

You can:
| Name | Mas-related G-protein coupled receptor member X1 |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Mrgprx1 |
| Synonym | Sensory neuron-specific G-protein coupled receptor 1 |
| Disease | N/A for non-human GPCRs |
| Length | 323 |
| Amino acid sequence | MDPTISSLSTESTTLNKTGHPSCRPILTLSFLVPIITLLGLAGNTIVLWLLGFRMRRKAISVYVLNLSLADSFFLCCHFIDSLMRIMNFYGIYAHKLSKEILGNAAIIPYISGLSILSAISTERCLSVLWPIWYHCHRPRNMSAIICVLIWVLSFLMGILDWFFSGFLGETHHHLWKNVDFIVTAFLIFLFMLLFGSSLALLVRILCGSRRKPLSRLYVTISLTVMVYLICGLPLGLYLFLLYWFGIHLHYPFCHIYQVTVLLSCVNSSANPIIYFLVGSFRHRKKHRSLKMVLKRALEETPEEDEYTDSHVQKPTEISERRC |
| UniProt | Q8R4G1 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3341575 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 521494 | 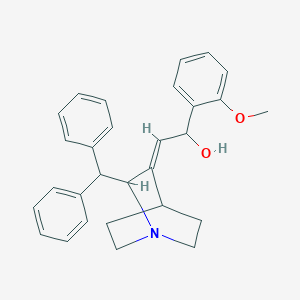 CHEMBL3730717 CHEMBL3730717 | C29H31NO2 | 425.572 | 3 / 1 | 4.9 | Yes |
| 442532 | 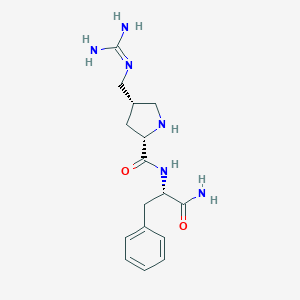 CHEMBL3343991 CHEMBL3343991 | C16H24N6O2 | 332.408 | 4 / 5 | -1.0 | Yes |
| 443621 | 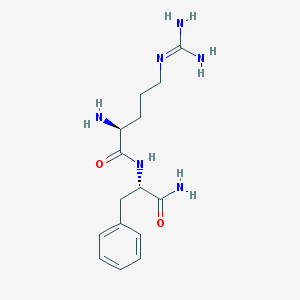 Rfamide Rfamide | C15H24N6O2 | 320.397 | 4 / 5 | -2.0 | Yes |
| 523547 | 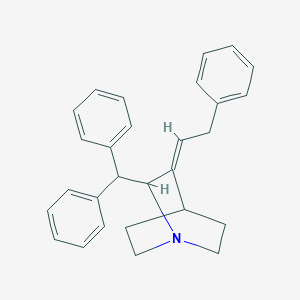 CHEMBL3731811 CHEMBL3731811 | C28H29N | 379.547 | 1 / 0 | 6.1 | No |
| 524417 | 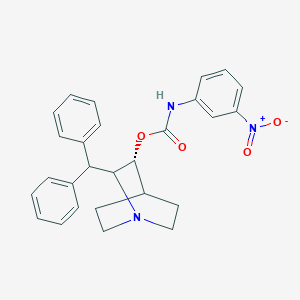 CHEMBL3732603 CHEMBL3732603 | C27H27N3O4 | 457.53 | 5 / 1 | 5.4 | No |
| 445680 | 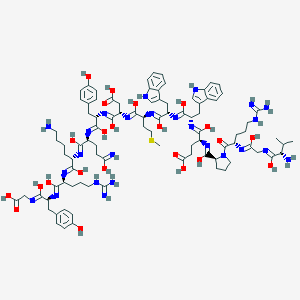 BAM(8-22) BAM(8-22) | C91H127N25O23S | 1971.23 | 42 / 30 | 1.4 | No |
| 524643 | 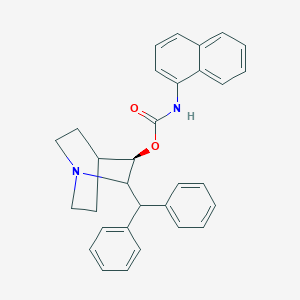 CHEMBL3730405 CHEMBL3730405 | C31H30N2O2 | 462.593 | 3 / 1 | 6.9 | No |
| 446904 | 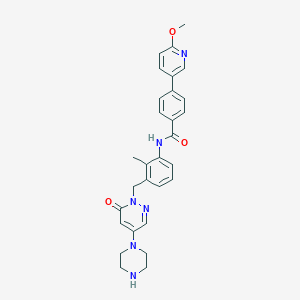 1103458-91-8 1103458-91-8 | C29H30N6O3 | 510.598 | 7 / 2 | 2.7 | No |
| 525387 | 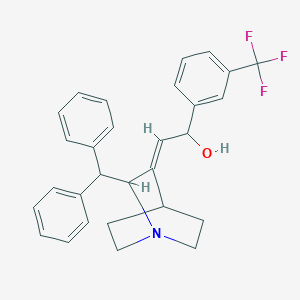 CHEMBL3730500 CHEMBL3730500 | C29H28F3NO | 463.544 | 5 / 1 | 5.9 | No |
| 447649 | 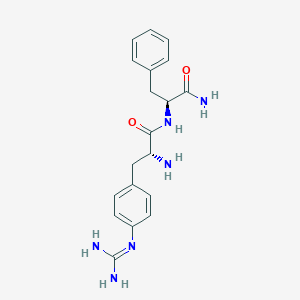 CHEMBL3343994 CHEMBL3343994 | C19H24N6O2 | 368.441 | 4 / 5 | -0.4 | Yes |
| 447653 | 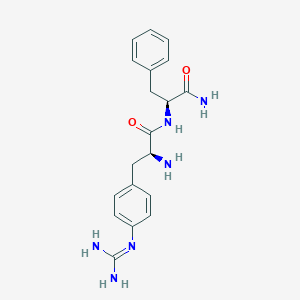 CHEMBL3343993 CHEMBL3343993 | C19H24N6O2 | 368.441 | 4 / 5 | -0.4 | Yes |
| 528780 | 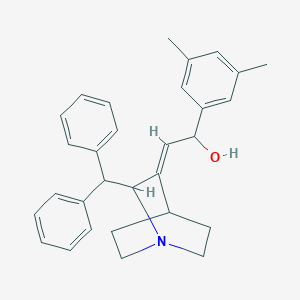 CHEMBL3730179 CHEMBL3730179 | C30H33NO | 423.6 | 2 / 1 | 5.7 | No |
| 528784 | 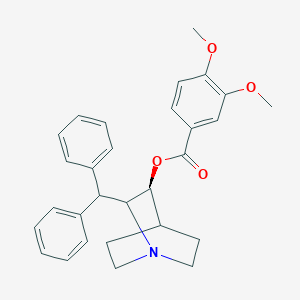 CHEMBL3727438 CHEMBL3727438 | C29H31NO4 | 457.57 | 5 / 0 | 5.8 | No |
| 529226 | 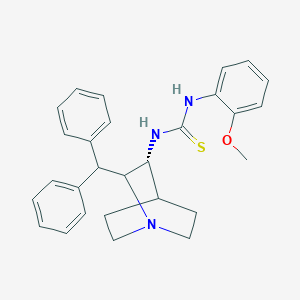 CHEMBL3728009 CHEMBL3728009 | C28H31N3OS | 457.636 | 3 / 2 | 5.6 | No |
| 530408 | 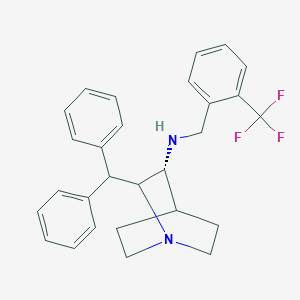 CHEMBL3728505 CHEMBL3728505 | C28H29F3N2 | 450.549 | 5 / 1 | 6.3 | No |
| 530835 | 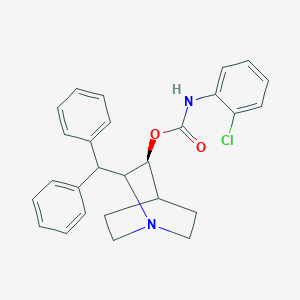 CHEMBL3733177 CHEMBL3733177 | C27H27ClN2O2 | 446.975 | 3 / 1 | 6.2 | No |
| 530836 | 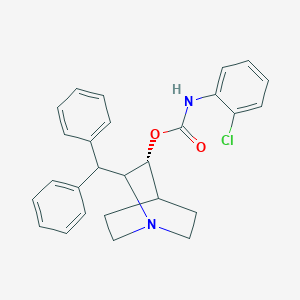 CHEMBL3730599 CHEMBL3730599 | C27H27ClN2O2 | 446.975 | 3 / 1 | 6.2 | No |
| 455205 | 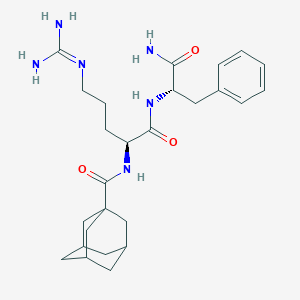 RF 9 RF 9 | C26H38N6O3 | 482.629 | 4 / 5 | 1.4 | Yes |
| 531242 | 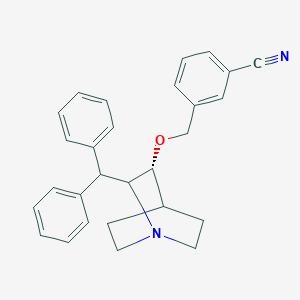 CHEMBL3729794 CHEMBL3729794 | C28H28N2O | 408.545 | 3 / 0 | 5.4 | No |
| 531821 | 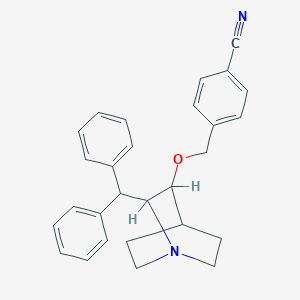 CHEMBL145103 CHEMBL145103 | C28H28N2O | 408.545 | 3 / 0 | 5.4 | No |
| 532327 | 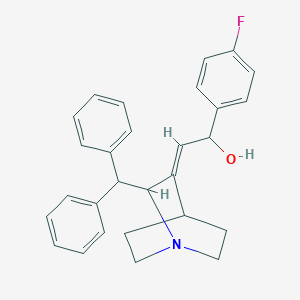 CHEMBL3727861 CHEMBL3727861 | C28H28FNO | 413.536 | 3 / 1 | 5.1 | No |
| 532425 | 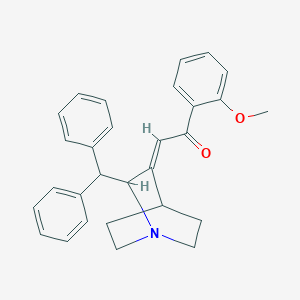 887109-77-5 887109-77-5 | C29H29NO2 | 423.556 | 3 / 0 | 5.3 | No |
| 532548 | 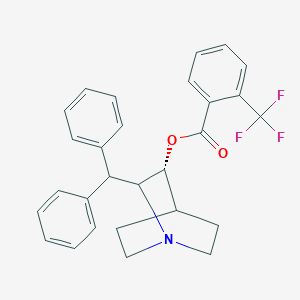 CHEMBL3731844 CHEMBL3731844 | C28H26F3NO2 | 465.516 | 6 / 0 | 6.8 | No |
| 458062 | 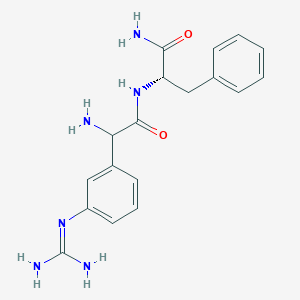 CHEMBL3343992 CHEMBL3343992 | C18H22N6O2 | 354.414 | 4 / 5 | -0.3 | Yes |
| 458448 | 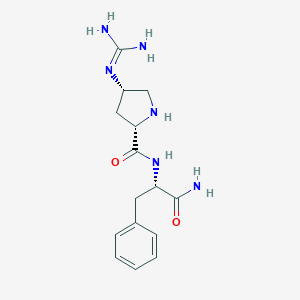 CHEMBL3343987 CHEMBL3343987 | C15H22N6O2 | 318.381 | 4 / 5 | -1.3 | Yes |
| 458964 | 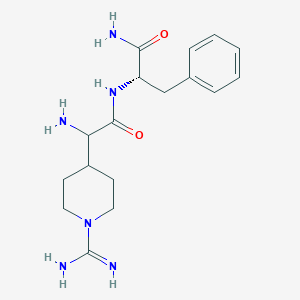 CHEMBL3343988 CHEMBL3343988 | C17H26N6O2 | 346.435 | 4 / 5 | -0.6 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218