

You can:
| Name | Neuropeptide Y receptor type 5 |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Npy5r |
| Synonym | food intake receptor neuropeptide Y receptor type 5 NPY-Y5 receptor NPY5-R NPYY5-R [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 445 |
| Amino acid sequence | MEFKLEEHFNKTFVTENNTAAARNAAFPAWEDYRGSVDDLQYFLIGLYTFVSLLGFMGNLLILMAVMKKRNQKTTVNFLIGNLAFSDILVVLFCSPFTLTSVLLDQWMFGKAMCHIMPFLQCVSVLVSTLILISIAIVRYHMIKHPISNNLTANHGYFLIATVWTLGFAICSPLPVFHSLVELKETFGSALLSSKYLCVESWPSDSYRIAFTISLLLVQYILPLVCLTVSHTSVCRSISCGLSHKENRLEENEMINLTLQPSKKSRNQAKTPSTQKWSYSFIRKHRRRYSKKTACVLPAPAGPSQGKHLAVPENPASVRSQLSPSSKVIPGVPICFEVKPEESSDAHEMRVKRSITRIKKRSRSVFYRLTILILVFAVSWMPLHVFHVVTDFNDNLISNRHFKLVYCICHLLGMMSCCLNPILYGFLNNGIKADLRALIHCLHMS |
| UniProt | Q63634 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL2548 |
| IUPHAR | 308 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 9460 | 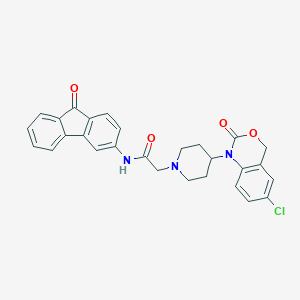 CHEMBL541891 CHEMBL541891 | C28H24ClN3O4 | 501.967 | 5 / 1 | 4.5 | No |
| 9984 | 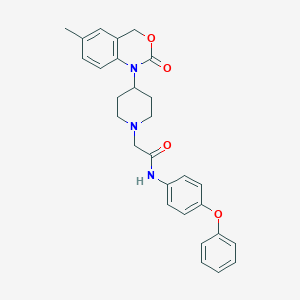 CHEMBL539084 CHEMBL539084 | C28H29N3O4 | 471.557 | 5 / 1 | 4.6 | Yes |
| 15542 | 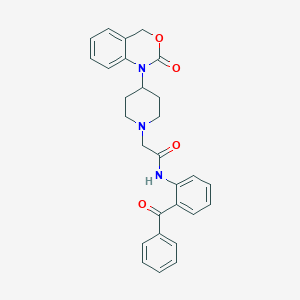 CHEMBL534710 CHEMBL534710 | C28H27N3O4 | 469.541 | 5 / 1 | 4.6 | Yes |
| 25968 | 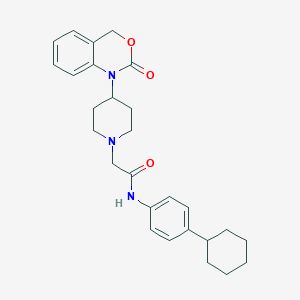 CHEMBL536496 CHEMBL536496 | C27H33N3O3 | 447.579 | 4 / 1 | 5.3 | No |
| 28464 | 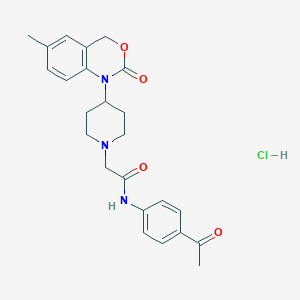 CHEMBL537187 CHEMBL537187 | C24H28ClN3O4 | 457.955 | 5 / 2 | N/A | N/A |
| 32154 | 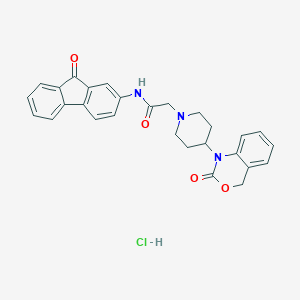 CHEMBL556951 CHEMBL556951 | C28H26ClN3O4 | 503.983 | 5 / 2 | N/A | No |
| 33455 | 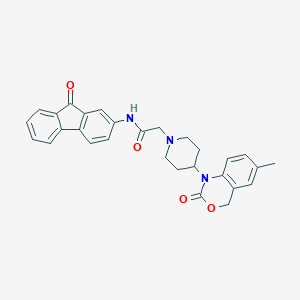 CHEMBL538823 CHEMBL538823 | C29H27N3O4 | 481.552 | 5 / 1 | 4.3 | Yes |
| 34200 | 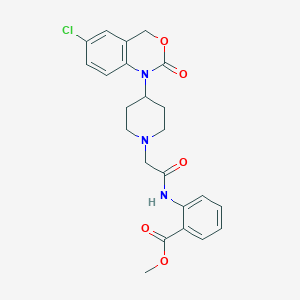 CHEMBL372724 CHEMBL372724 | C23H24ClN3O5 | 457.911 | 6 / 1 | 3.8 | Yes |
| 36129 | 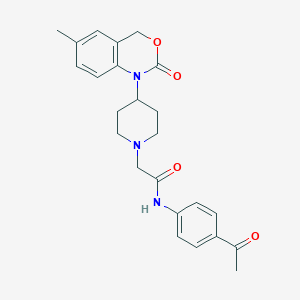 CHEMBL537187 CHEMBL537187 | C24H27N3O4 | 421.497 | 5 / 1 | 2.8 | Yes |
| 553439 | 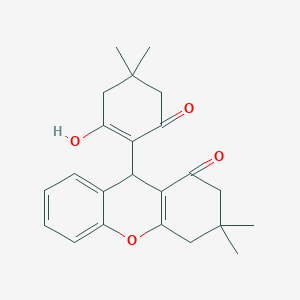 CHEMBL491762 CHEMBL491762 | C23H26O4 | 366.457 | 4 / 1 | 3.6 | Yes |
| 39565 | 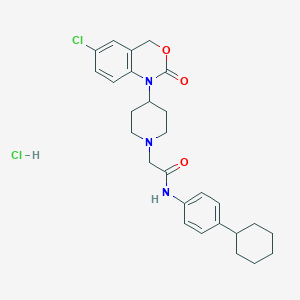 CHEMBL536045 CHEMBL536045 | C27H33Cl2N3O3 | 518.479 | 4 / 2 | N/A | No |
| 43003 | 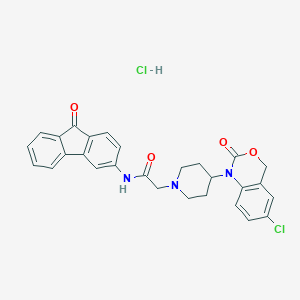 CHEMBL541891 CHEMBL541891 | C28H25Cl2N3O4 | 538.425 | 5 / 2 | N/A | No |
| 46404 | 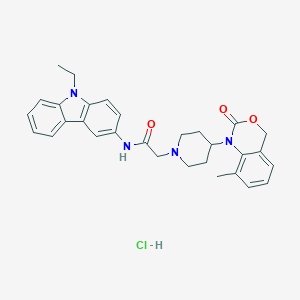 CHEMBL540365 CHEMBL540365 | C30H33ClN4O3 | 533.069 | 4 / 2 | N/A | No |
| 555670 | 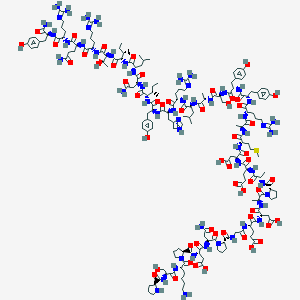 CID 57339554 CID 57339554 | C180H276N54O55S | 4108.57 | 63 / 58 | -18.4 | No |
| 52015 | 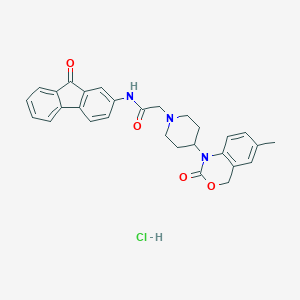 CHEMBL538823 CHEMBL538823 | C29H28ClN3O4 | 518.01 | 5 / 2 | N/A | No |
| 52790 | 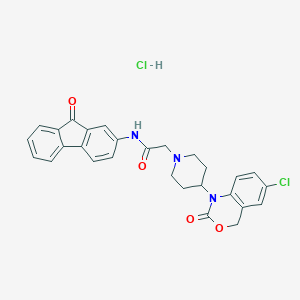 CHEMBL555513 CHEMBL555513 | C28H25Cl2N3O4 | 538.425 | 5 / 2 | N/A | No |
| 57217 | 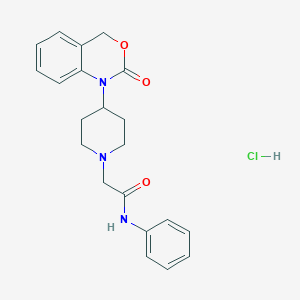 CHEMBL534709 CHEMBL534709 | C21H24ClN3O3 | 401.891 | 4 / 2 | N/A | N/A |
| 60473 | 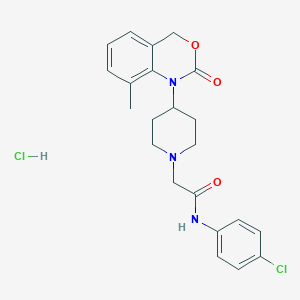 CHEMBL534483 CHEMBL534483 | C22H25Cl2N3O3 | 450.36 | 4 / 2 | N/A | N/A |
| 69958 | 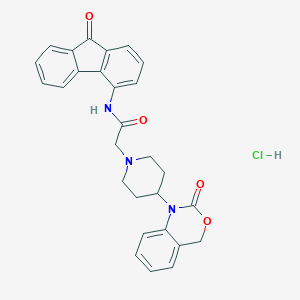 CHEMBL536275 CHEMBL536275 | C28H26ClN3O4 | 503.983 | 5 / 2 | N/A | No |
| 72355 | 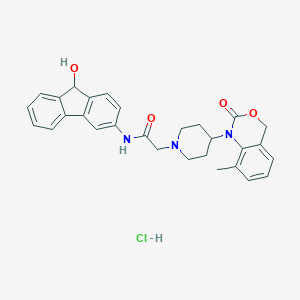 CHEMBL535824 CHEMBL535824 | C29H30ClN3O4 | 520.026 | 5 / 3 | N/A | No |
| 83419 | 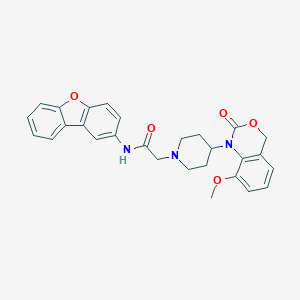 CHEMBL365777 CHEMBL365777 | C28H27N3O5 | 485.54 | 6 / 1 | 4.5 | Yes |
| 93258 | 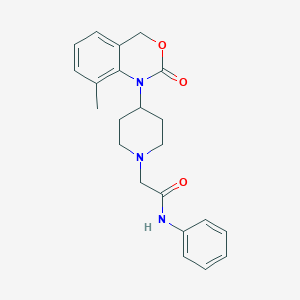 CHEMBL538323 CHEMBL538323 | C22H25N3O3 | 379.46 | 4 / 1 | 3.1 | Yes |
| 555811 | 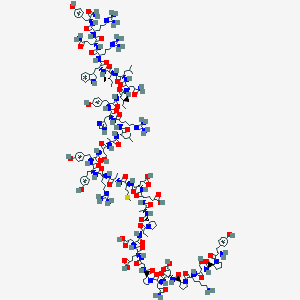 NPY D-Trp32 NPY D-Trp32 | C196H290N56O56S | 4358.87 | 65 / 59 | -15.6 | No |
| 102557 | 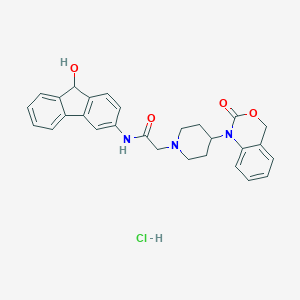 CHEMBL557802 CHEMBL557802 | C28H28ClN3O4 | 505.999 | 5 / 3 | N/A | No |
| 106173 | 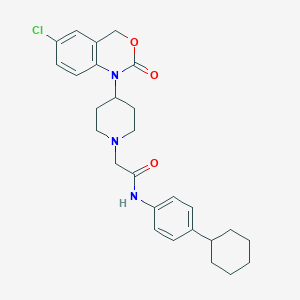 CHEMBL536045 CHEMBL536045 | C27H32ClN3O3 | 482.021 | 4 / 1 | 5.9 | No |
| 555908 | 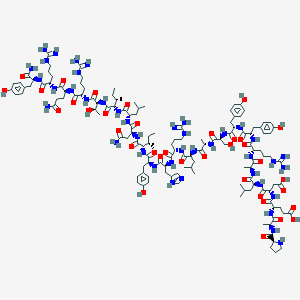 CID 16142386 CID 16142386 | C135H209N41O36 | 2982.41 | 42 / 48 | -6.6 | No |
| 119886 | 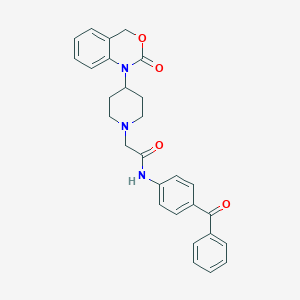 CHEMBL179972 CHEMBL179972 | C28H27N3O4 | 469.541 | 5 / 1 | 4.1 | Yes |
| 555970 |  BDBM82300 BDBM82300 | C195H302N58O57S+4 | 4402.97 | 61 / 65 | -14.8 | No |
| 555992 | 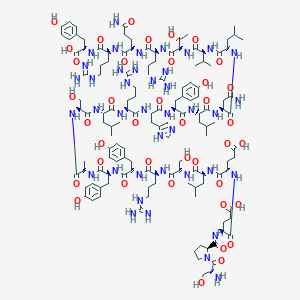 PYY13-36, porcine PYY13-36, porcine | C135H208N40O39 | 3015.39 | 45 / 49 | -8.3 | No |
| 128902 | 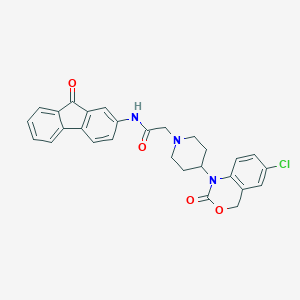 CHEMBL555513 CHEMBL555513 | C28H24ClN3O4 | 501.967 | 5 / 1 | 4.5 | No |
| 131697 | 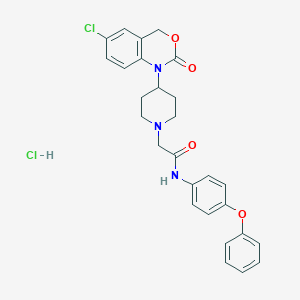 CHEMBL537858 CHEMBL537858 | C27H27Cl2N3O4 | 528.43 | 5 / 2 | N/A | No |
| 139400 | 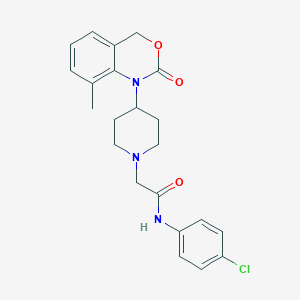 CHEMBL534483 CHEMBL534483 | C22H24ClN3O3 | 413.902 | 4 / 1 | 3.7 | Yes |
| 142571 | 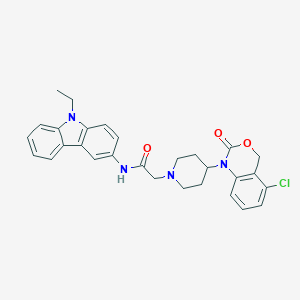 CHEMBL557817 CHEMBL557817 | C29H29ClN4O3 | 517.026 | 4 / 1 | 5.1 | No |
| 144050 | 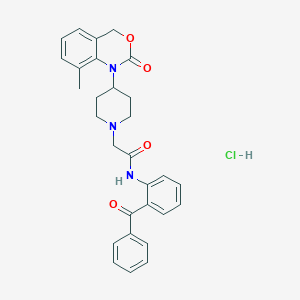 CHEMBL553250 CHEMBL553250 | C29H30ClN3O4 | 520.026 | 5 / 2 | N/A | No |
| 556028 | 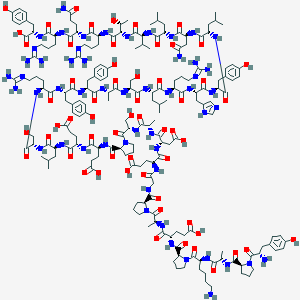 CID 77077859 CID 77077859 | C190H287N53O58 | 4241.7 | 65 / 59 | -15.5 | No |
| 154403 | 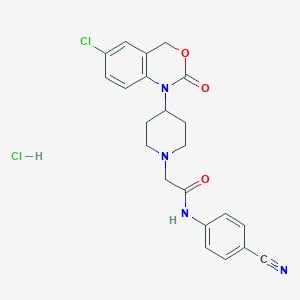 CHEMBL557583 CHEMBL557583 | C22H22Cl2N4O3 | 461.343 | 5 / 2 | N/A | N/A |
| 158556 | 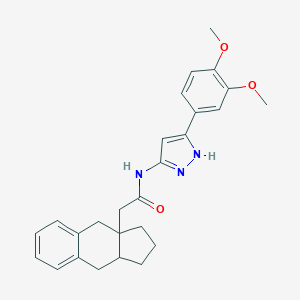 CHEMBL60319 CHEMBL60319 | C26H29N3O3 | 431.536 | 4 / 2 | 5.0 | Yes |
| 164760 | 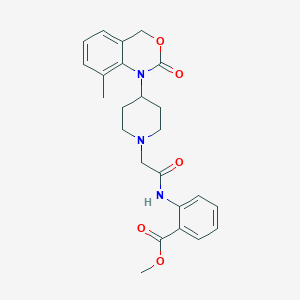 CHEMBL193554 CHEMBL193554 | C24H27N3O5 | 437.496 | 6 / 1 | 3.5 | Yes |
| 168415 | 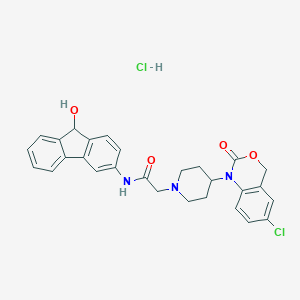 CHEMBL557172 CHEMBL557172 | C28H27Cl2N3O4 | 540.441 | 5 / 3 | N/A | No |
| 168725 | 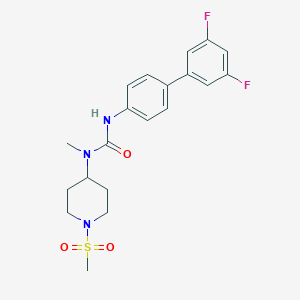 CHEMBL403414 CHEMBL403414 | C20H23F2N3O3S | 423.479 | 6 / 1 | 2.8 | Yes |
| 169743 | 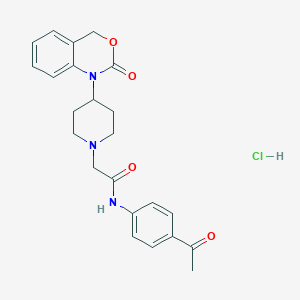 CHEMBL537412 CHEMBL537412 | C23H26ClN3O4 | 443.928 | 5 / 2 | N/A | N/A |
| 174043 | 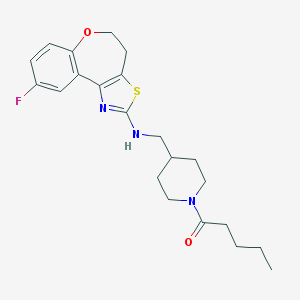 CHEMBL82374 CHEMBL82374 | C22H28FN3O2S | 417.543 | 6 / 1 | 4.7 | Yes |
| 556171 | 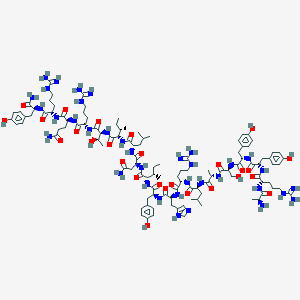 BDBM50368404 BDBM50368404 | C112H174N36O27 | 2456.85 | 33 / 41 | -3.6 | No |
| 556190 | 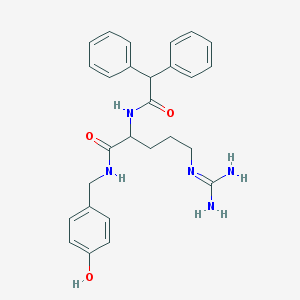 CHEMBL44246 CHEMBL44246 | C27H31N5O3 | 473.577 | 4 / 5 | 2.5 | Yes |
| 188027 | 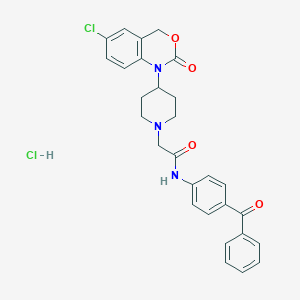 CHEMBL541121 CHEMBL541121 | C28H27Cl2N3O4 | 540.441 | 5 / 2 | N/A | No |
| 204031 | 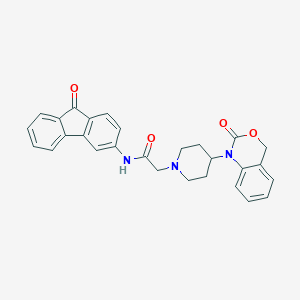 CHEMBL559210 CHEMBL559210 | C28H25N3O4 | 467.525 | 5 / 1 | 3.9 | Yes |
| 207965 | 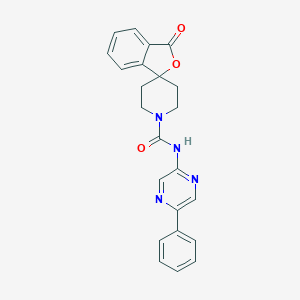 CHEMBL491930 CHEMBL491930 | C23H20N4O3 | 400.438 | 5 / 1 | 2.4 | Yes |
| 214928 | 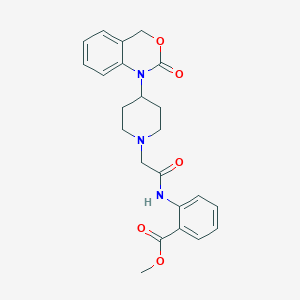 CHEMBL193770 CHEMBL193770 | C23H25N3O5 | 423.469 | 6 / 1 | 3.2 | Yes |
| 215731 | 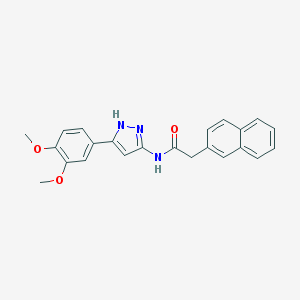 CHEMBL291666 CHEMBL291666 | C23H21N3O3 | 387.439 | 4 / 2 | 4.2 | Yes |
| 216027 | 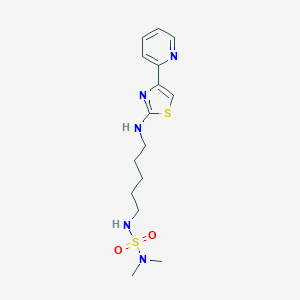 CHEMBL1909725 CHEMBL1909725 | C15H23N5O2S2 | 369.502 | 8 / 2 | 2.0 | Yes |
| 220888 | 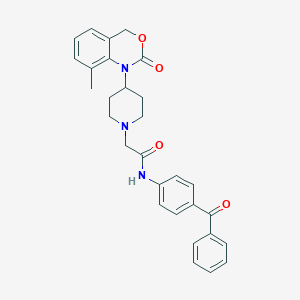 CHEMBL536276 CHEMBL536276 | C29H29N3O4 | 483.568 | 5 / 1 | 4.5 | Yes |
| 224377 | 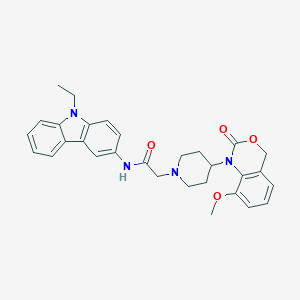 CHEMBL538569 CHEMBL538569 | C30H32N4O4 | 512.61 | 5 / 1 | 4.4 | No |
| 239979 | 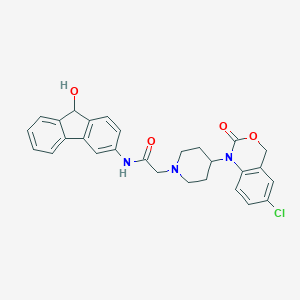 CHEMBL557172 CHEMBL557172 | C28H26ClN3O4 | 503.983 | 5 / 2 | 3.9 | No |
| 242749 | 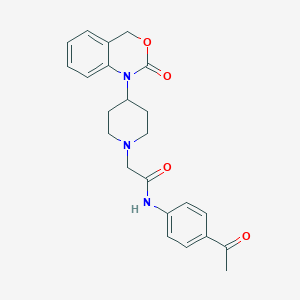 CHEMBL537412 CHEMBL537412 | C23H25N3O4 | 407.47 | 5 / 1 | 2.4 | Yes |
| 243531 | 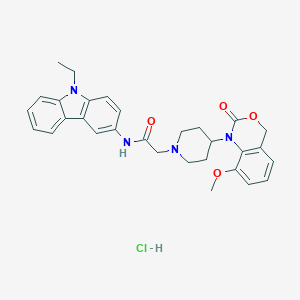 CHEMBL538569 CHEMBL538569 | C30H33ClN4O4 | 549.068 | 5 / 2 | N/A | No |
| 254864 | 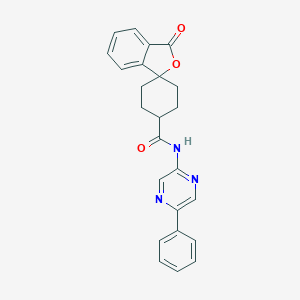 CHEMBL593934 CHEMBL593934 | C24H21N3O3 | 399.45 | 5 / 1 | 3.1 | Yes |
| 257002 | 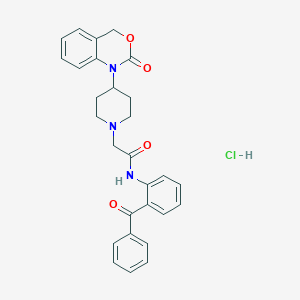 CHEMBL534710 CHEMBL534710 | C28H28ClN3O4 | 505.999 | 5 / 2 | N/A | No |
| 257230 | 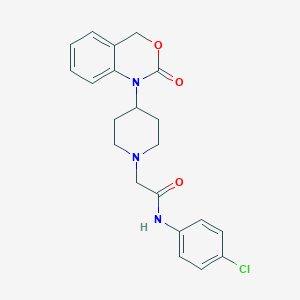 CHEMBL558393 CHEMBL558393 | C21H22ClN3O3 | 399.875 | 4 / 1 | 3.4 | Yes |
| 262749 | 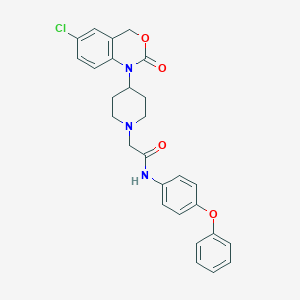 CHEMBL537858 CHEMBL537858 | C27H26ClN3O4 | 491.972 | 5 / 1 | 4.9 | Yes |
| 265031 | 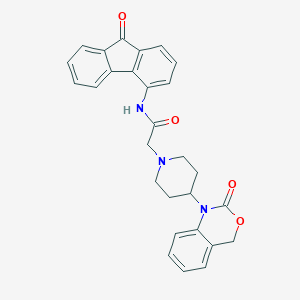 CHEMBL536275 CHEMBL536275 | C28H25N3O4 | 467.525 | 5 / 1 | 3.9 | Yes |
| 265991 | 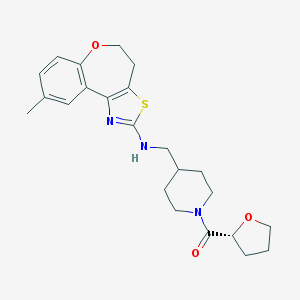 CHEMBL311229 CHEMBL311229 | C23H29N3O3S | 427.563 | 6 / 1 | 4.0 | Yes |
| 267524 | 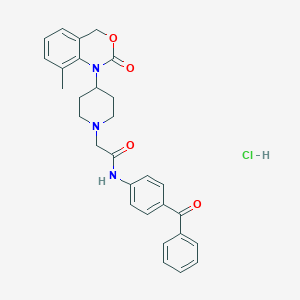 CHEMBL536276 CHEMBL536276 | C29H30ClN3O4 | 520.026 | 5 / 2 | N/A | No |
| 268586 | 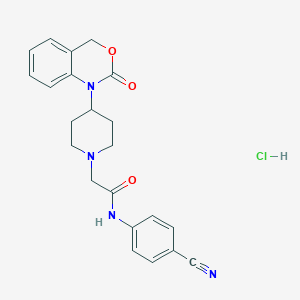 CHEMBL556999 CHEMBL556999 | C22H23ClN4O3 | 426.901 | 5 / 2 | N/A | N/A |
| 286714 | 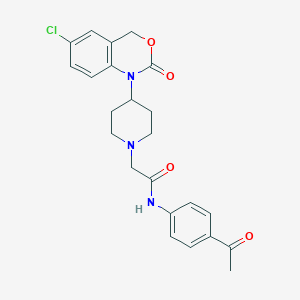 CHEMBL537191 CHEMBL537191 | C23H24ClN3O4 | 441.912 | 5 / 1 | 3.1 | Yes |
| 295051 | 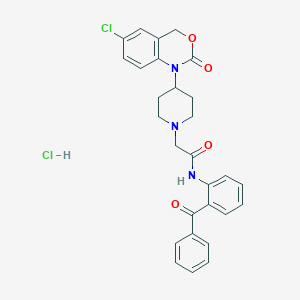 CHEMBL536729 CHEMBL536729 | C28H27Cl2N3O4 | 540.441 | 5 / 2 | N/A | No |
| 298732 | 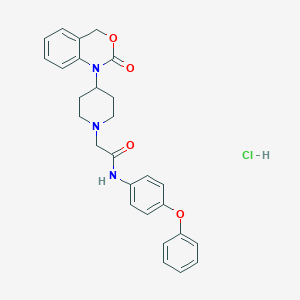 CHEMBL536048 CHEMBL536048 | C27H28ClN3O4 | 493.988 | 5 / 2 | N/A | N/A |
| 301873 | 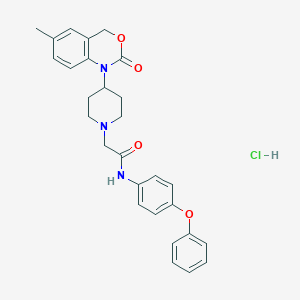 CHEMBL539084 CHEMBL539084 | C28H30ClN3O4 | 508.015 | 5 / 2 | N/A | No |
| 307175 | 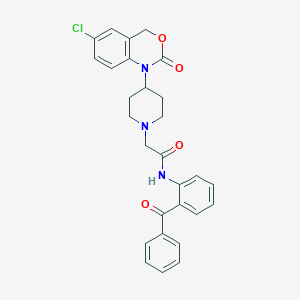 CHEMBL536729 CHEMBL536729 | C28H26ClN3O4 | 503.983 | 5 / 1 | 5.3 | No |
| 311856 | 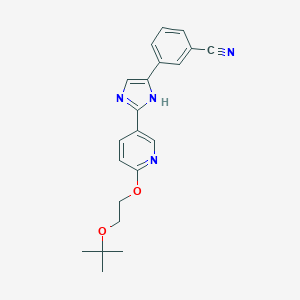 CHEMBL161724 CHEMBL161724 | C21H22N4O2 | 362.433 | 5 / 1 | 3.2 | Yes |
| 313873 | 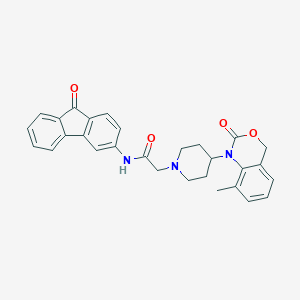 CHEMBL538079 CHEMBL538079 | C29H27N3O4 | 481.552 | 5 / 1 | 4.3 | Yes |
| 314939 | 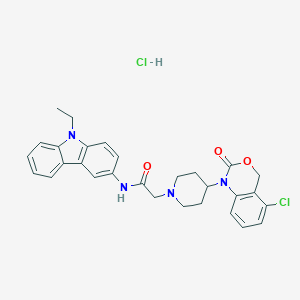 CHEMBL557817 CHEMBL557817 | C29H30Cl2N4O3 | 553.484 | 4 / 2 | N/A | No |
| 315264 | 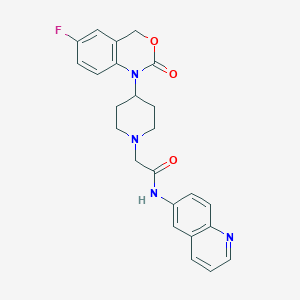 CHEMBL179154 CHEMBL179154 | C24H23FN4O3 | 434.471 | 6 / 1 | 3.1 | Yes |
| 317750 | 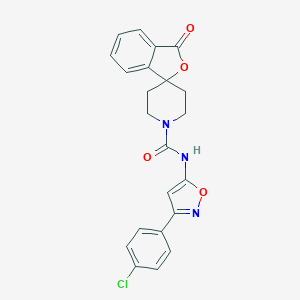 CHEMBL524085 CHEMBL524085 | C22H18ClN3O4 | 423.853 | 5 / 1 | 3.6 | Yes |
| 321530 | 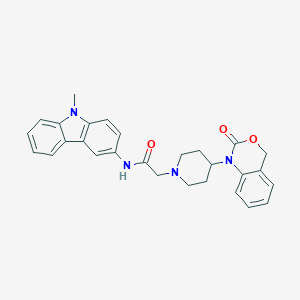 CHEMBL557540 CHEMBL557540 | C28H28N4O3 | 468.557 | 4 / 1 | 4.2 | Yes |
| 322775 | 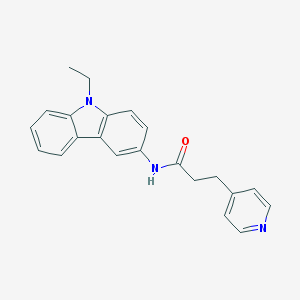 CHEMBL325475 CHEMBL325475 | C22H21N3O | 343.43 | 2 / 1 | 3.6 | Yes |
| 323992 | 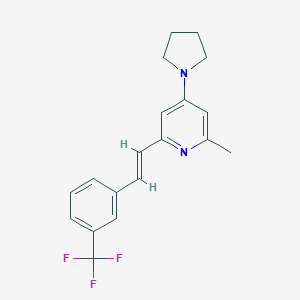 CHEMBL61488 CHEMBL61488 | C19H19F3N2 | 332.37 | 5 / 0 | 5.1 | No |
| 329864 | 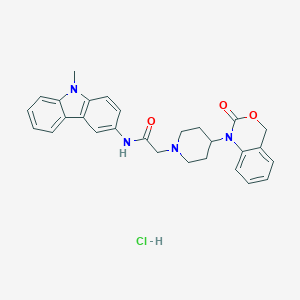 CHEMBL557540 CHEMBL557540 | C28H29ClN4O3 | 505.015 | 4 / 2 | N/A | No |
| 335785 | 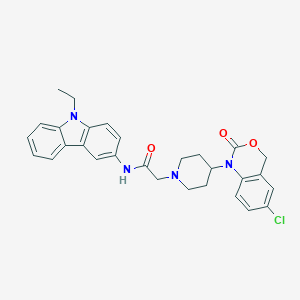 CHEMBL537634 CHEMBL537634 | C29H29ClN4O3 | 517.026 | 4 / 1 | 5.1 | No |
| 556868 | 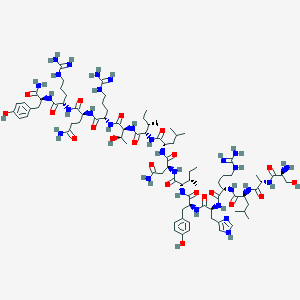 119019-65-7 119019-65-7 | C85H139N29O21 | 1903.23 | 26 / 32 | -4.2 | No |
| 341949 | 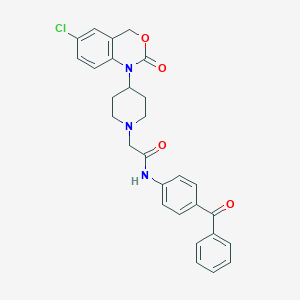 CHEMBL541121 CHEMBL541121 | C28H26ClN3O4 | 503.983 | 5 / 1 | 4.7 | No |
| 343347 | 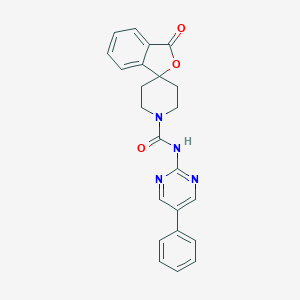 CHEMBL522388 CHEMBL522388 | C23H20N4O3 | 400.438 | 5 / 1 | 2.8 | Yes |
| 555006 | 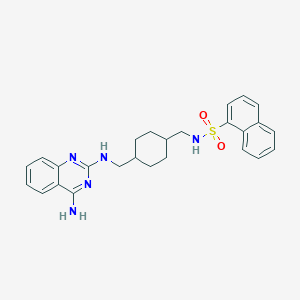 CGP-71683A CGP-71683A | C26H29N5O2S | 475.611 | 7 / 3 | 5.1 | No |
| 354148 | 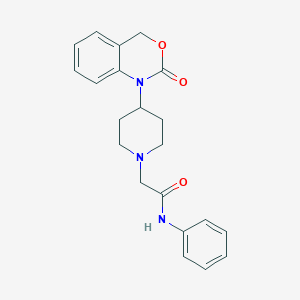 CHEMBL534709 CHEMBL534709 | C21H23N3O3 | 365.433 | 4 / 1 | 2.8 | Yes |
| 556950 | 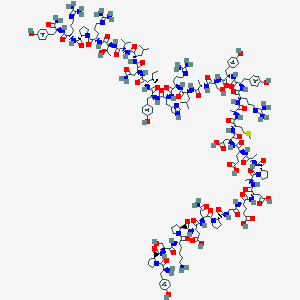 L31,P34-NPY,human L31,P34-NPY,human | C188H282N54O56S | 4226.71 | 64 / 57 | -16.6 | No |
| 357432 | 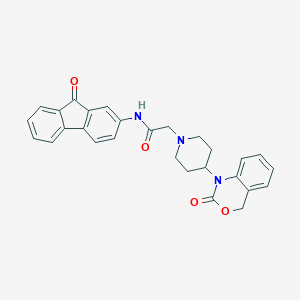 CHEMBL556951 CHEMBL556951 | C28H25N3O4 | 467.525 | 5 / 1 | 3.9 | Yes |
| 360308 | 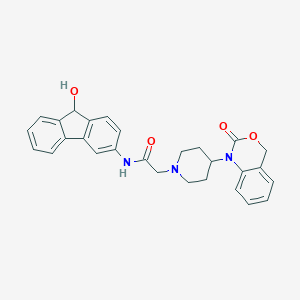 CHEMBL557802 CHEMBL557802 | C28H27N3O4 | 469.541 | 5 / 2 | 3.3 | Yes |
| 363170 | 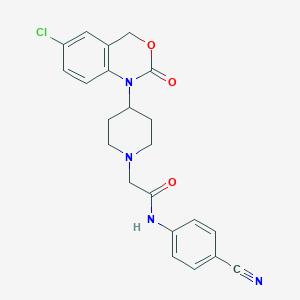 CHEMBL557583 CHEMBL557583 | C22H21ClN4O3 | 424.885 | 5 / 1 | 3.1 | Yes |
| 363676 | 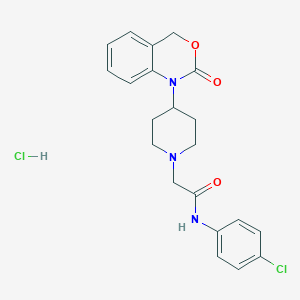 CHEMBL558393 CHEMBL558393 | C21H23Cl2N3O3 | 436.333 | 4 / 2 | N/A | N/A |
| 369354 | 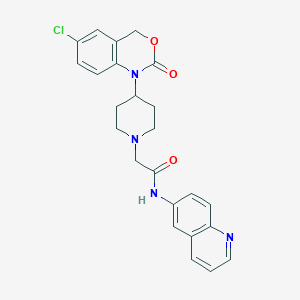 CHEMBL193382 CHEMBL193382 | C24H23ClN4O3 | 450.923 | 5 / 1 | 3.6 | Yes |
| 557062 | 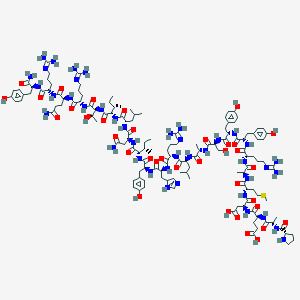 PYY13-36, rat PYY13-36, rat | C134H207N41O36S | 3000.44 | 43 / 44 | -8.7 | No |
| 383181 | 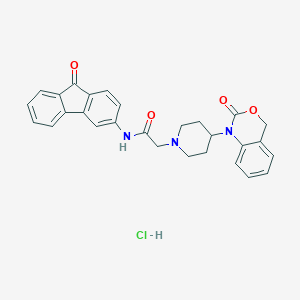 CHEMBL559210 CHEMBL559210 | C28H26ClN3O4 | 503.983 | 5 / 2 | N/A | No |
| 392543 | 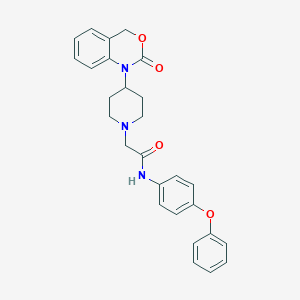 CHEMBL536048 CHEMBL536048 | C27H27N3O4 | 457.53 | 5 / 1 | 4.3 | Yes |
| 555258 | 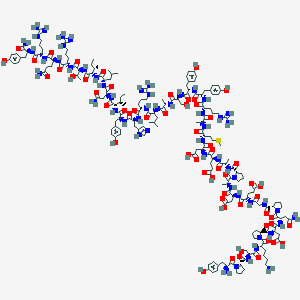 human Neuropeptide Y human Neuropeptide Y | C189H285N55O57S | 4271.75 | 65 / 63 | -16.2 | No |
| 395091 | 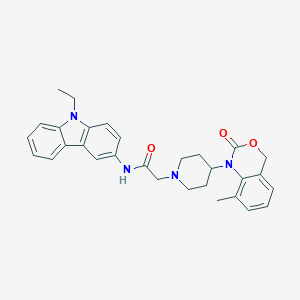 CHEMBL540365 CHEMBL540365 | C30H32N4O3 | 496.611 | 4 / 1 | 4.8 | Yes |
| 397587 | 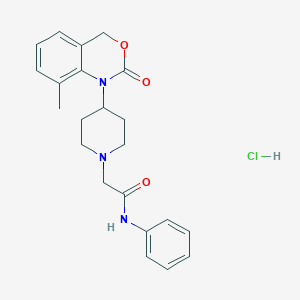 CHEMBL538323 CHEMBL538323 | C22H26ClN3O3 | 415.918 | 4 / 2 | N/A | N/A |
| 401978 | 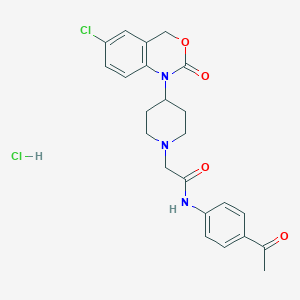 CHEMBL537191 CHEMBL537191 | C23H25Cl2N3O4 | 478.37 | 5 / 2 | N/A | N/A |
| 408232 | 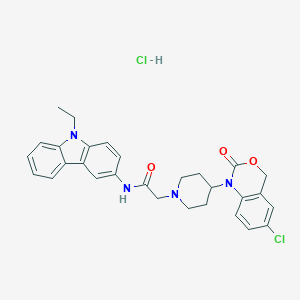 CHEMBL537634 CHEMBL537634 | C29H30Cl2N4O3 | 553.484 | 4 / 2 | N/A | No |
| 557217 | 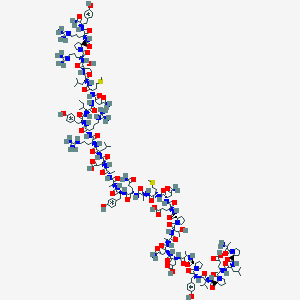 PP,SALMON PP,SALMON | C185H289N53O54S2+2 | 4183.78 | 59 / 56 | -12.7 | No |
| 421018 | 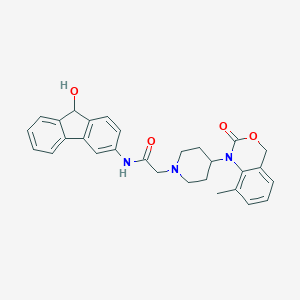 CHEMBL535824 CHEMBL535824 | C29H29N3O4 | 483.568 | 5 / 2 | 3.6 | Yes |
| 423788 | 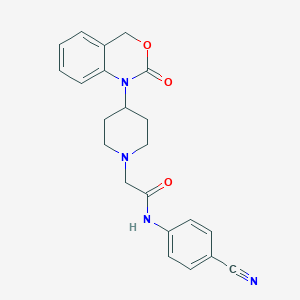 CHEMBL556999 CHEMBL556999 | C22H22N4O3 | 390.443 | 5 / 1 | 2.5 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218