

You can:
| Name | Oxoeicosanoid receptor 1 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | OXER1 |
| Synonym | R527 oxoeicosanoid (OXE) receptor 1 OXE receptor hGPCR48 GPR170 [ Show all ] |
| Disease | N/A |
| Length | 423 |
| Amino acid sequence | MLCHRGGQLIVPIIPLCPEHSCRGRRLQNLLSGPWPKQPMELHNLSSPSPSLSSSVLPPSFSPSPSSAPSAFTTVGGSSGGPCHPTSSSLVSAFLAPILALEFVLGLVGNSLALFIFCIHTRPWTSNTVFLVSLVAADFLLISNLPLRVDYYLLHETWRFGAAACKVNLFMLSTNRTASVVFLTAIALNRYLKVVQPHHVLSRASVGAAARVAGGLWVGILLLNGHLLLSTFSGPSCLSYRVGTKPSASLRWHQALYLLEFFLPLALILFAIVSIGLTIRNRGLGGQAGPQRAMRVLAMVVAVYTICFLPSIIFGMASMVAFWLSACRSLDLCTQLFHGSLAFTYLNSVLDPVLYCFSSPNFLHQSRALLGLTRGRQGPVSDESSYQPSRQWRYREASRKAEAIGKLKVQGEVSLEKEGSSQG |
| UniProt | Q8TDS5 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q8TDS5 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q8TDS5. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL1628461 |
| IUPHAR | 271 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 3884 | 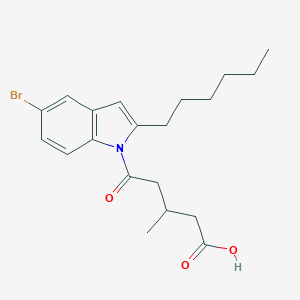 CHEMBL3115784 CHEMBL3115784 | C20H26BrNO3 | 408.336 | 3 / 1 | 5.6 | No |
| 5392 | 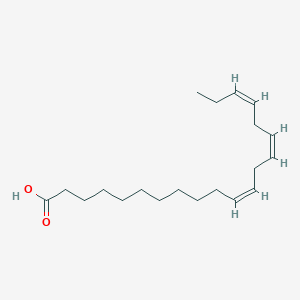 Eicosatrienoic acid Eicosatrienoic acid | C20H34O2 | 306.49 | 2 / 1 | 6.9 | No |
| 11310 | 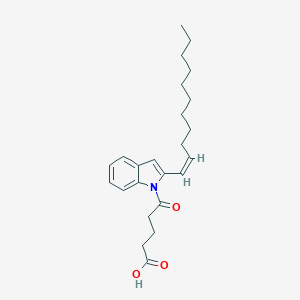 CHEMBL3115800 CHEMBL3115800 | C24H33NO3 | 383.532 | 3 / 1 | 6.9 | No |
| 45909 | 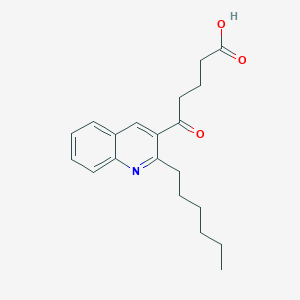 CHEMBL3115773 CHEMBL3115773 | C20H25NO3 | 327.424 | 4 / 1 | 4.5 | Yes |
| 47160 | 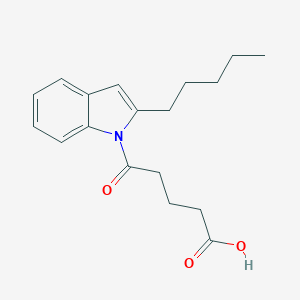 CHEMBL3115795 CHEMBL3115795 | C18H23NO3 | 301.386 | 3 / 1 | 3.9 | Yes |
| 553599 | 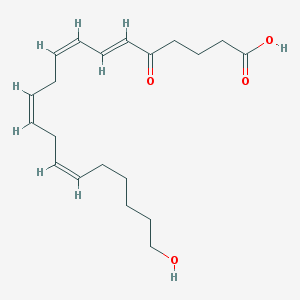 5-oxo-20-HETE 5-oxo-20-HETE | C20H30O4 | 334.456 | 4 / 2 | 3.5 | Yes |
| 68666 | 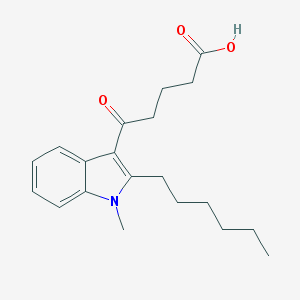 CHEMBL3115789 CHEMBL3115789 | C20H27NO3 | 329.44 | 3 / 1 | 4.2 | Yes |
| 446265 | 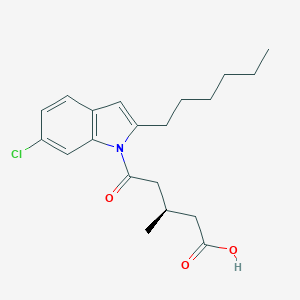 CHEMBL3341973 CHEMBL3341973 | C20H26ClNO3 | 363.882 | 3 / 1 | 5.5 | No |
| 116071 | 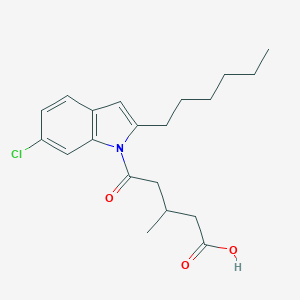 CHEMBL3115774 CHEMBL3115774 | C20H26ClNO3 | 363.882 | 3 / 1 | 5.5 | No |
| 118811 | 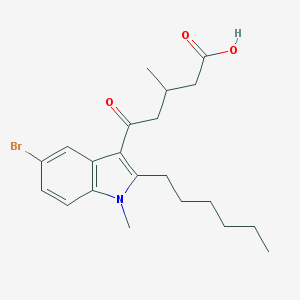 CHEMBL3115793 CHEMBL3115793 | C21H28BrNO3 | 422.363 | 3 / 1 | 5.4 | No |
| 119816 | 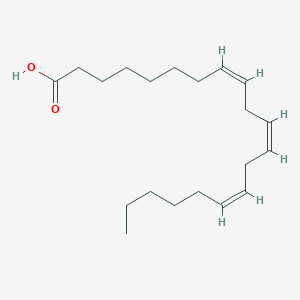 Dihomo-gamma-linolenic acid Dihomo-gamma-linolenic acid | C20H34O2 | 306.49 | 2 / 1 | 7.2 | No |
| 134066 | 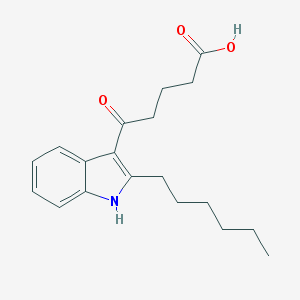 CHEMBL3115788 CHEMBL3115788 | C19H25NO3 | 315.413 | 3 / 2 | 4.3 | Yes |
| 138015 | 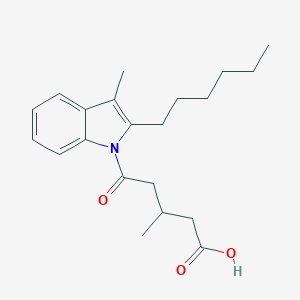 CHEMBL3115783 CHEMBL3115783 | C21H29NO3 | 343.467 | 3 / 1 | 5.3 | No |
| 146452 | 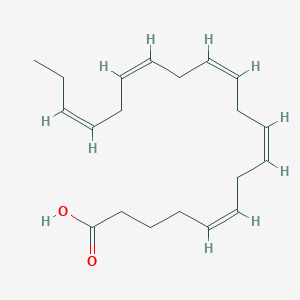 Eicosapentaenoic acid Eicosapentaenoic acid | C20H30O2 | 302.458 | 2 / 1 | 5.6 | No |
| 147553 | 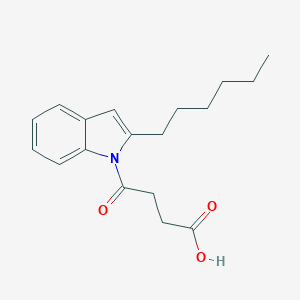 CHEMBL3115779 CHEMBL3115779 | C18H23NO3 | 301.386 | 3 / 1 | 4.1 | Yes |
| 554048 | 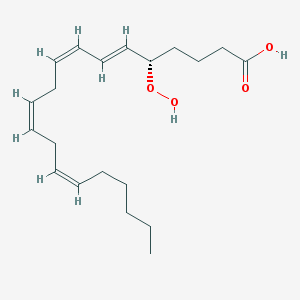 5S-HpETE 5S-HpETE | C20H32O4 | 336.472 | 4 / 2 | 5.3 | No |
| 167555 | 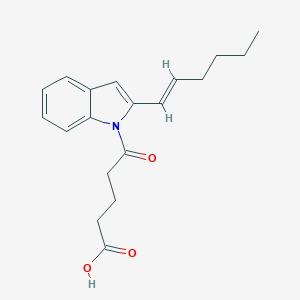 CHEMBL3115796 CHEMBL3115796 | C19H23NO3 | 313.397 | 3 / 1 | 4.2 | Yes |
| 554127 | 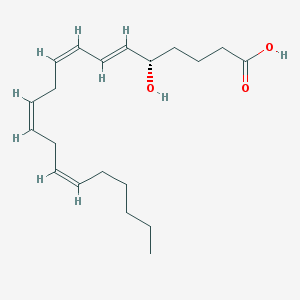 5(S)-HETE 5(S)-HETE | C20H32O3 | 320.473 | 3 / 2 | 5.2 | No |
| 554152 | 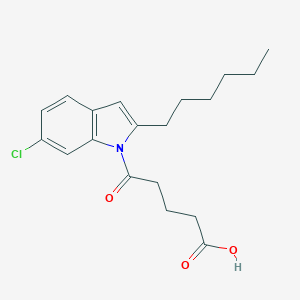 compound 39 [PMID: 23581530] compound 39 [PMID: 23581530] | C19H24ClNO3 | 349.855 | 3 / 1 | 5.1 | No |
| 185052 | 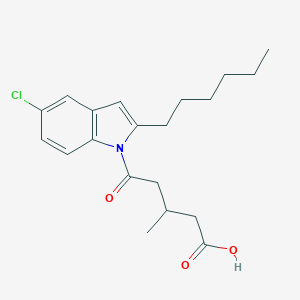 CHEMBL3115785 CHEMBL3115785 | C20H26ClNO3 | 363.882 | 3 / 1 | 5.5 | No |
| 190177 | 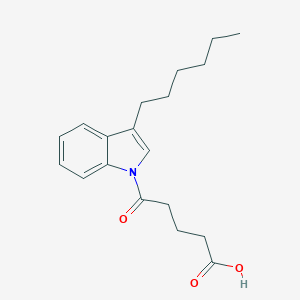 CHEMBL2332557 CHEMBL2332557 | C19H25NO3 | 315.413 | 3 / 1 | 4.6 | Yes |
| 201455 | 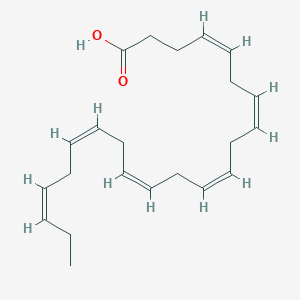 Docosahexaenoic acid Docosahexaenoic acid | C22H32O2 | 328.496 | 2 / 1 | 6.2 | No |
| 203203 | 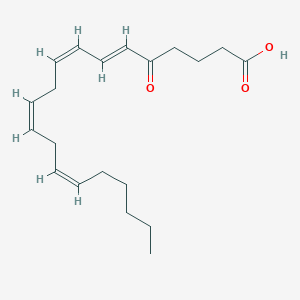 5-Oxo-ETE 5-Oxo-ETE | C20H30O3 | 318.457 | 3 / 1 | 5.1 | No |
| 208777 | 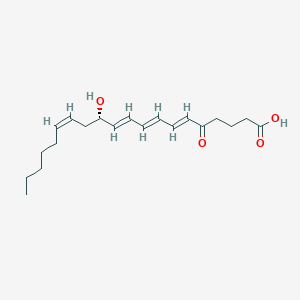 CHEMBL3115777 CHEMBL3115777 | C20H30O4 | 334.456 | 4 / 2 | 4.0 | Yes |
| 208778 | 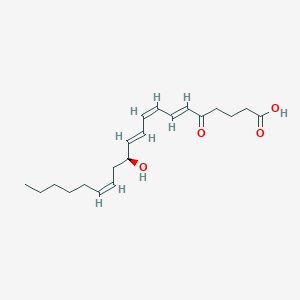 5-oxo-12-HETE 5-oxo-12-HETE | C20H30O4 | 334.456 | 4 / 2 | 4.0 | Yes |
| 234513 | 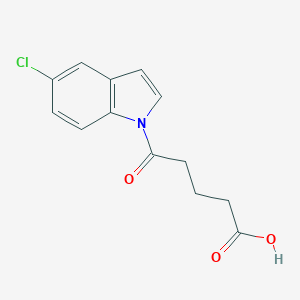 CHEMBL2332561 CHEMBL2332561 | C13H12ClNO3 | 265.693 | 3 / 1 | 2.3 | Yes |
| 249354 | 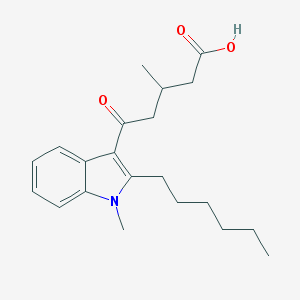 CHEMBL3115791 CHEMBL3115791 | C21H29NO3 | 343.467 | 3 / 1 | 4.7 | Yes |
| 451803 | 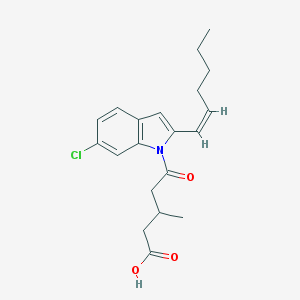 CHEMBL3311230 CHEMBL3311230 | C20H24ClNO3 | 361.866 | 3 / 1 | 5.3 | No |
| 451804 | 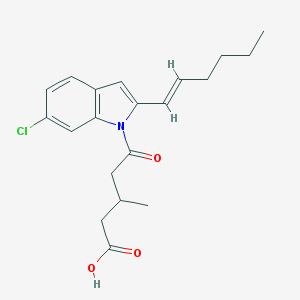 CHEMBL3311229 CHEMBL3311229 | C20H24ClNO3 | 361.866 | 3 / 1 | 5.3 | No |
| 267654 | 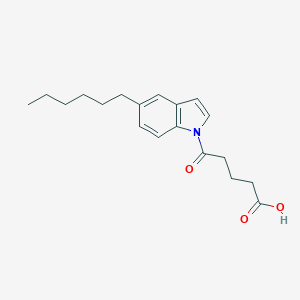 CHEMBL2332558 CHEMBL2332558 | C19H25NO3 | 315.413 | 3 / 1 | 4.6 | Yes |
| 285249 | 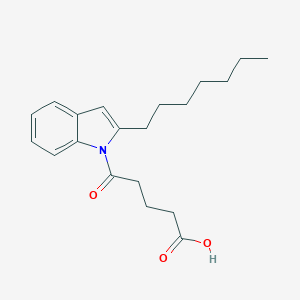 CHEMBL3115797 CHEMBL3115797 | C20H27NO3 | 329.44 | 3 / 1 | 5.0 | Yes |
| 288543 | 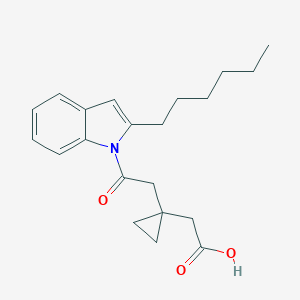 CHEMBL3115780 CHEMBL3115780 | C21H27NO3 | 341.451 | 3 / 1 | 5.1 | No |
| 289576 | 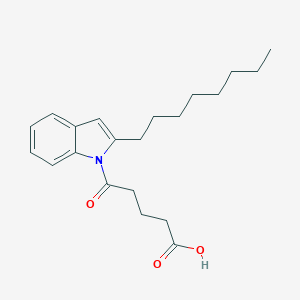 CHEMBL3115798 CHEMBL3115798 | C21H29NO3 | 343.467 | 3 / 1 | 5.6 | No |
| 294219 | 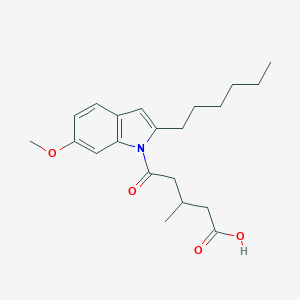 CHEMBL3115787 CHEMBL3115787 | C21H29NO4 | 359.466 | 4 / 1 | 4.9 | Yes |
| 297218 | 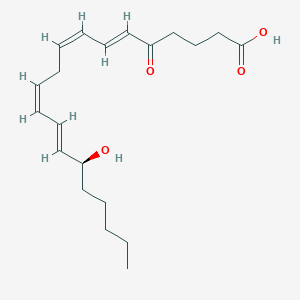 5-Oxo-15-hete 5-Oxo-15-hete | C20H30O4 | 334.456 | 4 / 2 | 3.9 | Yes |
| 298139 | 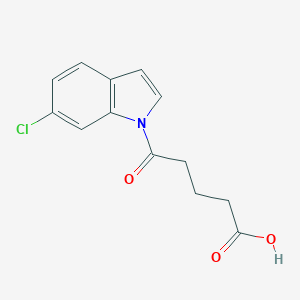 CHEMBL2332562 CHEMBL2332562 | C13H12ClNO3 | 265.693 | 3 / 1 | 2.3 | Yes |
| 305524 | 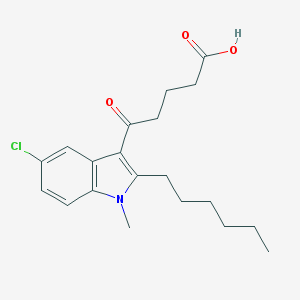 CHEMBL3115790 CHEMBL3115790 | C20H26ClNO3 | 363.882 | 3 / 1 | 4.9 | Yes |
| 310339 | 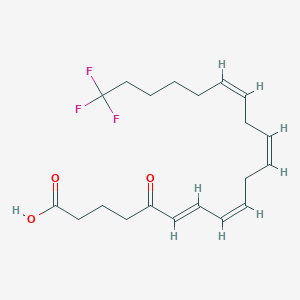 CHEMBL1760545 CHEMBL1760545 | C20H27F3O3 | 372.428 | 6 / 1 | 5.3 | No |
| 322423 | 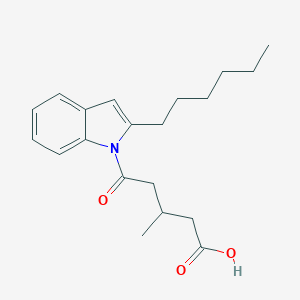 CHEMBL3115782 CHEMBL3115782 | C20H27NO3 | 329.44 | 3 / 1 | 4.9 | Yes |
| 322905 | 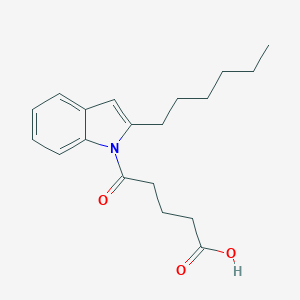 CHEMBL2332567 CHEMBL2332567 | C19H25NO3 | 315.413 | 3 / 1 | 4.5 | Yes |
| 554953 | 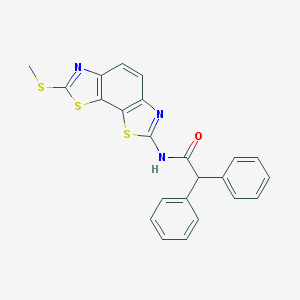 Gue 1654 Gue 1654 | C23H17N3OS3 | 447.589 | 6 / 1 | 6.5 | No |
| 554964 | 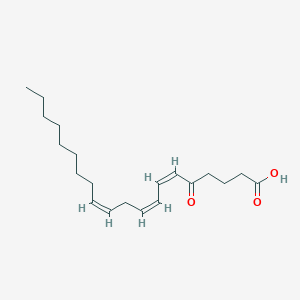 5-oxo-ETrE 5-oxo-ETrE | C20H32O3 | 320.473 | 3 / 1 | 5.8 | No |
| 350047 | 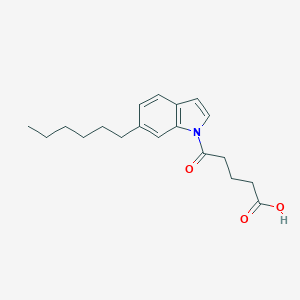 CHEMBL2332559 CHEMBL2332559 | C19H25NO3 | 315.413 | 3 / 1 | 4.6 | Yes |
| 387271 | 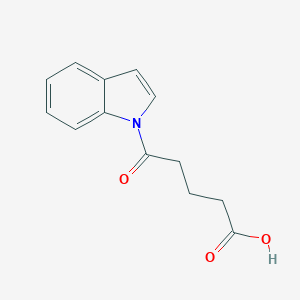 CHEMBL2332566 CHEMBL2332566 | C13H13NO3 | 231.251 | 3 / 1 | 1.7 | Yes |
| 457170 | 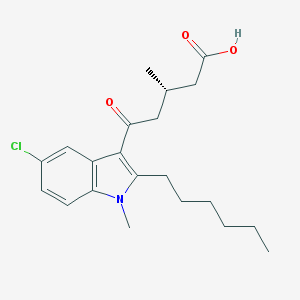 CHEMBL3341972 CHEMBL3341972 | C21H28ClNO3 | 377.909 | 3 / 1 | 5.4 | No |
| 457171 | 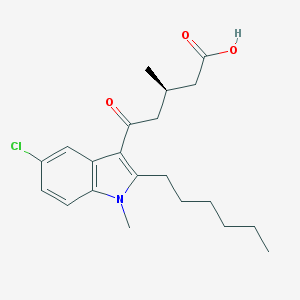 CHEMBL3341971 CHEMBL3341971 | C21H28ClNO3 | 377.909 | 3 / 1 | 5.4 | No |
| 389226 | 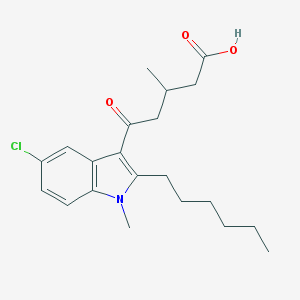 CHEMBL3115775 CHEMBL3115775 | C21H28ClNO3 | 377.909 | 3 / 1 | 5.4 | No |
| 394130 | 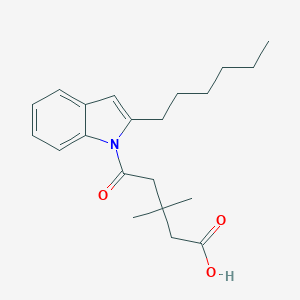 CHEMBL3115781 CHEMBL3115781 | C21H29NO3 | 343.467 | 3 / 1 | 5.3 | No |
| 405748 | 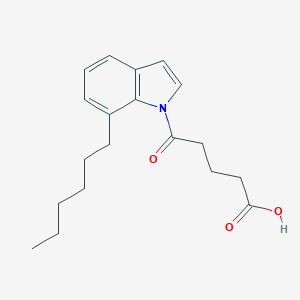 CHEMBL2332560 CHEMBL2332560 | C19H25NO3 | 315.413 | 3 / 1 | 4.6 | Yes |
| 409965 | 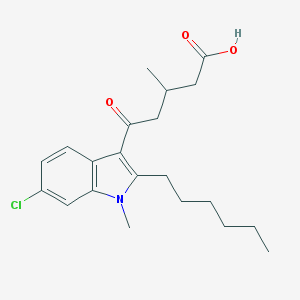 CHEMBL3115792 CHEMBL3115792 | C21H28ClNO3 | 377.909 | 3 / 1 | 5.4 | No |
| 411556 | 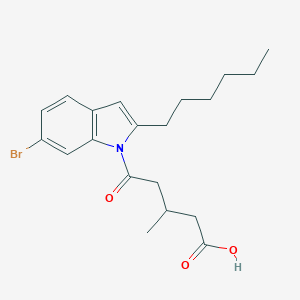 CHEMBL3115786 CHEMBL3115786 | C20H26BrNO3 | 408.336 | 3 / 1 | 5.6 | No |
| 411850 | 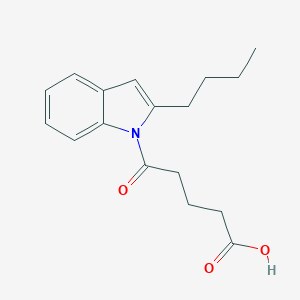 CHEMBL3115794 CHEMBL3115794 | C17H21NO3 | 287.359 | 3 / 1 | 3.4 | Yes |
| 416608 | 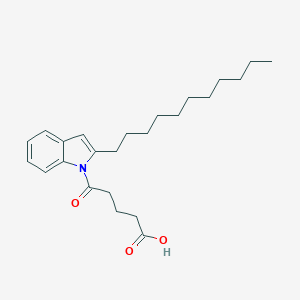 CHEMBL3115799 CHEMBL3115799 | C24H35NO3 | 385.548 | 3 / 1 | 7.2 | No |
| 555373 | 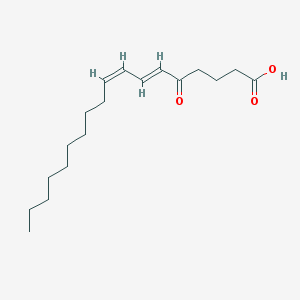 5-oxo-ODE 5-oxo-ODE | C18H30O3 | 294.435 | 3 / 1 | 5.4 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218