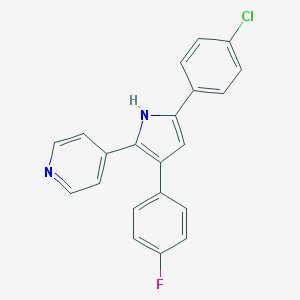 CHEMBL328126 CHEMBL328126 | C21H14ClFN2 | 348.805 | 2 / 1 | 5.3 | No |
| 174425 | 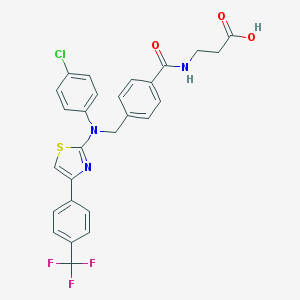 aminothiazole, 16 aminothiazole, 16 | C27H21ClF3N3O3S | 559.988 | 9 / 2 | 6.4 | No |
| 179331 | 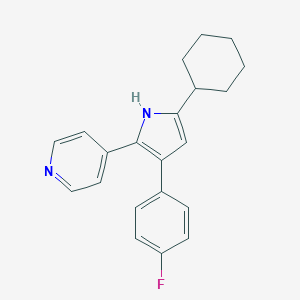 CHEMBL348401 CHEMBL348401 | C21H21FN2 | 320.411 | 2 / 1 | 5.2 | No |
| 182887 | 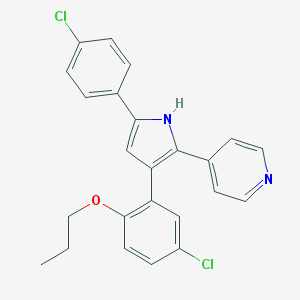 CHEMBL155686 CHEMBL155686 | C24H20Cl2N2O | 423.337 | 2 / 1 | 6.7 | No |
| 202842 | 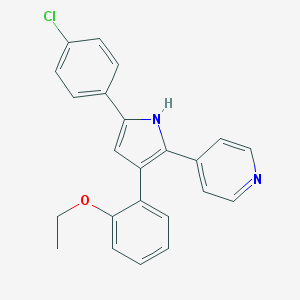 CHEMBL346698 CHEMBL346698 | C23H19ClN2O | 374.868 | 2 / 1 | 5.6 | No |
| 210352 | 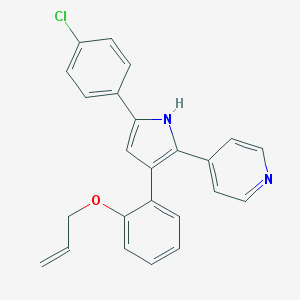 CHEMBL158535 CHEMBL158535 | C24H19ClN2O | 386.879 | 2 / 1 | 5.8 | No |
| 216743 | 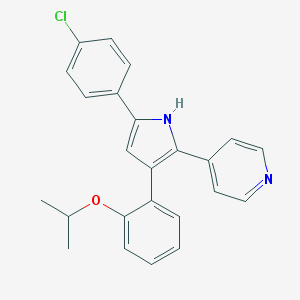 CHEMBL160561 CHEMBL160561 | C24H21ClN2O | 388.895 | 2 / 1 | 6.0 | No |
| 232570 | 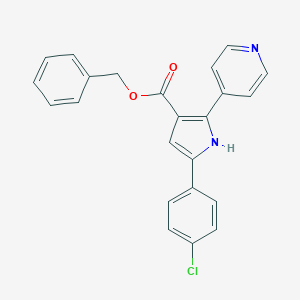 CHEMBL351104 CHEMBL351104 | C23H17ClN2O2 | 388.851 | 3 / 1 | 4.9 | Yes |
| 233418 | 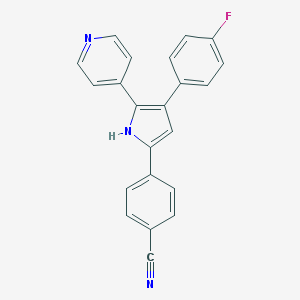 CHEMBL158989 CHEMBL158989 | C22H14FN3 | 339.373 | 3 / 1 | 4.4 | Yes |
| 238453 | 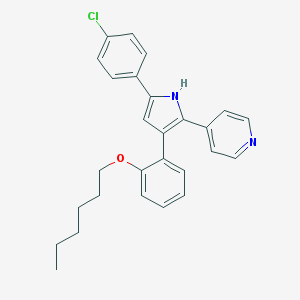 CHEMBL345834 CHEMBL345834 | C27H27ClN2O | 430.976 | 2 / 1 | 7.5 | No |
| 271926 | 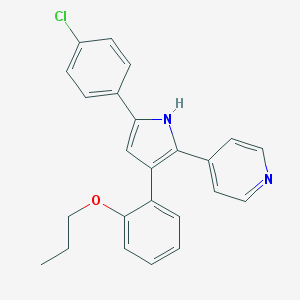 CHEMBL156458 CHEMBL156458 | C24H21ClN2O | 388.895 | 2 / 1 | 6.1 | No |
| 286050 | 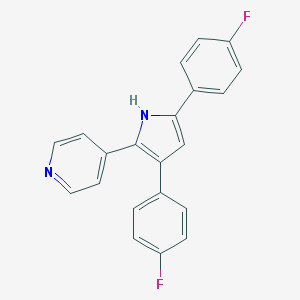 CHEMBL157529 CHEMBL157529 | C21H14F2N2 | 332.354 | 3 / 1 | 4.8 | Yes |
| 289599 | 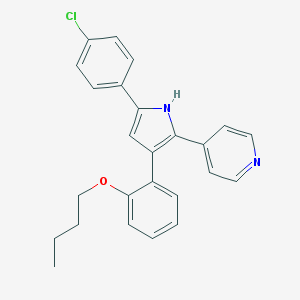 CHEMBL158676 CHEMBL158676 | C25H23ClN2O | 402.922 | 2 / 1 | 6.4 | No |
| 308065 | 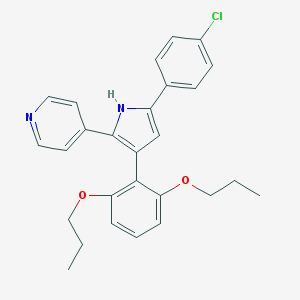 CHEMBL347462 CHEMBL347462 | C27H27ClN2O2 | 446.975 | 3 / 1 | 6.9 | No |
| 317401 | 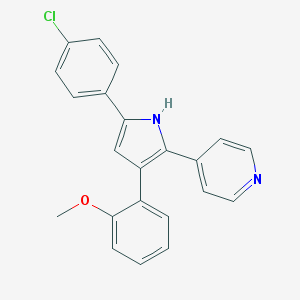 CHEMBL160380 CHEMBL160380 | C22H17ClN2O | 360.841 | 2 / 1 | 5.2 | No |
| 324258 | 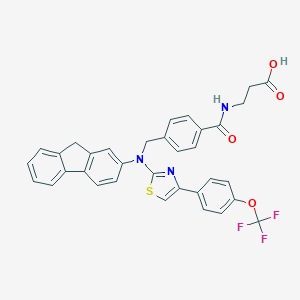 aminothiazole, 6 aminothiazole, 6 | C34H26F3N3O4S | 629.654 | 10 / 2 | 7.7 | No |
| 324917 | 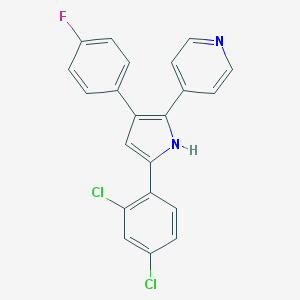 CHEMBL160638 CHEMBL160638 | C21H13Cl2FN2 | 383.247 | 2 / 1 | 5.9 | No |
| 345680 | 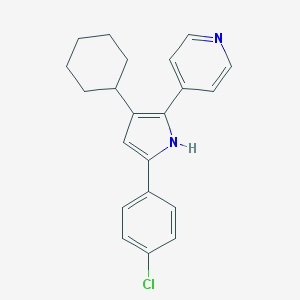 CHEMBL160735 CHEMBL160735 | C21H21ClN2 | 336.863 | 1 / 1 | 6.1 | No |
| 352494 | 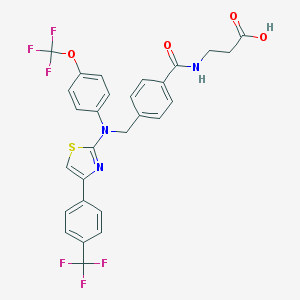 aminothiazole, 19 aminothiazole, 19 | C28H21F6N3O4S | 609.543 | 13 / 2 | 7.0 | No |
| 372260 | 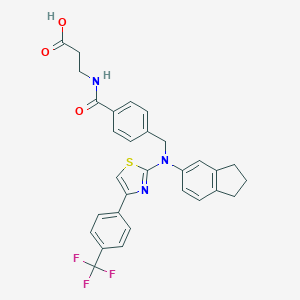 aminothiazole, 7 aminothiazole, 7 | C30H26F3N3O3S | 565.611 | 9 / 2 | 6.6 | No |
| 378633 | 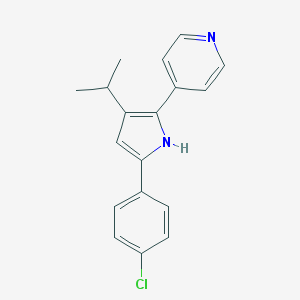 CHEMBL158548 CHEMBL158548 | C18H17ClN2 | 296.798 | 1 / 1 | 4.7 | Yes |
| 389205 | 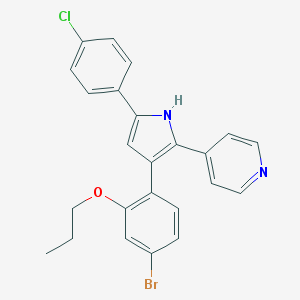 CHEMBL160287 CHEMBL160287 | C24H20BrClN2O | 467.791 | 2 / 1 | 6.8 | No |
| 389545 | 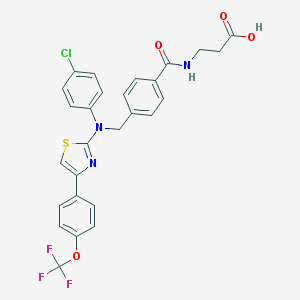 aminothiazole, 17 aminothiazole, 17 | C27H21ClF3N3O4S | 575.987 | 10 / 2 | 6.7 | No |
| 393869 | 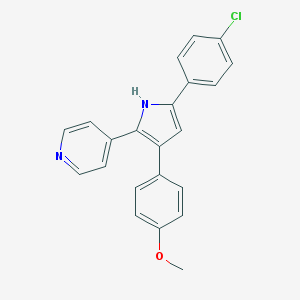 CHEMBL155694 CHEMBL155694 | C22H17ClN2O | 360.841 | 2 / 1 | 5.2 | No |
| 398107 | 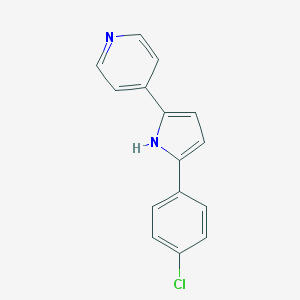 CHEMBL160372 CHEMBL160372 | C15H11ClN2 | 254.717 | 1 / 1 | 3.6 | Yes |
| 401176 | 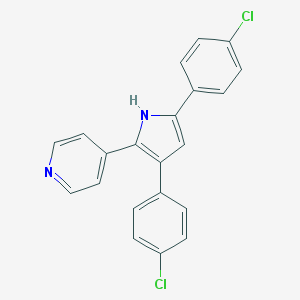 CHEMBL160490 CHEMBL160490 | C21H14Cl2N2 | 365.257 | 1 / 1 | 5.8 | No |
| 415232 | 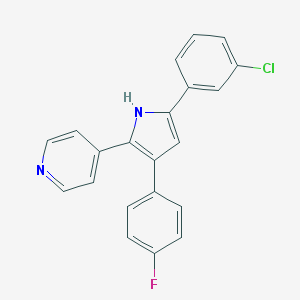 CHEMBL160281 CHEMBL160281 | C21H14ClFN2 | 348.805 | 2 / 1 | 5.3 | No |
| 421525 | 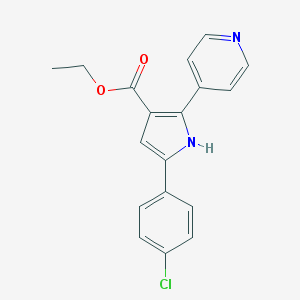 CHEMBL345863 CHEMBL345863 | C18H15ClN2O2 | 326.78 | 3 / 1 | 3.8 | Yes |
| 423283 | 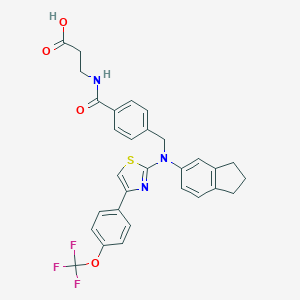 aminothiazole, 8 aminothiazole, 8 | C30H26F3N3O4S | 581.61 | 10 / 2 | 6.9 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218