

You can:
| Name | Adenosine receptor A3 |
|---|---|
| Species | Mus musculus (Mouse) |
| Gene | Adora3 |
| Synonym | A3 receptor A3AR Adenosine receptor A3 ARA3 TGPCR1 |
| Disease | N/A for non-human GPCRs |
| Length | 319 |
| Amino acid sequence | MEADNTTETDWLNITYITMEAAIGLCAVVGNMLVIWVVKLNPTLRTTTVYFIVSLALADIAVGVLVIPLAIAVSLQVKMHFYACLFMSCVLLIFTHASIMSLLAIAVHRYLRVKLTVRYRTVTTQRRIWLFLGLCWLVSFLVGLTPMFGWNRKATLASSQNSSTLLCHFRSVVSLDYMVFFSFITWILVPLVVMCIIYLDIFYIIRNKLSQNLTGFRETRAFYGREFKTAKSLFLVLFLFALCWLPLSIINFVSYFDVKIPDVAMCLGILLSHANSMMNPIVYACKIKKFKETYFLILRAVRLCQTSDSLDSNMEQTTE |
| UniProt | Q61618 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL1075269 |
| IUPHAR | 21 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 1200 | 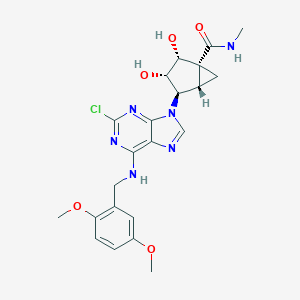 CHEMBL178846 CHEMBL178846 | C22H25ClN6O5 | 488.929 | 9 / 4 | 1.3 | Yes |
| 4749 | 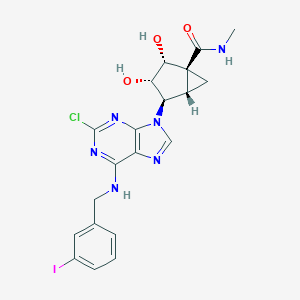 CHEMBL27376 CHEMBL27376 | C20H20ClIN6O3 | 554.773 | 7 / 4 | 2.0 | No |
| 9822 | 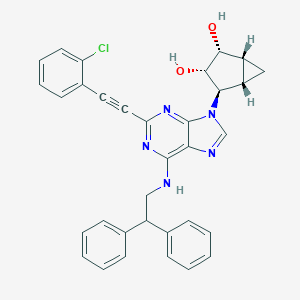 CHEMBL2089350 CHEMBL2089350 | C33H28ClN5O2 | 562.07 | 6 / 3 | 5.6 | No |
| 29001 | 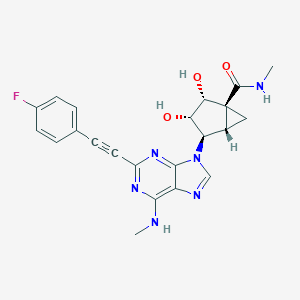 CHEMBL2064639 CHEMBL2064639 | C22H21FN6O3 | 436.447 | 8 / 4 | 1.1 | Yes |
| 443540 | 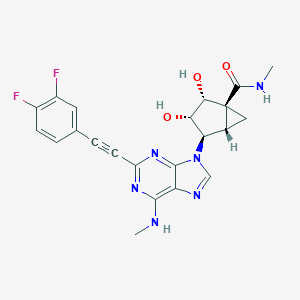 CHEMBL2064645 CHEMBL2064645 | C22H20F2N6O3 | 454.438 | 9 / 4 | 1.2 | Yes |
| 468737 | 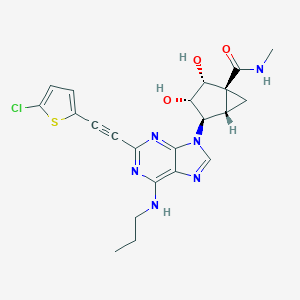 CHEMBL3612932 CHEMBL3612932 | C22H23ClN6O3S | 486.975 | 8 / 4 | 2.5 | Yes |
| 443589 | 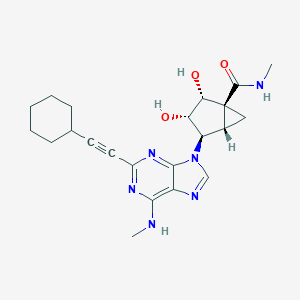 CHEMBL3407788 CHEMBL3407788 | C22H28N6O3 | 424.505 | 7 / 4 | 1.9 | Yes |
| 52516 | 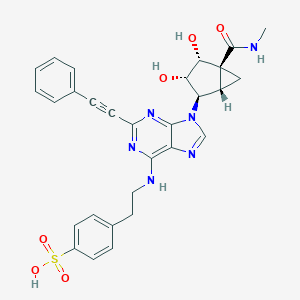 CHEMBL2402025 CHEMBL2402025 | C29H28N6O6S | 588.639 | 10 / 5 | 1.7 | No |
| 53566 | 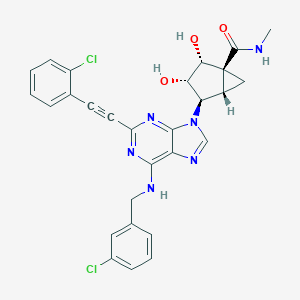 CHEMBL2064655 CHEMBL2064655 | C28H24Cl2N6O3 | 563.439 | 7 / 4 | 3.7 | No |
| 553504 | 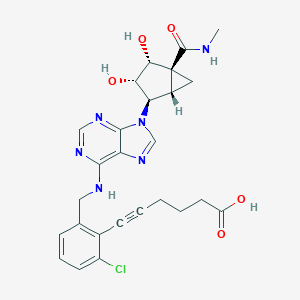 MRS5151 MRS5151 | C26H27ClN6O5 | 538.989 | 9 / 5 | 1.6 | No |
| 60680 | 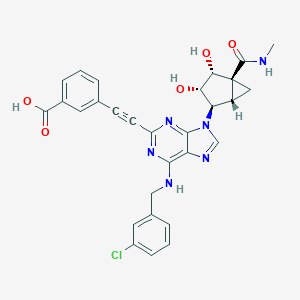 CHEMBL2064659 CHEMBL2064659 | C29H25ClN6O5 | 573.006 | 9 / 5 | 2.6 | No |
| 471403 | 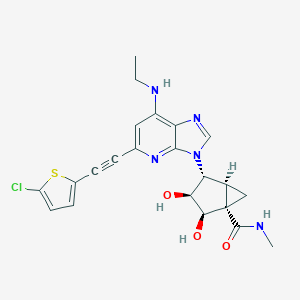 CHEMBL3612940 CHEMBL3612940 | C22H22ClN5O3S | 471.96 | 7 / 4 | 2.3 | Yes |
| 471532 | 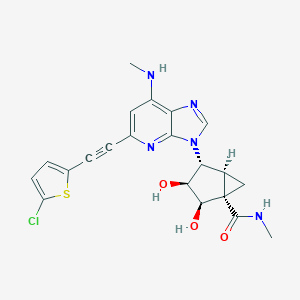 CHEMBL3612939 CHEMBL3612939 | C21H20ClN5O3S | 457.933 | 7 / 4 | 2.0 | Yes |
| 82293 | 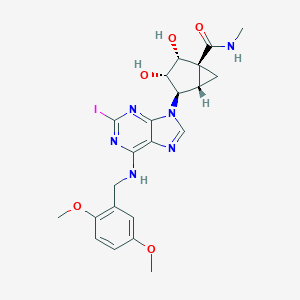 CHEMBL255907 CHEMBL255907 | C22H25IN6O5 | 580.383 | 9 / 4 | 1.0 | No |
| 90123 | 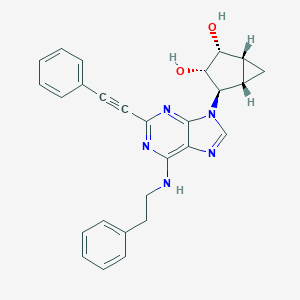 CHEMBL2089346 CHEMBL2089346 | C27H25N5O2 | 451.53 | 6 / 3 | 3.5 | Yes |
| 92067 | 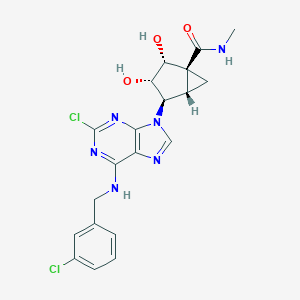 CHEMBL175543 CHEMBL175543 | C20H20Cl2N6O3 | 463.319 | 7 / 4 | 2.0 | Yes |
| 101160 | 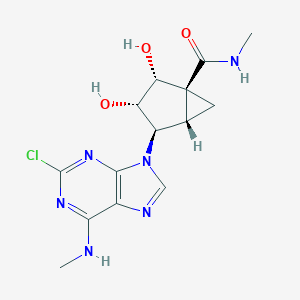 CHEMBL283057 CHEMBL283057 | C14H17ClN6O3 | 352.779 | 7 / 4 | -0.1 | Yes |
| 445708 | 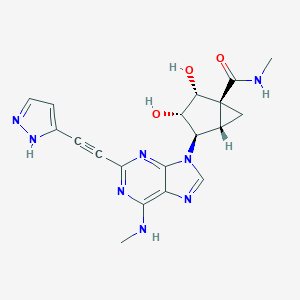 CHEMBL3407776 CHEMBL3407776 | C19H20N8O3 | 408.422 | 8 / 5 | -0.7 | Yes |
| 104062 | 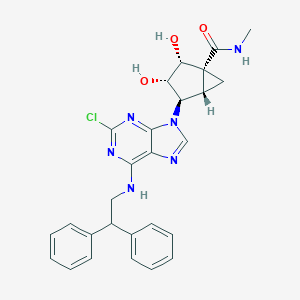 CHEMBL368872 CHEMBL368872 | C27H27ClN6O3 | 519.002 | 7 / 4 | 3.3 | No |
| 446271 | 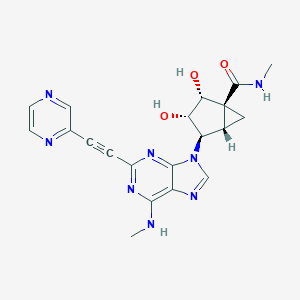 CHEMBL3407774 CHEMBL3407774 | C20H20N8O3 | 420.433 | 9 / 4 | -1.1 | Yes |
| 446400 | 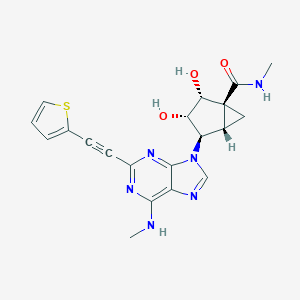 CHEMBL3407783 CHEMBL3407783 | C20H20N6O3S | 424.479 | 8 / 4 | 0.7 | Yes |
| 121529 | 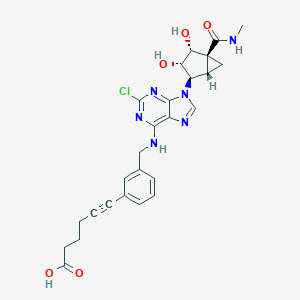 CHEMBL256520 CHEMBL256520 | C26H27ClN6O5 | 538.989 | 9 / 5 | 1.9 | No |
| 123577 | 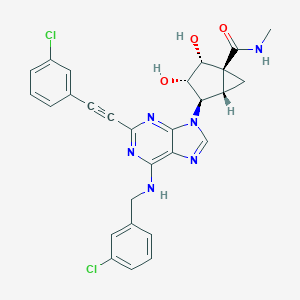 CHEMBL2064656 CHEMBL2064656 | C28H24Cl2N6O3 | 563.439 | 7 / 4 | 3.7 | No |
| 124379 | 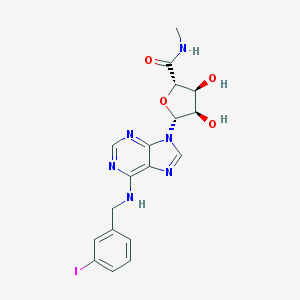 IB-MECA IB-MECA | C18H19IN6O4 | 510.292 | 8 / 4 | 0.9 | No |
| 126967 | 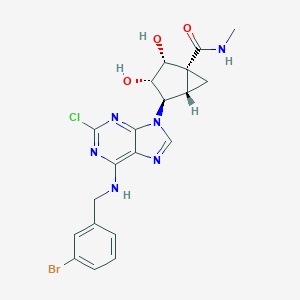 CHEMBL425741 CHEMBL425741 | C20H20BrClN6O3 | 507.773 | 7 / 4 | 2.1 | No |
| 127342 | 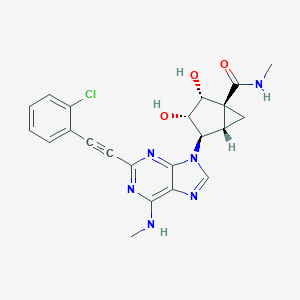 CHEMBL2064640 CHEMBL2064640 | C22H21ClN6O3 | 452.899 | 7 / 4 | 1.6 | Yes |
| 525275 | 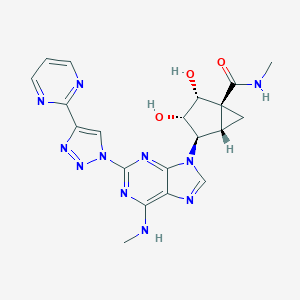 CHEMBL3747670 CHEMBL3747670 | C20H21N11O3 | 463.462 | 11 / 4 | -1.3 | No |
| 130397 | 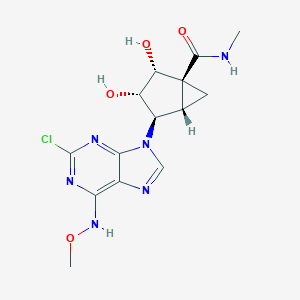 CHEMBL402607 CHEMBL402607 | C14H17ClN6O4 | 368.778 | 8 / 4 | -0.1 | Yes |
| 446928 | 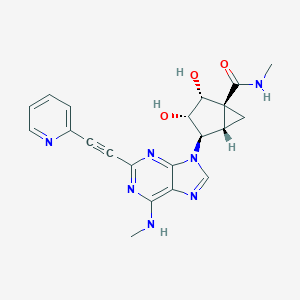 CHEMBL2064636 CHEMBL2064636 | C21H21N7O3 | 419.445 | 8 / 4 | -0.1 | Yes |
| 133229 | 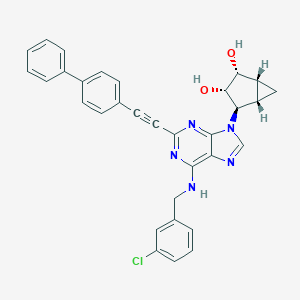 CHEMBL2089345 CHEMBL2089345 | C32H26ClN5O2 | 548.043 | 6 / 3 | 5.3 | No |
| 525382 | 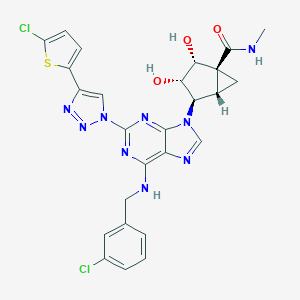 CHEMBL3746886 CHEMBL3746886 | C26H23Cl2N9O3S | 612.49 | 10 / 4 | 3.1 | No |
| 525399 | 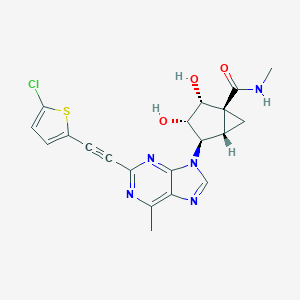 CHEMBL3798200 CHEMBL3798200 | C20H18ClN5O3S | 443.906 | 7 / 3 | 1.7 | Yes |
| 553959 | 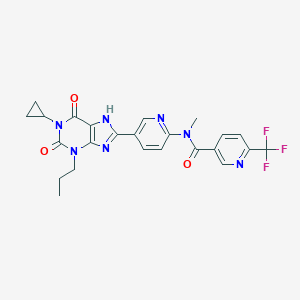 ATL802 ATL802 | C24H22F3N7O3 | 513.481 | 9 / 1 | 2.9 | No |
| 137491 | 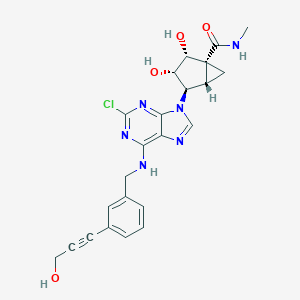 CHEMBL178644 CHEMBL178644 | C23H23ClN6O4 | 482.925 | 8 / 5 | 0.9 | Yes |
| 138946 | 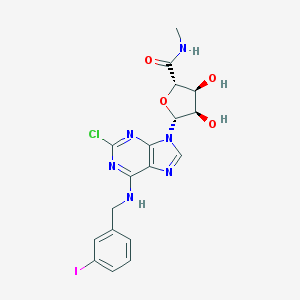 2-Cl-IB-Meca 2-Cl-IB-Meca | C18H18ClIN6O4 | 544.734 | 8 / 4 | 1.9 | No |
| 549649 | 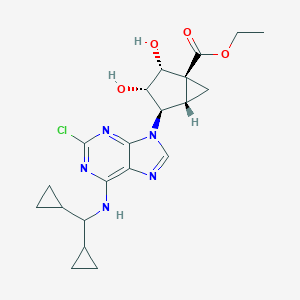 CHEMBL3955519 CHEMBL3955519 | C21H26ClN5O4 | 447.92 | 8 / 3 | 2.4 | Yes |
| 525655 | 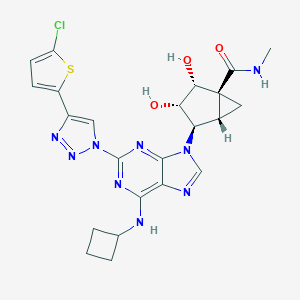 CHEMBL3745851 CHEMBL3745851 | C23H24ClN9O3S | 542.015 | 10 / 4 | 1.9 | No |
| 539713 | 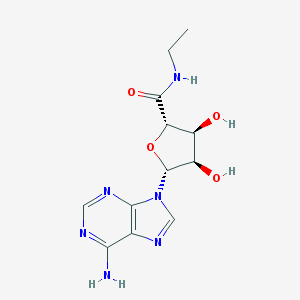 NECA NECA | C12H16N6O4 | 308.298 | 8 / 4 | -0.7 | Yes |
| 447756 | 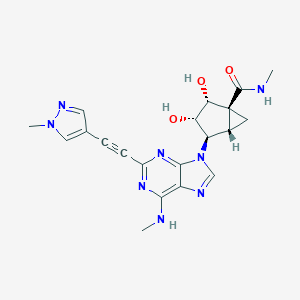 CHEMBL3407777 CHEMBL3407777 | C20H22N8O3 | 422.449 | 8 / 4 | -0.7 | Yes |
| 448302 | 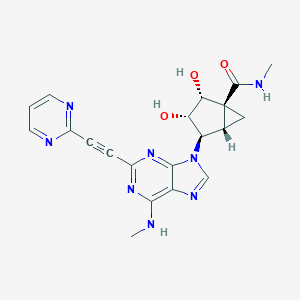 CHEMBL3407773 CHEMBL3407773 | C20H20N8O3 | 420.433 | 9 / 4 | -0.7 | Yes |
| 172068 | 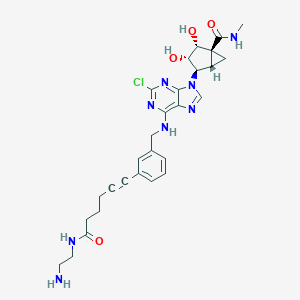 CHEMBL256312 CHEMBL256312 | C28H33ClN8O4 | 581.074 | 9 / 6 | 0.7 | No |
| 550070 | 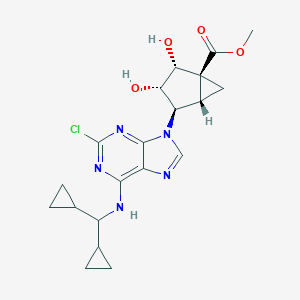 CHEMBL3970041 CHEMBL3970041 | C20H24ClN5O4 | 433.893 | 8 / 3 | 2.0 | Yes |
| 182179 | 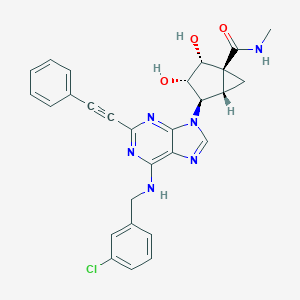 CHEMBL2064653 CHEMBL2064653 | C28H25ClN6O3 | 528.997 | 7 / 4 | 3.1 | No |
| 188776 | 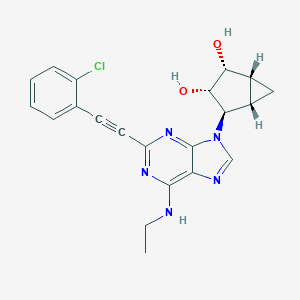 CHEMBL2089340 CHEMBL2089340 | C21H20ClN5O2 | 409.874 | 6 / 3 | 2.6 | Yes |
| 196792 | 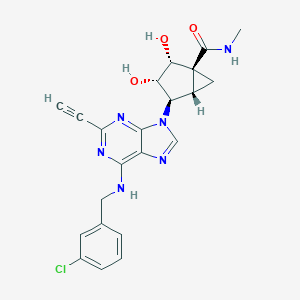 CHEMBL402574 CHEMBL402574 | C22H21ClN6O3 | 452.899 | 7 / 4 | 1.3 | Yes |
| 199449 | 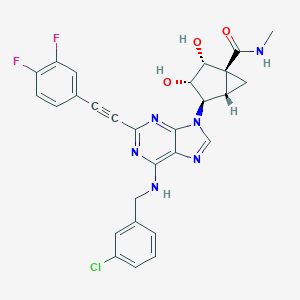 MRS 5698 MRS 5698 | C28H23ClF2N6O3 | 564.978 | 9 / 4 | 3.3 | No |
| 221197 | 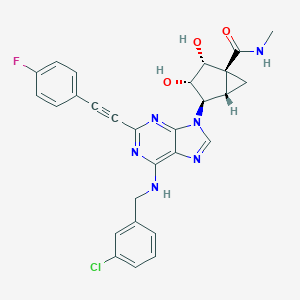 CHEMBL2064654 CHEMBL2064654 | C28H24ClFN6O3 | 546.987 | 8 / 4 | 3.2 | No |
| 528214 | 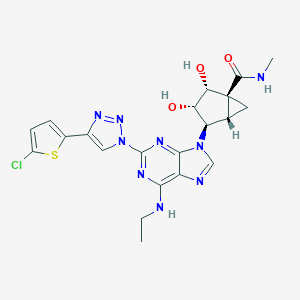 CHEMBL3612944 CHEMBL3612944 | C21H22ClN9O3S | 515.977 | 10 / 4 | 1.4 | No |
| 238309 | 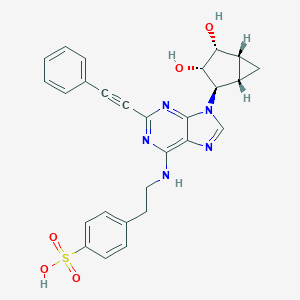 CHEMBL2402023 CHEMBL2402023 | C27H25N5O5S | 531.587 | 9 / 4 | 2.3 | No |
| 253418 | 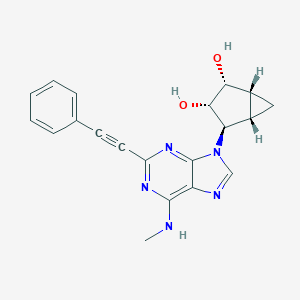 CHEMBL2089335 CHEMBL2089335 | C20H19N5O2 | 361.405 | 6 / 3 | 1.6 | Yes |
| 551262 | 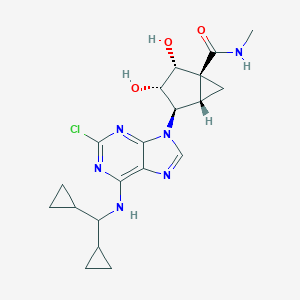 CHEMBL3976121 CHEMBL3976121 | C20H25ClN6O3 | 432.909 | 7 / 4 | 1.4 | Yes |
| 274618 | 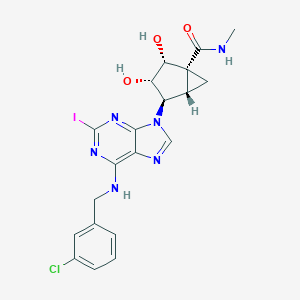 CHEMBL175444 CHEMBL175444 | C20H20ClIN6O3 | 554.773 | 7 / 4 | 1.7 | No |
| 452720 | 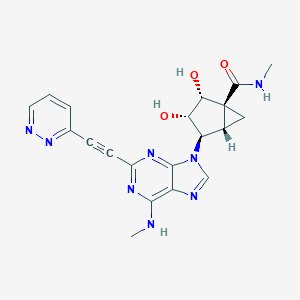 CHEMBL3407775 CHEMBL3407775 | C20H20N8O3 | 420.433 | 9 / 4 | -1.1 | Yes |
| 497444 | 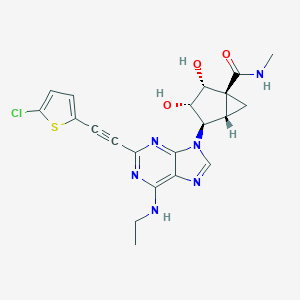 CHEMBL3612931 CHEMBL3612931 | C21H21ClN6O3S | 472.948 | 8 / 4 | 2.0 | Yes |
| 279243 | 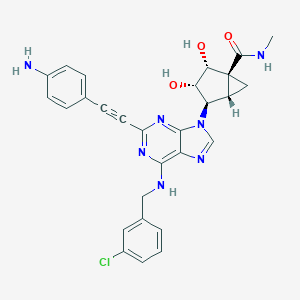 CHEMBL2064658 CHEMBL2064658 | C28H26ClN7O3 | 544.012 | 8 / 5 | 2.4 | No |
| 529471 | 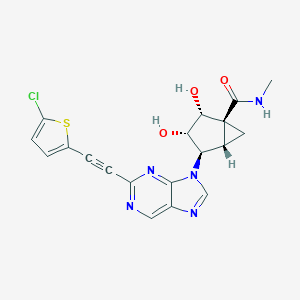 CHEMBL3799738 CHEMBL3799738 | C19H16ClN5O3S | 429.879 | 7 / 3 | 1.3 | Yes |
| 293840 | 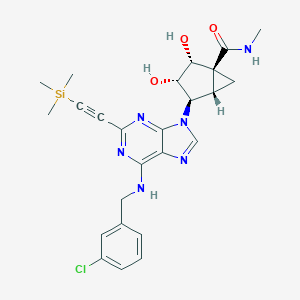 CHEMBL257405 CHEMBL257405 | C25H29ClN6O3Si | 525.081 | 7 / 4 | N/A | No |
| 453336 | 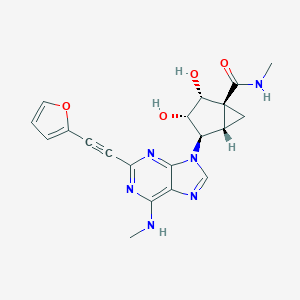 CHEMBL3407778 CHEMBL3407778 | C20H20N6O4 | 408.418 | 8 / 4 | 0.1 | Yes |
| 453817 | 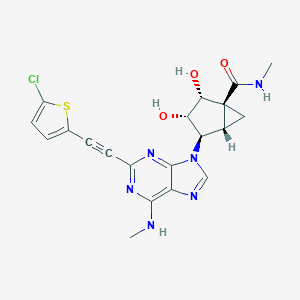 CHEMBL3407784 CHEMBL3407784 | C20H19ClN6O3S | 458.921 | 8 / 4 | 1.7 | Yes |
| 314972 | 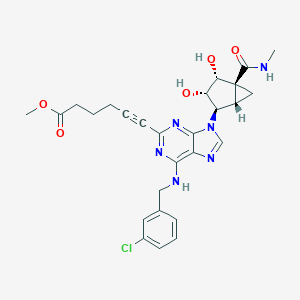 CHEMBL436649 CHEMBL436649 | C27H29ClN6O5 | 553.016 | 9 / 4 | 1.9 | No |
| 327782 | 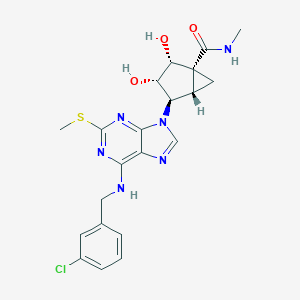 CHEMBL175814 CHEMBL175814 | C21H23ClN6O3S | 474.964 | 8 / 4 | 1.9 | Yes |
| 350086 | 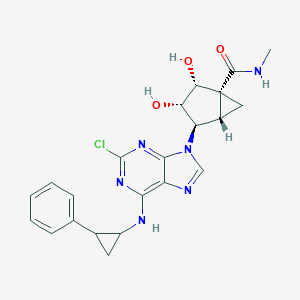 CHEMBL411861 CHEMBL411861 | C22H23ClN6O3 | 454.915 | 7 / 4 | 1.8 | Yes |
| 351324 | 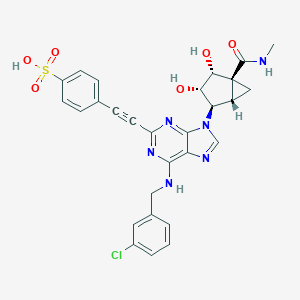 CHEMBL2401954 CHEMBL2401954 | C28H25ClN6O6S | 609.054 | 10 / 5 | 1.8 | No |
| 359728 | 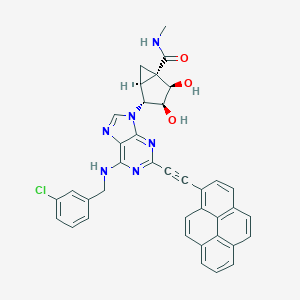 CHEMBL2064661 CHEMBL2064661 | C38H29ClN6O3 | 653.139 | 7 / 4 | 6.2 | No |
| 365915 | 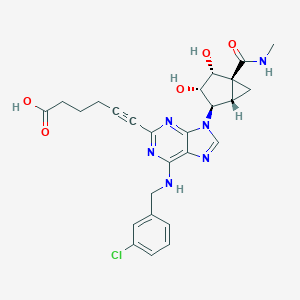 CHEMBL257833 CHEMBL257833 | C26H27ClN6O5 | 538.989 | 9 / 5 | 1.6 | No |
| 375339 | 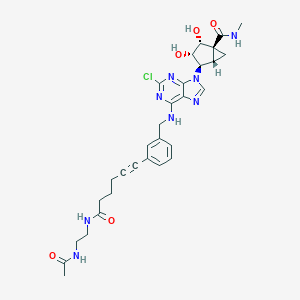 CHEMBL257573 CHEMBL257573 | C30H35ClN8O5 | 623.111 | 9 / 6 | 1.0 | No |
| 569367 | 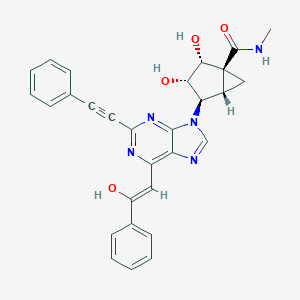 CHEMBL3800106 CHEMBL3800106 | C29H25N5O4 | 507.55 | 7 / 4 | 2.4 | No |
| 392275 | 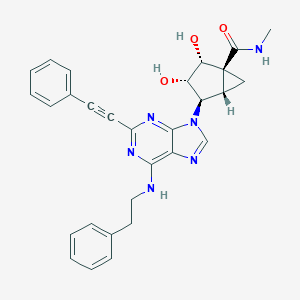 CHEMBL2402024 CHEMBL2402024 | C29H28N6O3 | 508.582 | 7 / 4 | 2.9 | No |
| 392917 | 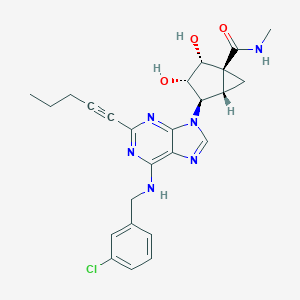 CHEMBL258421 CHEMBL258421 | C25H27ClN6O3 | 494.98 | 7 / 4 | 2.8 | Yes |
| 532642 | 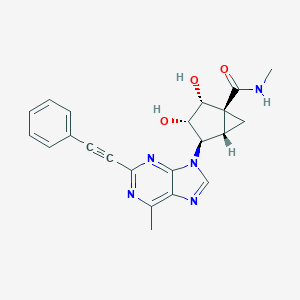 CHEMBL3800048 CHEMBL3800048 | C22H21N5O3 | 403.442 | 6 / 3 | 1.1 | Yes |
| 395905 | 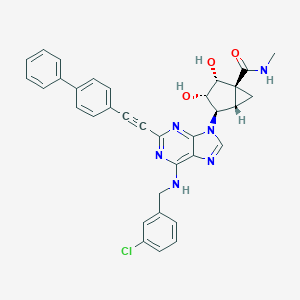 CHEMBL2064660 CHEMBL2064660 | C34H29ClN6O3 | 605.095 | 7 / 4 | 4.7 | No |
| 417308 | 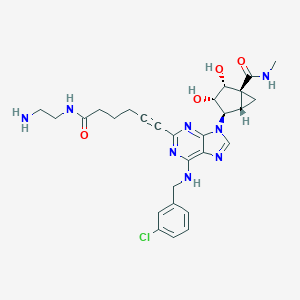 CHEMBL257785 CHEMBL257785 | C28H33ClN8O4 | 581.074 | 9 / 6 | 0.4 | No |
| 430313 | 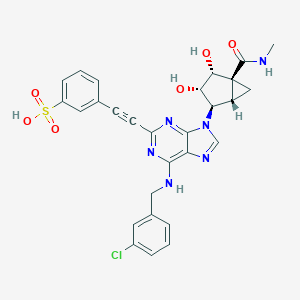 CHEMBL2401955 CHEMBL2401955 | C28H25ClN6O6S | 609.054 | 10 / 5 | 1.8 | No |
| 431158 | 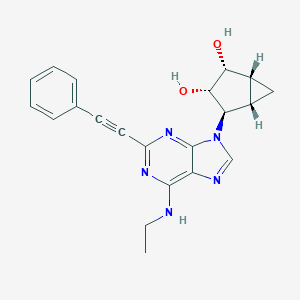 CHEMBL2089339 CHEMBL2089339 | C21H21N5O2 | 375.432 | 6 / 3 | 1.9 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218