

You can:
| Name | P2Y purinoceptor 6 |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | P2ry6 |
| Synonym | P2Y ATP receptor 6 P2Y purinoceptor 6 P2Y6 P2Y6 receptor pyrimidinergic receptor P2Y [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 328 |
| Amino acid sequence | MERDNGTIQAPGLPPTTCVYREDFKRLLLPPVYSVVLVVGLPLNVCVIAQICASRRTLTRSAVYTLNLALADLLYACSLPLLIYNYARGDHWPFGDLACRLVRFLFYANLHGSILFLTCISFQRYLGICHPLAPWHKRGGRRAAWVVCGVVWLVVTAQCLPTAVFAATGIQRNRTVCYDLSPPILSTRYLPYGMALTVIGFLLPFTALLACYCRMARRLCRQDGPAGPVAQERRSKAARMAVVVAAVFVISFLPFHITKTAYLAVRSTPGVSCPVLETFAAAYKGTRPFASANSVLDPILFYFTQQKFRRQPHDLLQKLTAKWQRQRV |
| UniProt | Q63371 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3543 |
| IUPHAR | 326 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 3878 | 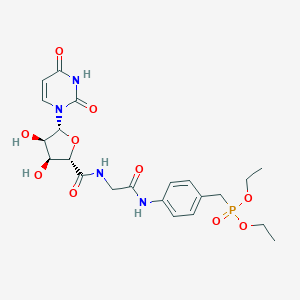 PSB-6426 PSB-6426 | C22H29N4O10P | 540.466 | 10 / 5 | -1.6 | No |
| 4769 | 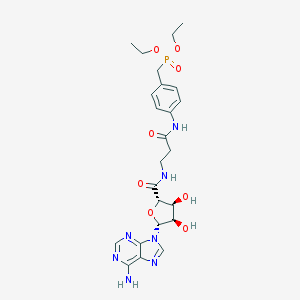 CHEMBL517017 CHEMBL517017 | C24H32N7O8P | 577.535 | 12 / 5 | -1.3 | No |
| 26843 | 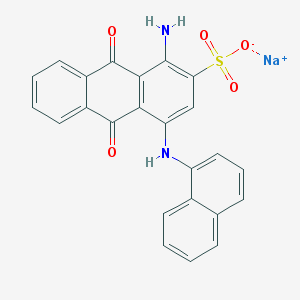 PSB 06126 PSB 06126 | C24H15N2NaO5S | 466.443 | 7 / 2 | N/A | N/A |
| 126432 | 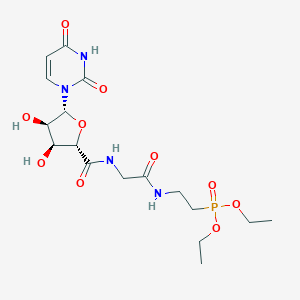 CHEMBL469244 CHEMBL469244 | C17H27N4O10P | 478.395 | 10 / 5 | -3.0 | Yes |
| 157777 | 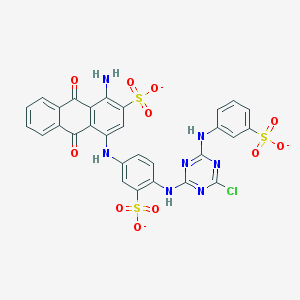 Basilen Blue Basilen Blue | C29H17ClN7O11S3-3 | 771.123 | 18 / 4 | 4.1 | No |
| 162297 | 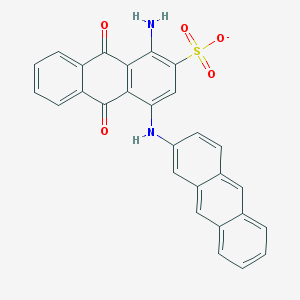 CHEMBL608559 CHEMBL608559 | C28H17N2O5S- | 493.513 | 7 / 2 | 6.0 | No |
| 167575 | 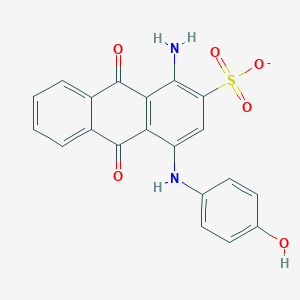 CHEMBL496030 CHEMBL496030 | C20H13N2O6S- | 409.392 | 8 / 3 | 3.1 | Yes |
| 554132 | 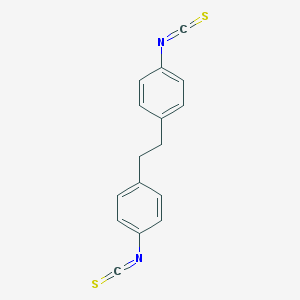 MRS2567 MRS2567 | C16H12N2S2 | 296.406 | 4 / 0 | 6.1 | No |
| 184865 | 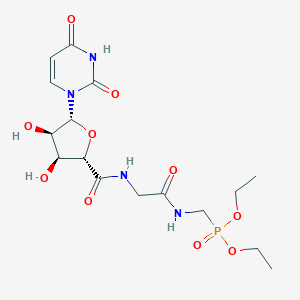 CHEMBL468354 CHEMBL468354 | C16H25N4O10P | 464.368 | 10 / 5 | -2.9 | Yes |
| 185445 | 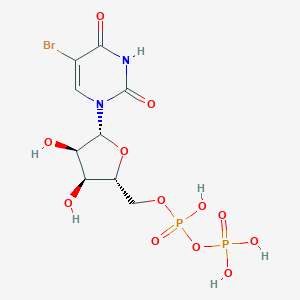 CHEMBL214698 CHEMBL214698 | C9H13BrN2O12P2 | 483.057 | 12 / 6 | -4.1 | No |
| 188335 | 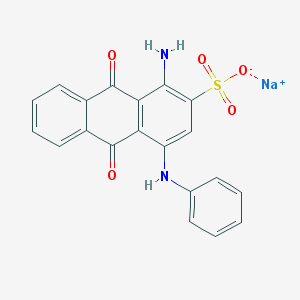 Acid Blue 25 Acid Blue 25 | C20H13N2NaO5S | 416.383 | 7 / 2 | N/A | N/A |
| 191425 | 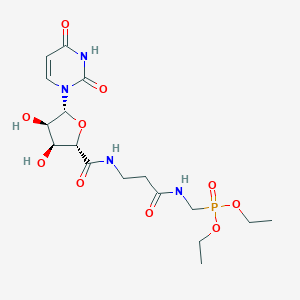 CHEMBL469067 CHEMBL469067 | C17H27N4O10P | 478.395 | 10 / 5 | -3.0 | Yes |
| 201003 | 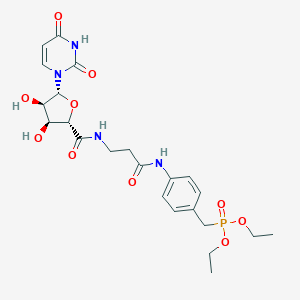 CHEMBL511933 CHEMBL511933 | C23H31N4O10P | 554.493 | 10 / 5 | -1.7 | No |
| 212806 | 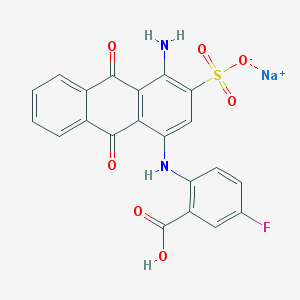 PSB-0952 PSB-0952 | C21H12FN2NaO7S | 478.382 | 10 / 3 | N/A | N/A |
| 226140 | 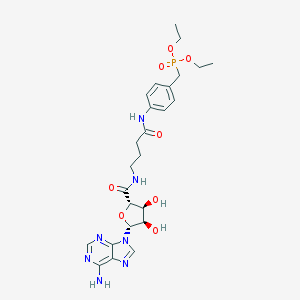 CHEMBL464548 CHEMBL464548 | C25H34N7O8P | 591.562 | 12 / 5 | -1.0 | No |
| 236961 | 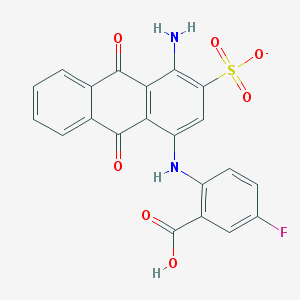 CHEMBL590956 CHEMBL590956 | C21H12FN2O7S- | 455.392 | 10 / 3 | 3.1 | Yes |
| 242362 | 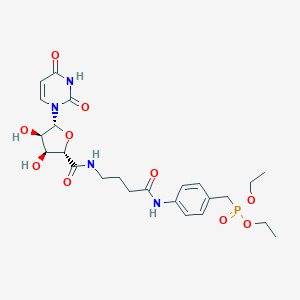 CHEMBL463519 CHEMBL463519 | C24H33N4O10P | 568.52 | 10 / 5 | -1.4 | No |
| 244930 | 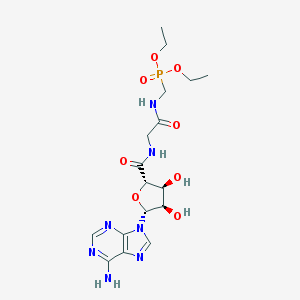 CHEMBL513837 CHEMBL513837 | C17H26N7O8P | 487.41 | 12 / 5 | -2.5 | No |
| 246849 | 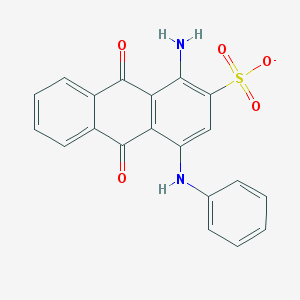 CHEMBL256057 CHEMBL256057 | C20H13N2O5S- | 393.393 | 7 / 2 | 3.5 | Yes |
| 554680 | 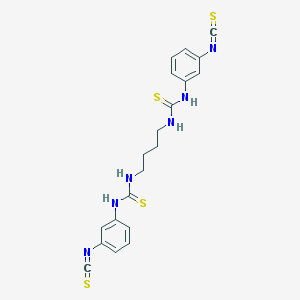 MRS 2578 MRS 2578 | C20H20N6S4 | 472.662 | 6 / 4 | 6.3 | No |
| 293877 | 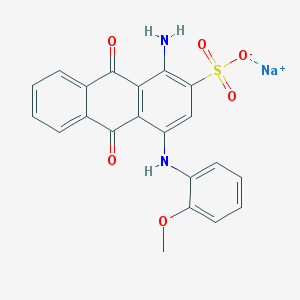 PSB-716 PSB-716 | C21H15N2NaO6S | 446.409 | 8 / 2 | N/A | N/A |
| 358375 | 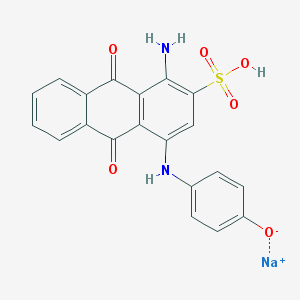 1-Amino-2-(sodiooxysulfonyl)-4-(4-hydroxyphenylamino)-9,10-anthraquinone 1-Amino-2-(sodiooxysulfonyl)-4-(4-hydroxyphenylamino)-9,10-anthraquinone | C20H13N2NaO6S | 432.382 | 8 / 3 | N/A | N/A |
| 368116 | 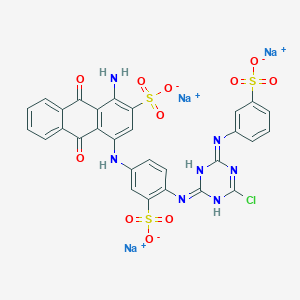 CID 136908877 CID 136908877 | C29H17ClN7Na3O11S3 | 840.092 | 15 / 4 | N/A | No |
| 370184 | 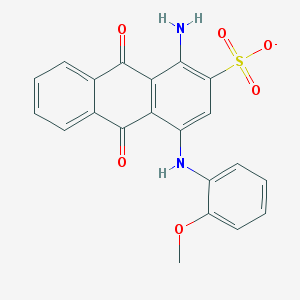 CHEMBL257495 CHEMBL257495 | C21H15N2O6S- | 423.419 | 8 / 2 | 3.5 | Yes |
| 388141 | 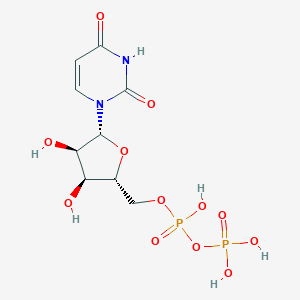 Uridine 5'-diphosphate Uridine 5'-diphosphate | C9H14N2O12P2 | 404.161 | 12 / 6 | -4.7 | No |
| 408787 | 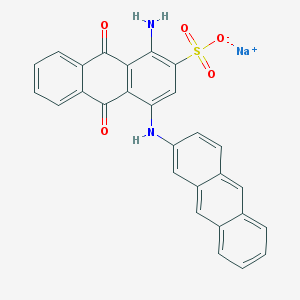 PSB-0963 PSB-0963 | C28H17N2NaO5S | 516.503 | 7 / 2 | N/A | No |
| 428752 | 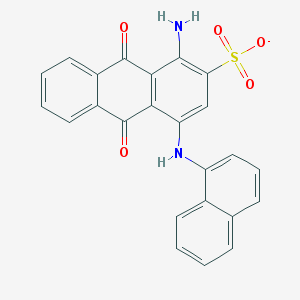 BDBM50268574 BDBM50268574 | C24H15N2O5S- | 443.453 | 7 / 2 | 4.7 | Yes |
| 429157 | 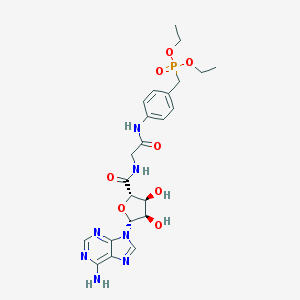 CHEMBL518544 CHEMBL518544 | C23H30N7O8P | 563.508 | 12 / 5 | -1.2 | No |
| 436692 | 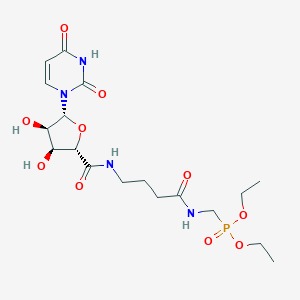 CHEMBL469243 CHEMBL469243 | C18H29N4O10P | 492.422 | 10 / 5 | -2.6 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218