

You can:
| Name | D(1A) dopamine receptor |
|---|---|
| Species | Mus musculus (Mouse) |
| Gene | Drd1 |
| Synonym | D1 receptor D1A DADR Dopamine D1 receptor Dopamine-1A receptor [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 446 |
| Amino acid sequence | MAPNTSTMDETGLPVERDFSFRILTACFLSLLILSTLLGNTLVCAAVIRFRHLRSKVTNFFVISLAVSDLLVAVLVMPWKAVAEIAGFWPFGSFCNIWVAFDIMCSTASILNLCVISVDRYWAISSPFQYERKMTPKAAFILISVAWTLSVLISFIPVQLSWHKAKPTWPLDGNFTSLEDAEDDNCDTRLSRTYAISSSLISFYIPVAIMIVTYTSIYRIAQKQIRRISALERAAVHAKNCQTTTGNGNPVECSQSESSFKMSFKRETKVLKTLSVIMGVFVCCWLPFFISNCMVPFCGSEETQPFCIDSITFDVFVWFGWANSSLNPIIYAFNADFQKAFSTLLGCYRLCPTTNNAIETVSINNNGAVMFSSHHEPRGSISKDCNLVYLIPHAVGSSEDLKREEAGGIPKPLEKLSPALSVILDYDTDVSLEKIQPVTHSGQHST |
| UniProt | Q61616 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3071 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 10450 | 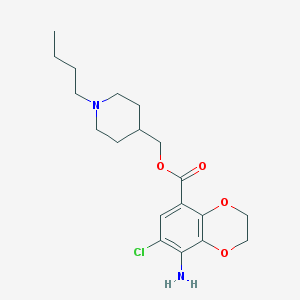 SB 204070 hydrochloride SB 204070 hydrochloride | C19H27ClN2O4 | 382.885 | 6 / 1 | 3.5 | Yes |
| 11787 | 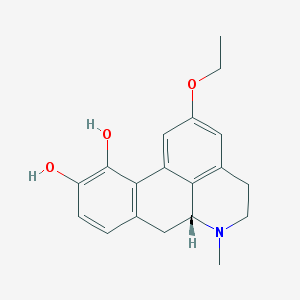 CHEMBL465055 CHEMBL465055 | C19H21NO3 | 311.381 | 4 / 2 | 2.6 | Yes |
| 13673 | 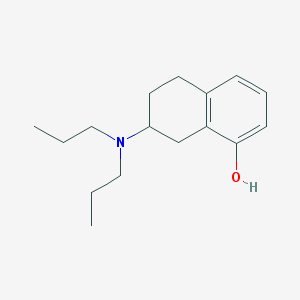 8-OH-Dpat 8-OH-Dpat | C16H25NO | 247.382 | 2 / 1 | 4.1 | Yes |
| 19664 | 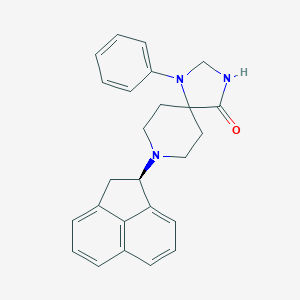 CHEMBL355202 CHEMBL355202 | C25H25N3O | 383.495 | 3 / 1 | 4.5 | Yes |
| 20722 | 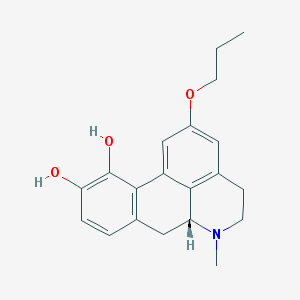 CHEMBL517491 CHEMBL517491 | C20H23NO3 | 325.408 | 4 / 2 | 3.2 | Yes |
| 23334 | 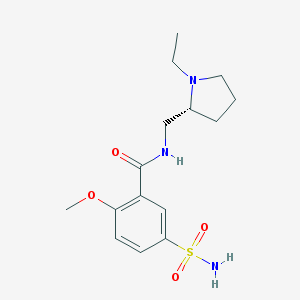 (+)-sulpiride (+)-sulpiride | C15H23N3O4S | 341.426 | 6 / 2 | 0.6 | Yes |
| 23344 | 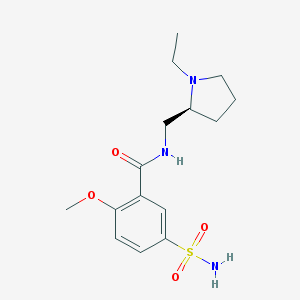 Levosulpiride Levosulpiride | C15H23N3O4S | 341.426 | 6 / 2 | 0.6 | Yes |
| 23362 | 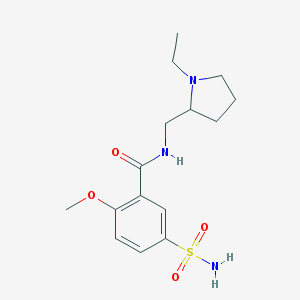 sulpiride sulpiride | C15H23N3O4S | 341.426 | 6 / 2 | 0.6 | Yes |
| 33960 | 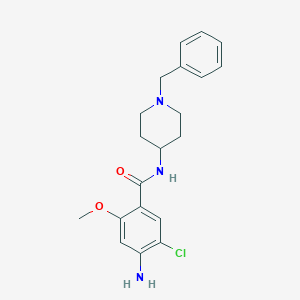 Clebopride Clebopride | C20H24ClN3O2 | 373.881 | 4 / 2 | 3.2 | Yes |
| 35762 | 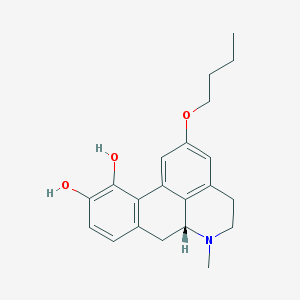 CHEMBL449461 CHEMBL449461 | C21H25NO3 | 339.435 | 4 / 2 | 3.5 | Yes |
| 558728 | 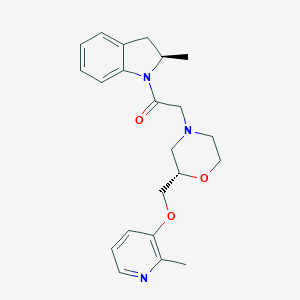 SCHEMBL15746093 SCHEMBL15746093 | C22H27N3O3 | 381.476 | 5 / 0 | 2.5 | Yes |
| 537188 | 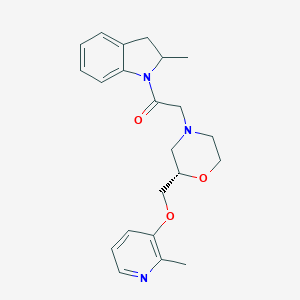 CHEMBL3941579 CHEMBL3941579 | C22H27N3O3 | 381.476 | 5 / 0 | 2.5 | Yes |
| 548428 | 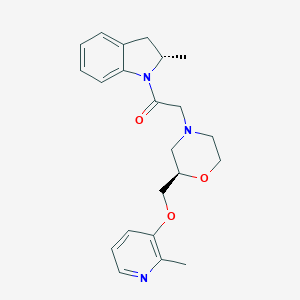 CHEMBL3956749 CHEMBL3956749 | C22H27N3O3 | 381.476 | 5 / 0 | 2.5 | Yes |
| 62248 | 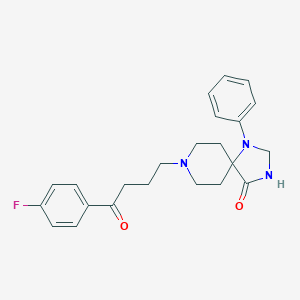 spiperone spiperone | C23H26FN3O2 | 395.478 | 5 / 1 | 3.0 | Yes |
| 69158 | 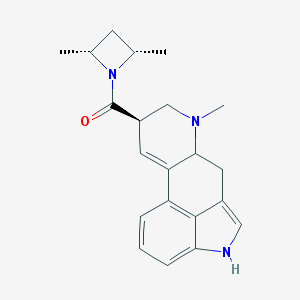 CHEMBL137781 CHEMBL137781 | C21H25N3O | 335.451 | 2 / 1 | 3.2 | Yes |
| 69173 | 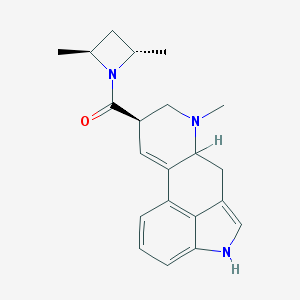 CHEMBL343755 CHEMBL343755 | C21H25N3O | 335.451 | 2 / 1 | 3.2 | Yes |
| 69182 | 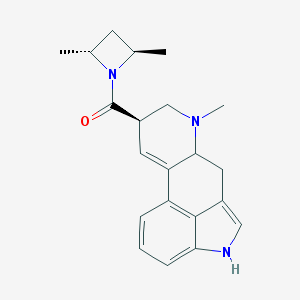 CHEMBL138989 CHEMBL138989 | C21H25N3O | 335.451 | 2 / 1 | 3.2 | Yes |
| 80078 | 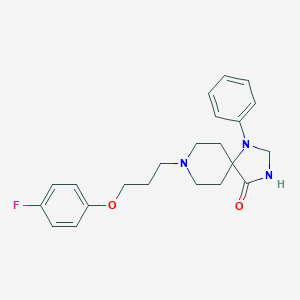 Spiramide Spiramide | C22H26FN3O2 | 383.467 | 5 / 1 | 3.4 | Yes |
| 83990 | 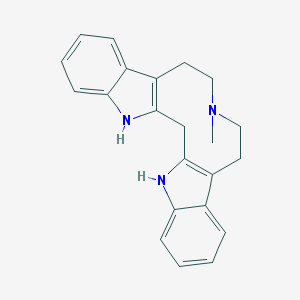 CHEMBL62529 CHEMBL62529 | C22H23N3 | 329.447 | 1 / 2 | 4.5 | Yes |
| 87048 | 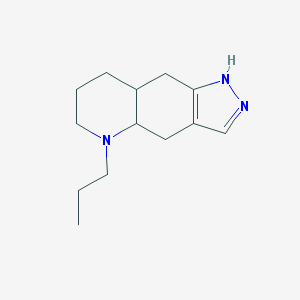 (A+/-)-Quinpirole dihydrochloride (A+/-)-Quinpirole dihydrochloride | C13H21N3 | 219.332 | 2 / 1 | 2.3 | Yes |
| 98029 | 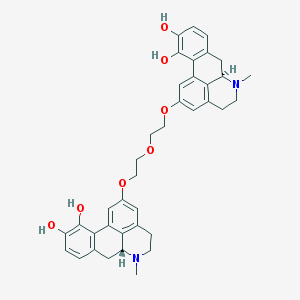 CHEMBL444149 CHEMBL444149 | C38H40N2O7 | 636.745 | 9 / 4 | 5.1 | No |
| 101531 | 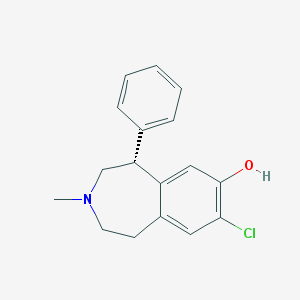 UNII-UGT5535REQ UNII-UGT5535REQ | C17H18ClNO | 287.787 | 2 / 1 | 4.0 | Yes |
| 119904 | 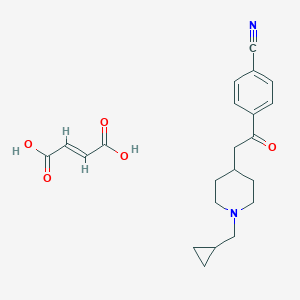 CHEMBL137776 CHEMBL137776 | C22H26N2O5 | 398.459 | 7 / 2 | N/A | N/A |
| 124416 | 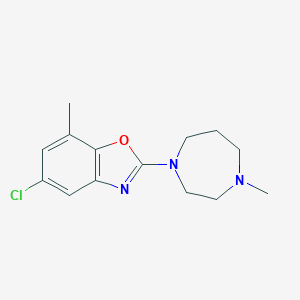 CHEMBL104985 CHEMBL104985 | C14H18ClN3O | 279.768 | 4 / 0 | 3.4 | Yes |
| 134694 | 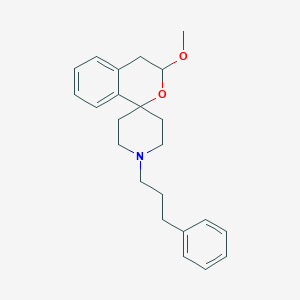 CHEMBL138458 CHEMBL138458 | C23H29NO2 | 351.49 | 3 / 0 | 4.3 | Yes |
| 136082 | 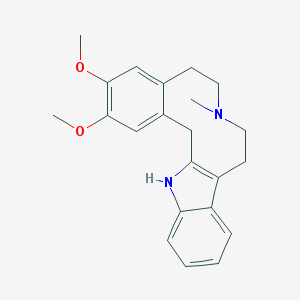 CHEMBL63063 CHEMBL63063 | C22H26N2O2 | 350.462 | 3 / 1 | 4.3 | Yes |
| 160024 | 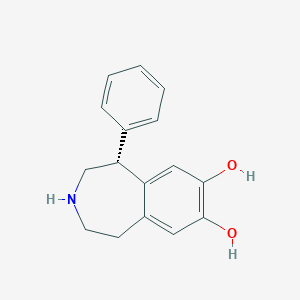 UNII-YZC2Y51NHJ UNII-YZC2Y51NHJ | C16H17NO2 | 255.317 | 3 / 3 | 2.5 | Yes |
| 162667 | 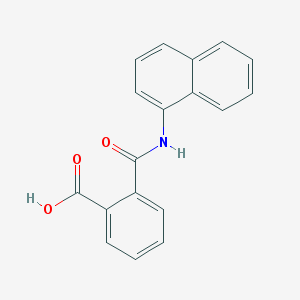 NAPTALAM NAPTALAM | C18H13NO3 | 291.306 | 3 / 2 | 3.5 | Yes |
| 172218 | 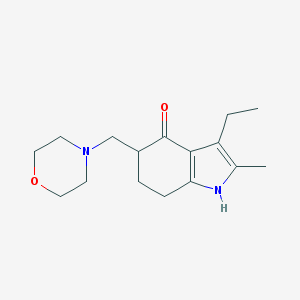 MOLINDONE MOLINDONE | C16H24N2O2 | 276.38 | 3 / 1 | 1.7 | Yes |
| 179916 | 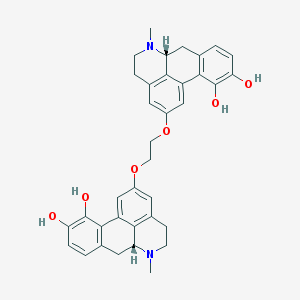 CHEMBL442926 CHEMBL442926 | C36H36N2O6 | 592.692 | 8 / 4 | 4.8 | No |
| 180085 | 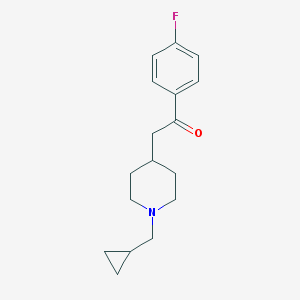 DUP-734 free base DUP-734 free base | C17H22FNO | 275.367 | 3 / 0 | 3.2 | Yes |
| 188203 | 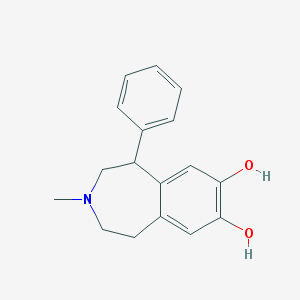 SKF-75670 SKF-75670 | C17H19NO2 | 269.344 | 3 / 2 | 3.0 | Yes |
| 191551 | 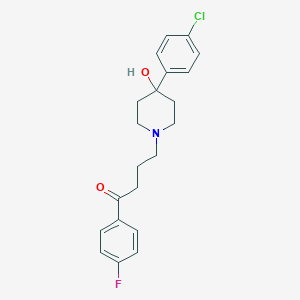 haloperidol haloperidol | C21H23ClFNO2 | 375.868 | 4 / 1 | 3.2 | Yes |
| 196621 | 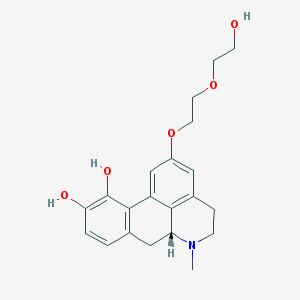 CHEMBL516151 CHEMBL516151 | C21H25NO5 | 371.433 | 6 / 3 | 1.4 | Yes |
| 208627 | 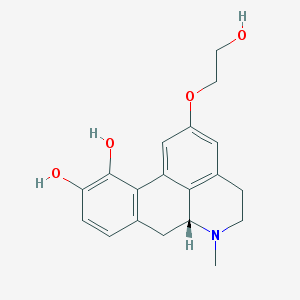 CHEMBL457275 CHEMBL457275 | C19H21NO4 | 327.38 | 5 / 3 | 1.6 | Yes |
| 224934 | 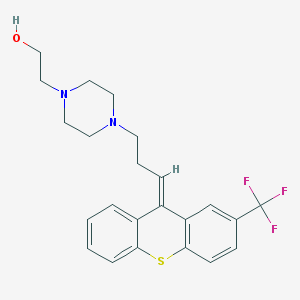 (E)-Flupenthixol (E)-Flupenthixol | C23H25F3N2OS | 434.521 | 7 / 1 | 4.5 | Yes |
| 542147 | 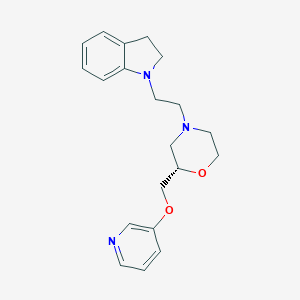 CHEMBL3947839 CHEMBL3947839 | C20H25N3O2 | 339.439 | 5 / 0 | 2.4 | Yes |
| 564615 | 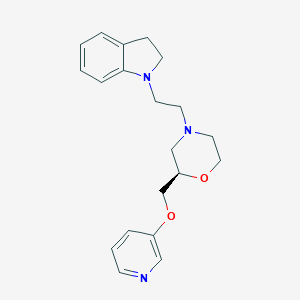 SCHEMBL14149778 SCHEMBL14149778 | C20H25N3O2 | 339.439 | 5 / 0 | 2.4 | Yes |
| 237435 | 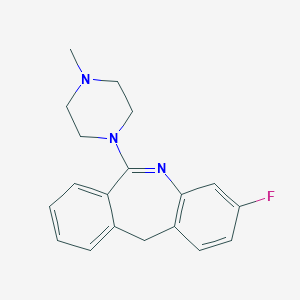 FLUPERLAPINE FLUPERLAPINE | C19H20FN3 | 309.388 | 3 / 0 | 2.9 | Yes |
| 244524 | 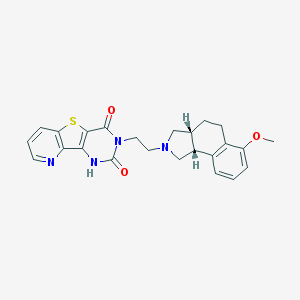 A-131701 A-131701 | C24H24N4O3S | 448.541 | 6 / 1 | 3.2 | Yes |
| 258011 | 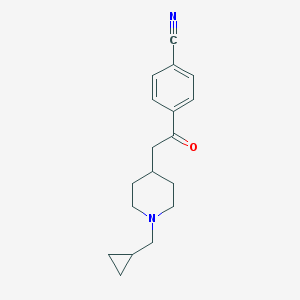 CHEMBL130356 CHEMBL130356 | C18H22N2O | 282.387 | 3 / 0 | 2.8 | Yes |
| 262517 | 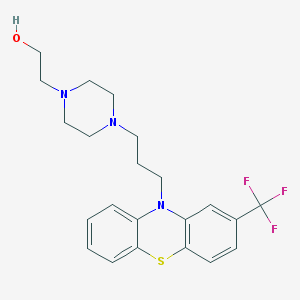 FLUPHENAZINE FLUPHENAZINE | C22H26F3N3OS | 437.525 | 8 / 1 | 4.4 | Yes |
| 264917 | 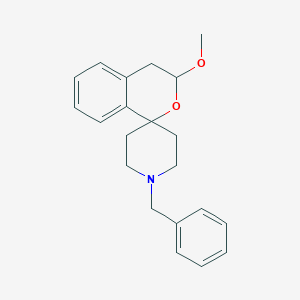 CHEMBL141209 CHEMBL141209 | C21H25NO2 | 323.436 | 3 / 0 | 3.5 | Yes |
| 543340 | 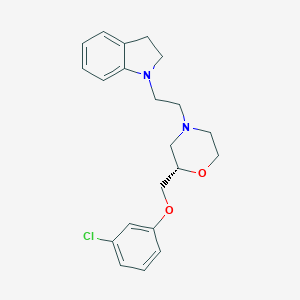 CHEMBL3946540 CHEMBL3946540 | C21H25ClN2O2 | 372.893 | 4 / 0 | 4.1 | Yes |
| 565814 | 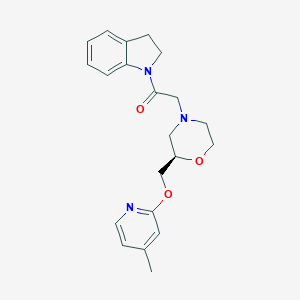 SCHEMBL14061586 SCHEMBL14061586 | C21H25N3O3 | 367.449 | 5 / 0 | 2.4 | Yes |
| 551320 | 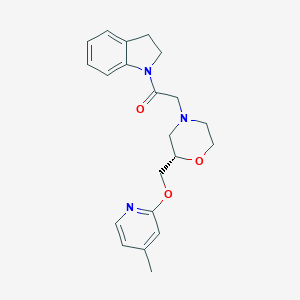 CHEMBL3970895 CHEMBL3970895 | C21H25N3O3 | 367.449 | 5 / 0 | 2.4 | Yes |
| 277636 | 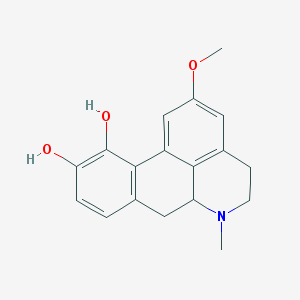 ACMC-20dim2 ACMC-20dim2 | C18H19NO3 | 297.354 | 4 / 2 | 2.3 | Yes |
| 281540 | 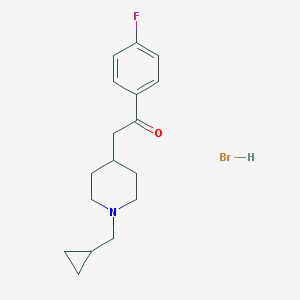 DuP 734 DuP 734 | C17H23BrFNO | 356.279 | 3 / 1 | N/A | N/A |
| 543761 | 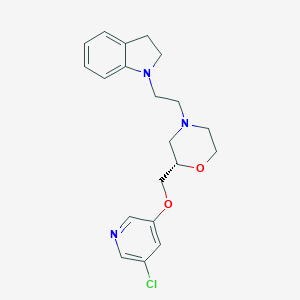 CHEMBL3963122 CHEMBL3963122 | C20H24ClN3O2 | 373.881 | 5 / 0 | 3.0 | Yes |
| 288329 | 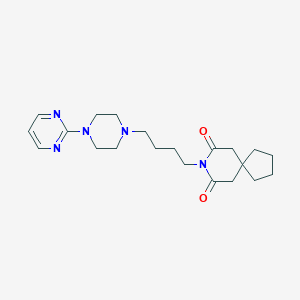 buspirone buspirone | C21H31N5O2 | 385.512 | 6 / 0 | 2.6 | Yes |
| 566326 | 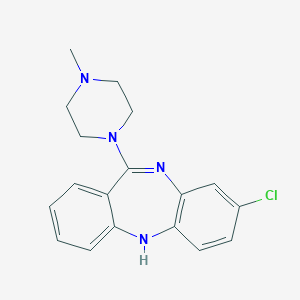 clozapine clozapine | C18H19ClN4 | 326.828 | 3 / 1 | 3.1 | Yes |
| 291132 | 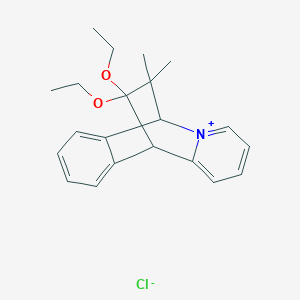 CHEMBL52124 CHEMBL52124 | C21H26ClNO2 | 359.894 | 3 / 0 | N/A | N/A |
| 544115 | 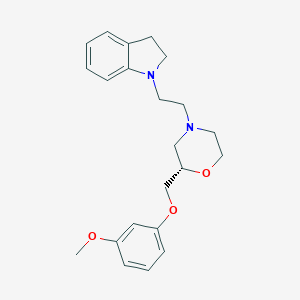 CHEMBL3986651 CHEMBL3986651 | C22H28N2O3 | 368.477 | 5 / 0 | 3.4 | Yes |
| 309256 | 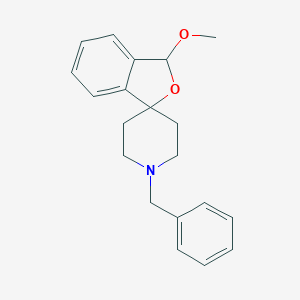 CHEMBL138809 CHEMBL138809 | C20H23NO2 | 309.409 | 3 / 0 | 3.0 | Yes |
| 311830 | 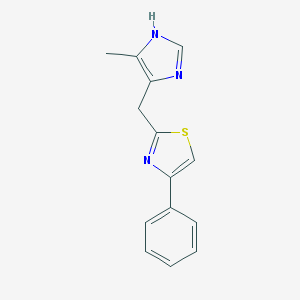 CHEMBL38465 CHEMBL38465 | C14H13N3S | 255.339 | 3 / 1 | 3.1 | Yes |
| 316286 | 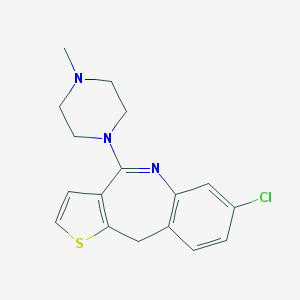 Tilozepine Tilozepine | C17H18ClN3S | 331.862 | 3 / 0 | 3.2 | Yes |
| 345204 | 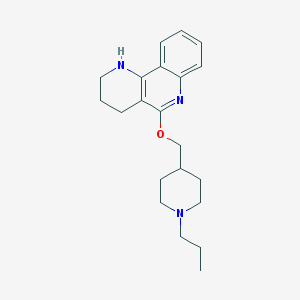 CHEMBL35226 CHEMBL35226 | C21H29N3O | 339.483 | 4 / 1 | 4.5 | Yes |
| 348829 | 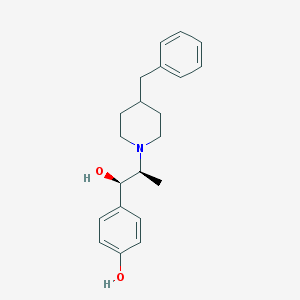 CHEMBL49623 CHEMBL49623 | C21H27NO2 | 325.452 | 3 / 2 | 3.9 | Yes |
| 348839 | 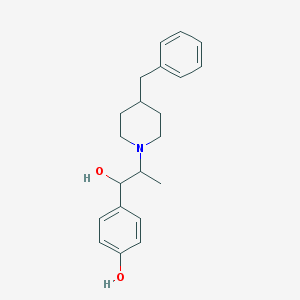 Ifenprodil Ifenprodil | C21H27NO2 | 325.452 | 3 / 2 | 3.9 | Yes |
| 350540 | 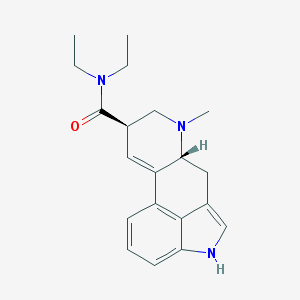 Lysergide Lysergide | C20H25N3O | 323.44 | 2 / 1 | 3.0 | Yes |
| 359019 | 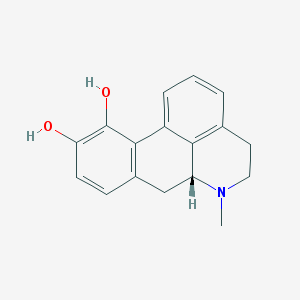 apomorphine apomorphine | C17H17NO2 | 267.328 | 3 / 2 | 2.3 | Yes |
| 368747 | 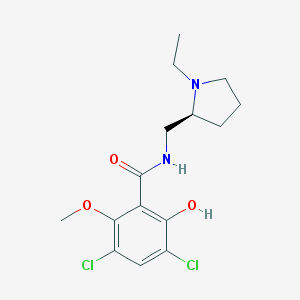 RACLOPRIDE RACLOPRIDE | C15H20Cl2N2O3 | 347.236 | 4 / 2 | 2.9 | Yes |
| 552689 | 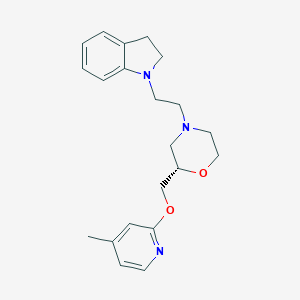 CHEMBL3941323 CHEMBL3941323 | C21H27N3O2 | 353.466 | 5 / 0 | 3.1 | Yes |
| 569390 | 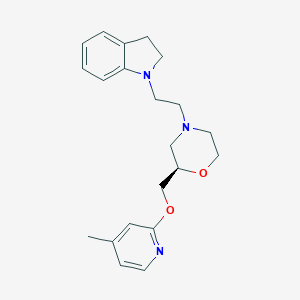 SCHEMBL14061568 SCHEMBL14061568 | C21H27N3O2 | 353.466 | 5 / 0 | 3.1 | Yes |
| 393307 | 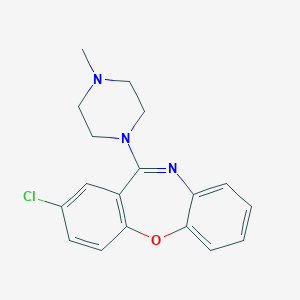 loxapine loxapine | C18H18ClN3O | 327.812 | 3 / 0 | 3.1 | Yes |
| 395991 | 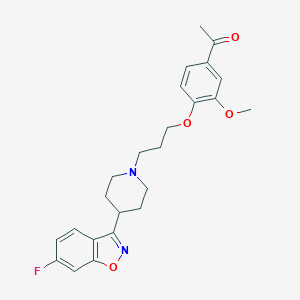 Iloperidone Iloperidone | C24H27FN2O4 | 426.488 | 7 / 0 | 4.1 | Yes |
| 546853 | 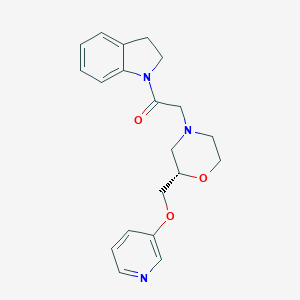 CHEMBL3936595 CHEMBL3936595 | C20H23N3O3 | 353.422 | 5 / 0 | 1.7 | Yes |
| 408085 | 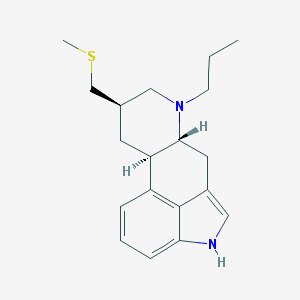 pergolide pergolide | C19H26N2S | 314.491 | 2 / 1 | 4.2 | Yes |
| 408440 | 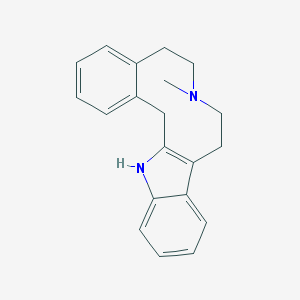 LE 300 LE 300 | C20H22N2 | 290.41 | 1 / 1 | 4.3 | Yes |
| 553113 | 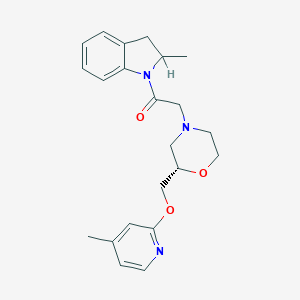 CHEMBL3963147 CHEMBL3963147 | C22H27N3O3 | 381.476 | 5 / 0 | 2.8 | Yes |
| 570601 | 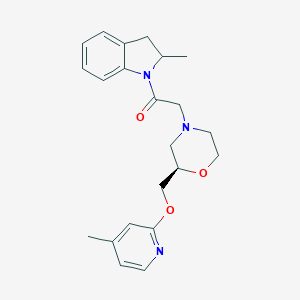 SCHEMBL15746102 SCHEMBL15746102 | C22H27N3O3 | 381.476 | 5 / 0 | 2.8 | Yes |
| 547590 | 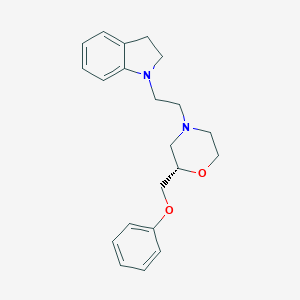 CHEMBL3946661 CHEMBL3946661 | C21H26N2O2 | 338.451 | 4 / 0 | 3.5 | Yes |
| 434066 | 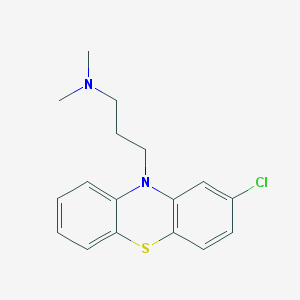 chlorpromazine chlorpromazine | C17H19ClN2S | 318.863 | 3 / 0 | 5.2 | No |
| 441221 | 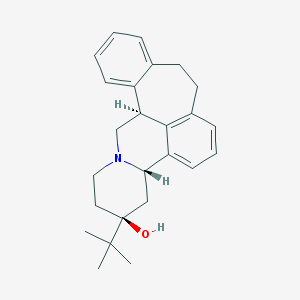 (+)-Butaclamol (+)-Butaclamol | C25H31NO | 361.529 | 2 / 1 | 5.0 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218